https://github.com/arm61/lipids_at_airdes
ESI for "Bayesian determination of the effect of a deep eutectic solvent on the structure of lipid monolayers"
Science Score: 23.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
○CITATION.cff file
-
○codemeta.json file
-
○.zenodo.json file
-
✓DOI references
Found 26 DOI reference(s) in README -
✓Academic publication links
Links to: arxiv.org, aps.org, zenodo.org -
○Academic email domains
-
○Institutional organization owner
-
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (9.6%) to scientific vocabulary
Repository
ESI for "Bayesian determination of the effect of a deep eutectic solvent on the structure of lipid monolayers"
Basic Info
Statistics
- Stars: 2
- Watchers: 3
- Forks: 1
- Open Issues: 0
- Releases: 0
Metadata Files
README.md
ESI for "Bayesian determination of the effect of a deep eutectic solvent on the structure of lipid monolayers"
Andrew R. McCluskey*, Adrian Sanchez-Fernandez, Karen J. Edler, Stephen C. Parker, Andrew J. Jackson, Richard A. Campbell, and Thomas Arnold*.
*a.r.mccluskey@bath.ac.uk/andrew.mccluskey@diamond.ac.uk & tom.arnold@esss.se
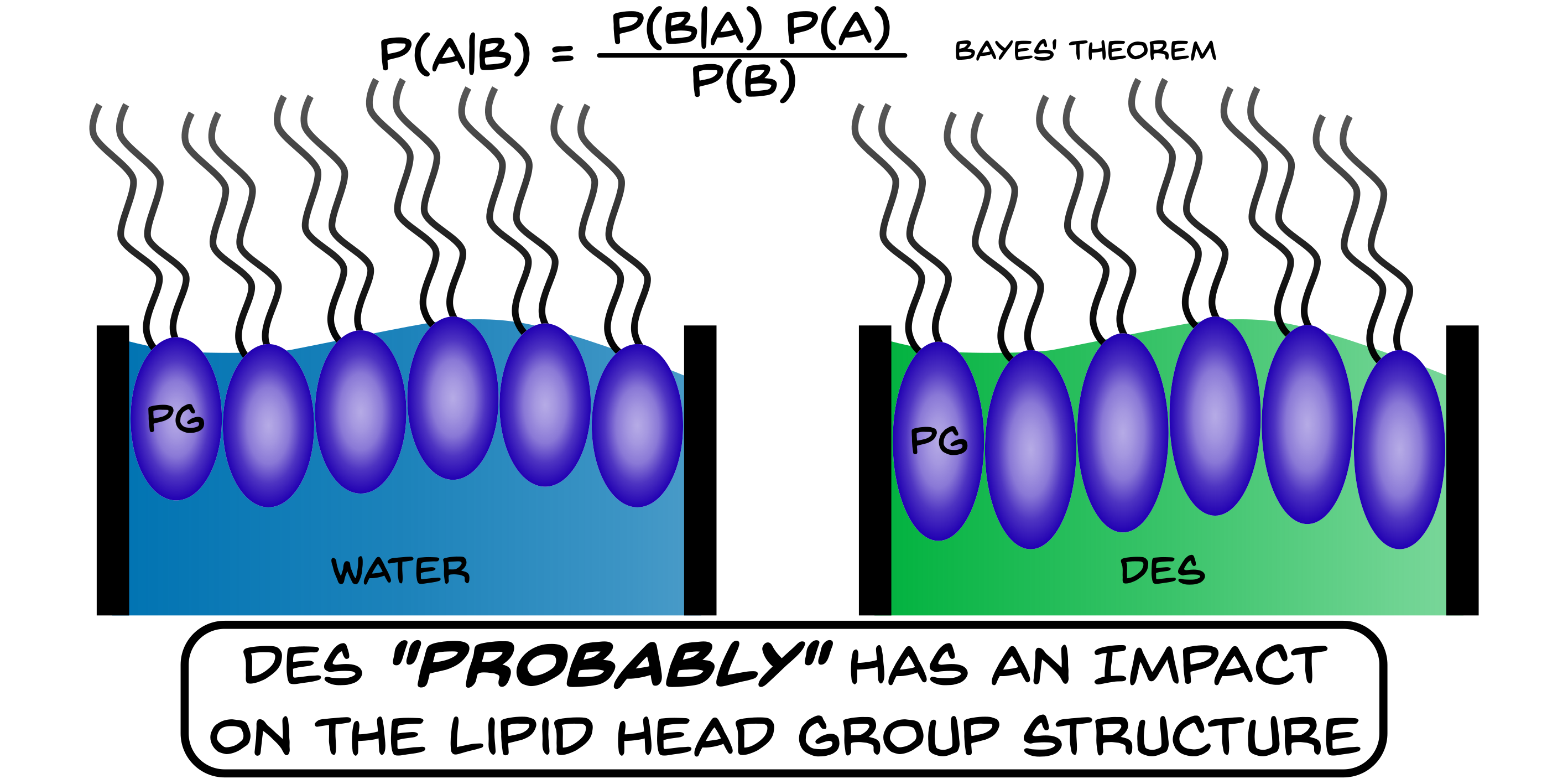 A novel reflectometry analysis method reveals the structure of lipid monolayers at the air-DES interface.
A novel reflectometry analysis method reveals the structure of lipid monolayers at the air-DES interface.
This is the electronic supplementary information (ESI) associated with the publication "Bayesian determination of the effect of a deep eutectic solvent on the structure of lipid monolayers". This ESI provides full details of the analyses performed in the work and access to an automated analysis workflow, through this we aim to provide better access to analysis reproducibility. The Supplementary Information document can be found in the reports folder, alongside a preprint copy of the publication. For more information about reproducible workflows in Python, check out Tania Allard's talk from Pycon2018.
Data
The reduced X-ray and neutron reflectometry data can be obtained from the University of Bath Research Data Archive.
DOI: 10.15125/BATH-00548
The full neutron reflectometry data can be obtained from the ILL Data Archive.
DOI: 10.5291/ILL-DATA.9-13-612
Analysis
This ESI aims to provide a fully reproducible workflow to the data analysis presented within the paper.
Requirements:
- anaconda or miniconda python
- make
- REVTeX
The supplied Makefile, will reproduce all of the analysis, plot the figures, and build a preprint version of the paper (reports/paper.pdf) when run. Be aware that the analyses within this work are non-trivial and take many hours to run so use caution before re-running.
If you still want to re-run all of the analysis, please download the experimental data zip file, and unzip it (in the lipids_at_airdes directory) using the following command:
unzip lipids_at_airdes_data.zip
Then run the following commands:
``` conda create --name paper_env python
source activate paper_env
pip install --upgrade pip
pip install -r config/requirements.txt
make clean # this will remove all of the output from previous runs
make ```
Figures
PDF versions of the figures, can be found in the reports/figures directory.
Citations
Paper: A. R. McCluskey, A. Sanchez-Fernandez, K. J. Edler, S. C. Parker, A. J. Jackson, R. A. Campbell and T. Arnold, Phys. Chem. Chem. Phys., 2019, 21(11), 61336141.
ESI: A. R. McCluskey, A. Sanchez-Fernandez, K. J. Edler, S. C. Parker, A. J. Jackson, R. A. Campbell and T. Arnold, lipidsatairdes (Version 1.0), http://doi.org/10.5281/zenodo.2577796.
Data: A. R. McCluskey, A. Sanchez-Fernandez, K. J. Edler, S. C. Parker, A. J. Jackson, R. A. Campbell and T. Arnold, Dataset for Bayesian determination of the effect of a deep eutectic solvent on the structure of lipid monolayers, https://doi.org/10.15125/BATH-00548.
BibTeX
``` @article{lipidsatairdes_paper, title={Bayesian determination of the effect of a deep eutectic solvent on the structure of lipid monolayers}, volume={21}, DOI={10.1039/C9CP00203K}, number={11}, journal={Phys. Chem. Chem. Phys.}, author={McCluskey, A. R. and Sanchez-Fernandez, A. and Edler, K. J. and Parker, S. C. and Jackson, A. J. and Campbell, R. A. and Arnold, T.}, year={2019}, pages={61336141} }
@misc{lipidsatairdesesi, title={lipidsat_airdes (Version 1.0)}, url={http://doi.org/10.5281/zenodo.2577796}, author={McCluskey, A. R. and Sanchez-Fernandez, A. and Edler, K. J. and Parker, S. C. and Jackson, A. J. and Campbell, R. A. and Arnold, T.}, year={2019} }
@misc{lipidsatairdes_data, title={Dataset for Bayesian determination of the effect of a deep eutectic solvent on the structure of lipid monolayers}, url={https://doi.org/10.15125/BATH-00548}, author={McCluskey, A. R. and Sanchez-Fernandez, A. and Edler, K. J. and Parker, S. C. and Jackson, A. J. and Campbell, R. A. and Arnold, T.}, year={2019} } ```
Acknowledgements
A. R. M. is grateful to the University of Bath and Diamond Light Source for co-funding a studentship (Studentship Number STU0149). The authors thank the European Spallation Source and the University of Bath Alumni Fund for supporting A. S.-F.
Project Organization
.
AUTHORS.md
LICENSE # CC-BY-SA-4.0 License
README.md # You are here
Makefile # Makefile to outline workflow
output # Files and data output by analysis scripts
dlpc #
dmpc #
dmpg #
dppc #
config # requirements.txt file
notebooks # Notebooks for analysis
reports # Paper and ESI
figures
src
models # mol_vol.py custom model for refnx
tools # helper.py script
visualization # Plotting scripts
Owner
- Name: Andrew McCluskey
- Login: arm61
- Kind: user
- Location: Copenhagen
- Company: European Spallation Source
- Website: https://mccluskey.scot
- Repositories: 8
- Profile: https://github.com/arm61
instrument data scientist @essneutron (he/him)
GitHub Events
Total
Last Year
Issues and Pull Requests
Last synced: about 1 year ago
All Time
- Total issues: 0
- Total pull requests: 5
- Average time to close issues: N/A
- Average time to close pull requests: about 4 hours
- Total issue authors: 0
- Total pull request authors: 3
- Average comments per issue: 0
- Average comments per pull request: 0.6
- Merged pull requests: 4
- Bot issues: 0
- Bot pull requests: 1
Past Year
- Issues: 0
- Pull requests: 0
- Average time to close issues: N/A
- Average time to close pull requests: N/A
- Issue authors: 0
- Pull request authors: 0
- Average comments per issue: 0
- Average comments per pull request: 0
- Merged pull requests: 0
- Bot issues: 0
- Bot pull requests: 0
Top Authors
Issue Authors
Pull Request Authors
- ajj (2)
- arm61 (2)
- dependabot[bot] (1)
Top Labels
Issue Labels
Pull Request Labels
Dependencies
- Cython ==0.29.2
- corner ==2.0.1
- h5py ==2.9.0
- jupyter_contrib_nbextensions ==0.5.1
- matplotlib ==3.0.2
- numpy ==1.22.0
- pandas ==0.23.4
- periodictable ==1.5.0
- refnx ==0.0.17
- scipy ==1.1.0
- seaborn ==0.9.0
- tqdm ==4.29.0
- xlrd ==1.2.0


