Science Score: 54.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
✓CITATION.cff file
Found CITATION.cff file -
✓codemeta.json file
Found codemeta.json file -
✓.zenodo.json file
Found .zenodo.json file -
○DOI references
-
○Academic publication links
-
✓Committers with academic emails
1 of 4 committers (25.0%) from academic institutions -
○Institutional organization owner
-
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (12.4%) to scientific vocabulary
Keywords
Repository
Ramachandran plotting tool
Basic Info
Statistics
- Stars: 20
- Watchers: 1
- Forks: 10
- Open Issues: 0
- Releases: 1
Topics
Metadata Files
README.md
Ramachandran plotting tool
Draws a Ramachandran plot based on the input PDB file (e.g. 1mbn.pdb). Makes use of a Gaussian KDE (kernel density
estimation) to plot the density of favoured torsion angles (φ and ψ).
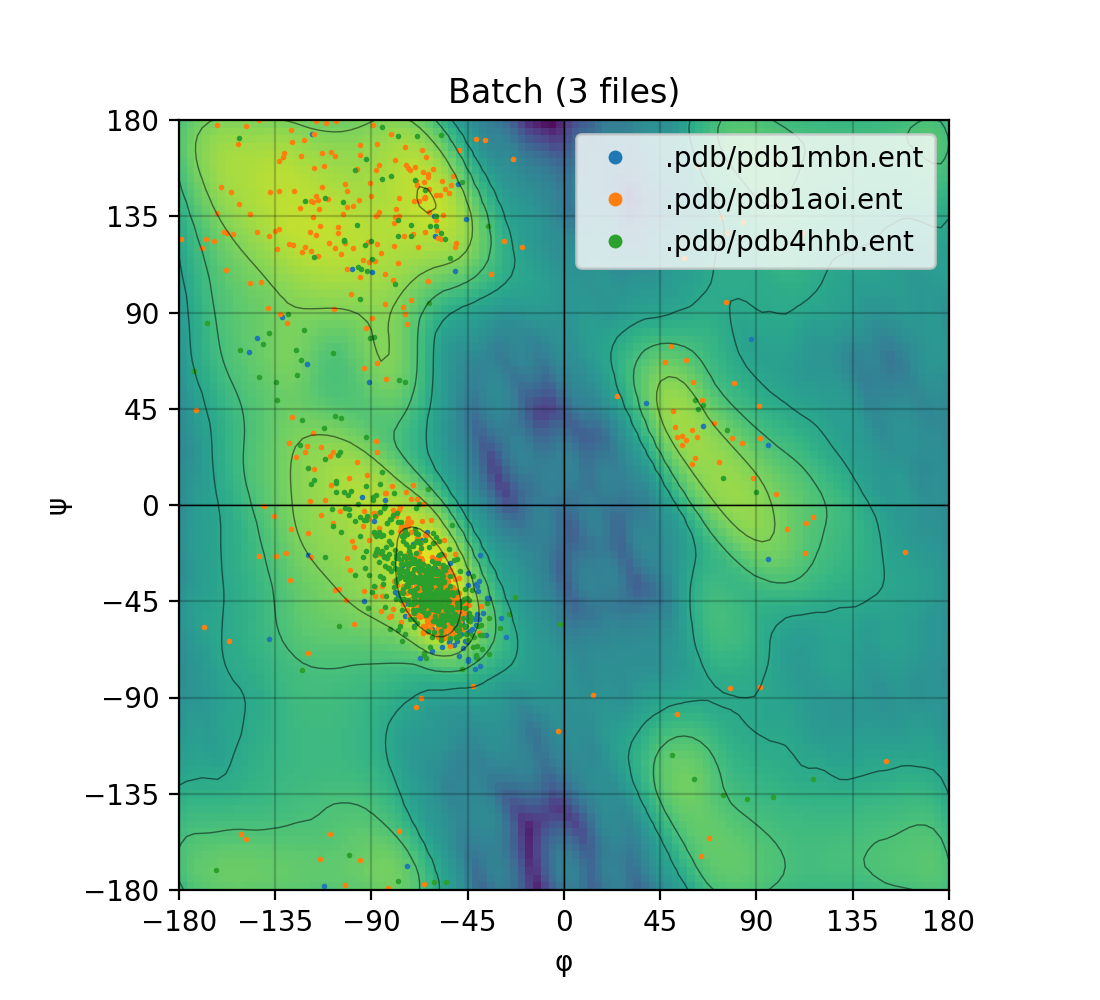
Single mode | Batch mode
:-------------------------:|:-------------------------:
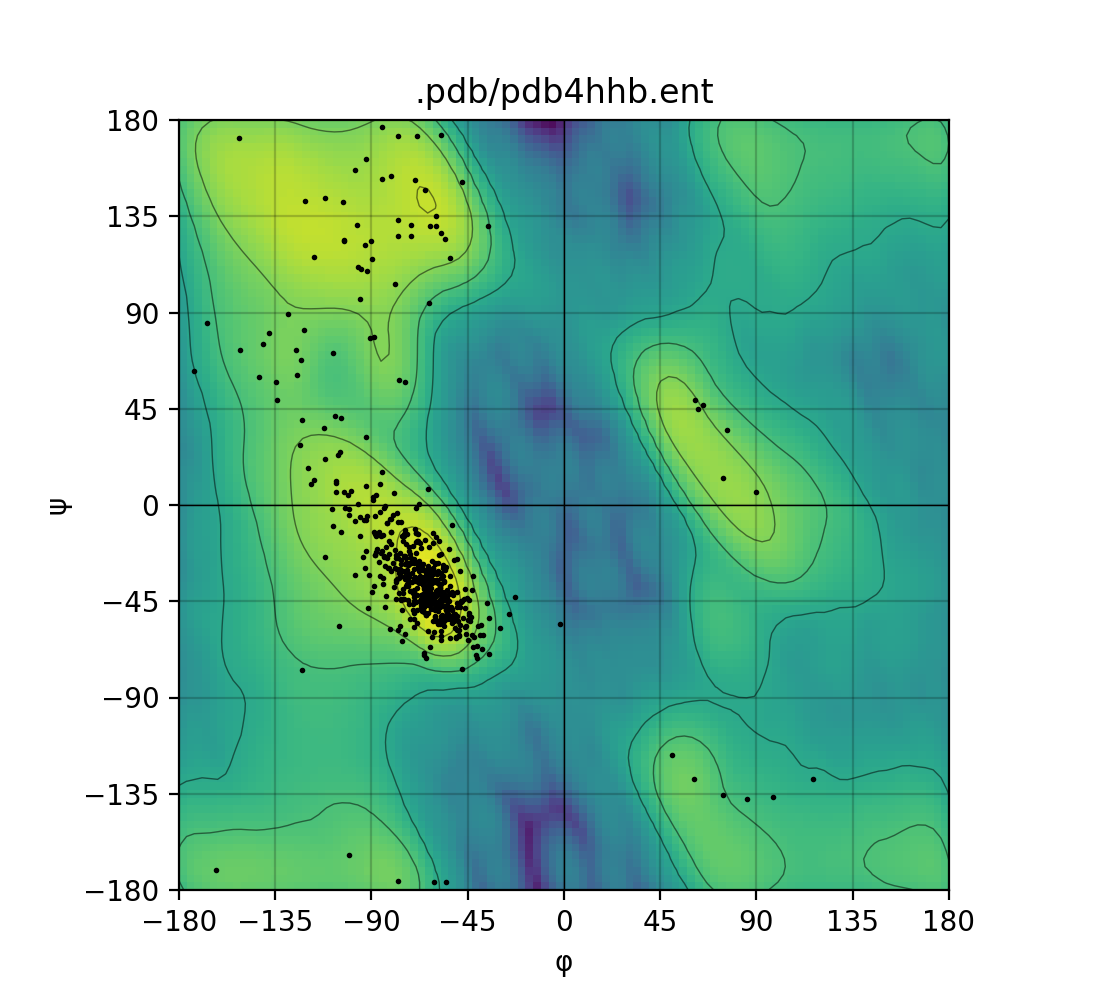 |
| 
Installation
RamachanDraw is hosted on PyPi.
bash
pip install ramachandraw
Usage
RamachanDraw includes useful functions to effortlessly draw a Ramachandran plot.
Fetch the PDB file
To draw a Ramachandran plot, we need a PDB file. RamachanDraw conveniently includes a function to automatically fetch and locally store the PDB file for the given PDB id.
Example
```python from ramachandraw.utils import fetch_pdb
fetchpdb(pdbid, verbose) ```
pdb_id (str|list|tuple): PDB id(s) corresponding to the PDB file(s) to be downloadedverbose (bool)(optional): set the verbosity, defaults toFalse- Returns: path(s) to PDB file(s)
Extract φ and ψ torsion angles
RamachanDraw extracts the φ and ψ torsion angles from the PDB file by taking advantage of the Biopython module. Additionally, aminoacid residues that were not drawn on the plot can be extract using the return_ignored argument.
Example
```python from ramachandraw.parser import getphipsi
phipsi(pdbfilepath, prune, hide_warnings) ```
pdb_id (str|list|tuple): PDB filepath(s)prune (bool)(optional): prunes aminoacids with missing torsion angle(s), defaults toTruehide_warnings (bool)(optional): sets the verbosity of the PDB structure parser, defaults toTrue- Returns: Dictionary with keys as aminoacid residues and values as (φ, ψ) angles.
Ramachandran plot
Makes use of the matplotlib module to draw a highly customizable Ramachandran plot.
Example
```python from ramachandraw.utils import plot
plot(pdb_filepath, cmap="viridis", alpha=0.75, dpi=100, save=True, show=False, filename="plot.png") ```
pdb_file (str|list|tuple): PDB id(s) corresponding to the PDB entry to be downloadedcmap (str)(optional): colormap to be used (frommatplotlib) for the density of the favoured ("allowed") regions; default is viridis.alpha (float)(optional): sets the opacity of the colormap (value between 0-1); default is 0.75.dpi (int)(optional): resolution (in dots per inch); default is100.save (bool)(optional):True: saves the plot locally; default is True.
show (bool)(optional):True: shows the plot using the Qt5Agg backend; default is False.
filename (str)(optional): filename to be used in case the plot is saved (i.e.save=True); default isplot.png.- Returns: Ramachandran plot (
matplotlib.axes.Axesobject) that can be further customized if needed
Example
Herein you will find an example from the PDB id corresponding to the myoglobin entry: 1MBN - in the Protein Data Bank.
Single PDB
```python from ramachandraw.parser import getphipsi from ramachandraw.utils import fetch_pdb, plot
PDB id
pdb_id = "1mbn"
Draw the Ramachandran plot
plot(fetchpdb(pdbid))
Generate a dictionary to store the (phi, psi) torsion angles
torsionangles = getphipsi(fetchpdb(pdb_id)) ```
Batch of PDBs
```python from ramachandraw.parser import getphipsi from ramachandraw.utils import fetch_pdb, plot
PDB id
pdb_ids = ["1mbn", "4hhb"]
Draw the Ramachandran plot
plot(fetchpdb(pdbids))
Generate a list of dictionaries to store the (phi, psi) torsion angles
torsionangles = getphipsi(fetchpdb(pdb_ids)) ```
Contributing
Feedback and constructive criticism is welcome. If necessary, open an issue in the issues tab.
License
Owner
- Name: Alexandre Cirilo
- Login: alxdrcirilo
- Kind: user
- Location: The Netherlands
- Repositories: 1
- Profile: https://github.com/alxdrcirilo
Citation (CITATION.cff)
# This CITATION.cff file was generated with cffinit.
# Visit https://bit.ly/cffinit to generate yours today!
cff-version: 1.2.0
title: Ramachandraw
version: 1.0.1
message: >-
If you use this software, please cite it using the
metadata from this file.
type: software
authors:
- given-names: Alexandre
family-names: Dias Cirilo
repository-code: 'https://github.com/alxdrcirilo/ramachandraw'
repository-artifact: 'https://pypi.org/project/ramachandraw/'
abstract: Ramachandran plotting tool.
keywords:
- bioinformatics
- ramachandran
- protein structure
license: MIT
GitHub Events
Total
- Issues event: 2
- Issue comment event: 5
Last Year
- Issues event: 2
- Issue comment event: 5
Committers
Last synced: about 3 years ago
All Time
- Total Commits: 24
- Total Committers: 4
- Avg Commits per committer: 6.0
- Development Distribution Score (DDS): 0.417
Top Committers
| Name | Commits | |
|---|---|---|
| alxdrcirilo | a****8@h****m | 14 |
| Alexandre Cirilo | 3****o@u****m | 8 |
| underasail | u****l@y****m | 1 |
| Noah Liguori-Bills | n****s@u****u | 1 |
Committer Domains (Top 20 + Academic)
Issues and Pull Requests
Last synced: 10 months ago
All Time
- Total issues: 6
- Total pull requests: 10
- Average time to close issues: 11 days
- Average time to close pull requests: 22 days
- Total issue authors: 6
- Total pull request authors: 5
- Average comments per issue: 1.83
- Average comments per pull request: 0.0
- Merged pull requests: 10
- Bot issues: 0
- Bot pull requests: 2
Past Year
- Issues: 2
- Pull requests: 3
- Average time to close issues: 2 days
- Average time to close pull requests: 5 minutes
- Issue authors: 2
- Pull request authors: 2
- Average comments per issue: 3.0
- Average comments per pull request: 0.0
- Merged pull requests: 3
- Bot issues: 0
- Bot pull requests: 1
Top Authors
Issue Authors
- 2young-2simple-sometimes-naive (1)
- Wenxi-Zhang (1)
- abhishekiiitdcb (1)
- bittner (1)
- noahlb123 (1)
- SShaminur (1)
Pull Request Authors
- alxdrcirilo (10)
- dependabot[bot] (4)
- Aquaplant (2)
- noahlb123 (1)
- underasail (1)
Top Labels
Issue Labels
Pull Request Labels
Packages
- Total packages: 1
-
Total downloads:
- pypi 237 last-month
- Total dependent packages: 0
- Total dependent repositories: 1
- Total versions: 4
- Total maintainers: 1
pypi.org: ramachandraw
Ramachandran plotting tool
- Documentation: https://ramachandraw.readthedocs.io/
- License: MIT
-
Latest release: 1.0.1
published about 2 years ago
Rankings
Maintainers (1)
Dependencies
- biopython >=1.75
- matplotlib >=3.1.2
- rich >=9.3





