https://github.com/bebatut/docker-galaxy-stable
:whale::bar_chart::books: Docker Images tracking the stable Galaxy releases.
Science Score: 10.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
○CITATION.cff file
-
○codemeta.json file
-
○.zenodo.json file
-
○DOI references
-
✓Academic publication links
Links to: zenodo.org -
○Academic email domains
-
○Institutional organization owner
-
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (12.6%) to scientific vocabulary
Last synced: 6 months ago
·
JSON representation
Repository
:whale::bar_chart::books: Docker Images tracking the stable Galaxy releases.
Basic Info
- Host: GitHub
- Owner: bebatut
- License: mit
- Language: Shell
- Default Branch: master
- Homepage: http://bgruening.github.io/docker-galaxy-stable
- Size: 17.9 MB
Statistics
- Stars: 0
- Watchers: 2
- Forks: 0
- Open Issues: 0
- Releases: 0
Fork of bgruening/docker-galaxy-stable
Created almost 9 years ago
· Last pushed almost 9 years ago
https://github.com/bebatut/docker-galaxy-stable/blob/master/
[](https://zenodo.org/badge/latestdoi/5466/bgruening/docker-galaxy-stable) [](https://travis-ci.org/bgruening/docker-galaxy-stable) [](https://quay.io/repository/bgruening/galaxy) [](https://gitter.im/bgruening/docker-galaxy-stable?utm_source=badge&utm_medium=badge&utm_campaign=pr-badge)   Galaxy Docker Image =================== The [Galaxy](http://www.galaxyproject.org) [Docker](http://www.docker.io) Image is an easy distributable full-fledged Galaxy installation, that can be used for testing, teaching and presenting new tools and features. One of the main goals is to make the access to entire tool suites as easy as possible. Usually, this includes the setup of a public available web-service that needs to be maintained, or that the Tool-user needs to either setup a Galaxy Server by its own or to have Admin access to a local Galaxy server. With docker, tool developers can create their own Image with all dependencies and the user only needs to run it within docker. The Image is based on [Ubuntu 14.04 LTS](http://releases.ubuntu.com/14.04/) and all recommended Galaxy requirements are installed. The following chart should illustrate the [Docker](http://www.docker.io) image hierarchy we have build to make is as easy as possible to build on different layers of our stack and create many exciting Galaxy flavors. 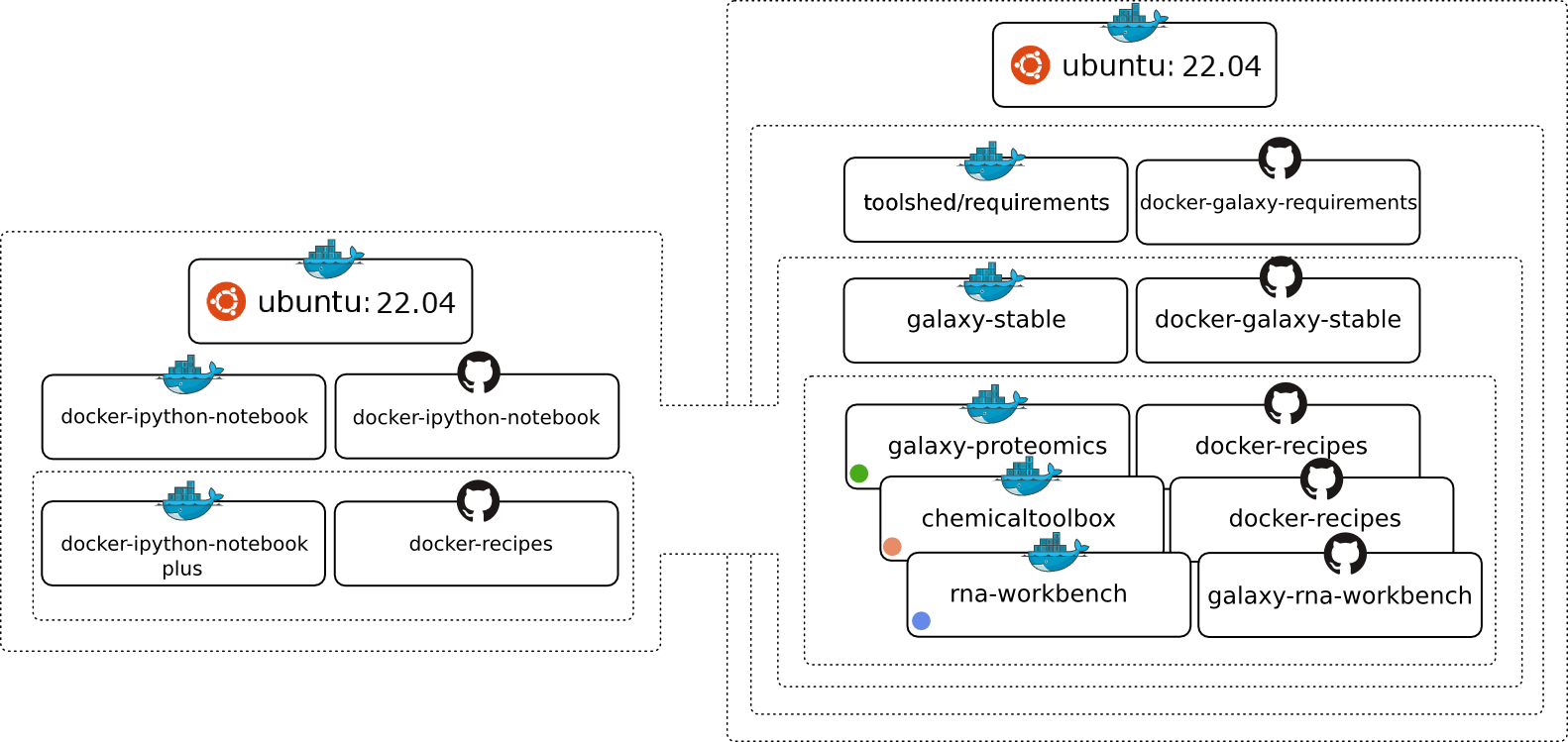 # Table of Contents - [Usage](#Usage) - [Upgrading images](#Upgrading-images) - [Enabling Interactive Environments in Galaxy](#Enabling-Interactive-Environments-in-Galaxy) - [Using passive mode FTP or SFTP](#Using-passive-mode-FTP-or-SFTP) - [Using Parent docker](#Using-Parent-docker) - [Galaxy Report Webapp](#Galaxy-Report-Webapp) - [Galaxy's config settings](#Galaxys-config-settings) - [Personalize your Galaxy](#Personalize-your-Galaxy) - [Deactivating services](#Deactivating-services) - [Restarting Galaxy](#Restarting-Galaxy) - [Advanced Logging](#Advanced-Logging) - [Using an external Slurm cluster](#Using-an-external-Slurm-cluster) - [Using an external Grid Engine cluster](#Using-an-external-Grid-Engine-cluster) - [Tips for Running Jobs Outside the Container](#Tips-for-Running-Jobs-Outside-the-Container) - [Enable Galaxy to use BioContainers (Docker)](#auto-exec-tools-in-docker) - [Magic Environment variables](#Magic-Environment-variables) - [Lite Mode](#Lite-Mode) - [Extending the Docker Image](#Extending-the-Docker-Image) - [List of Galaxy flavours](#List-of-Galaxy-flavours) - [Integrating non-Tool Shed tools into the container](#Integrating-non-Tool-Shed-tools-into-the-container) - [Users & Passwords](#Users-Passwords) - [Development](#Development) - [Requirements](#Requirements) - [History](#History) - [Support & Bug Reports](#Support-Bug-Reports) # Usage [[toc]](#toc) At first you need to install docker. Please follow the [very good instructions](https://docs.docker.com/installation/) from the Docker project. After the successful installation, all you need to do is: ```sh docker run -d -p 8080:80 -p 8021:21 -p 8022:22 bgruening/galaxy-stable ``` I will shortly explain the meaning of all the parameters. For a more detailed description please consult the [docker manual](http://docs.docker.io/), it's really worth reading. Let's start: - `docker run` will run the Image/Container for you. In case you do not have the Container stored locally, docker will download it for you. - `-p 8080:80` will make the port 80 (inside of the container) available on port 8080 on your host. Same holds for port 8021 and 8022, that can be used to transfer data via the FTP or SFTP protocol, respectively. Inside the container a nginx Webserver is running on port 80 and that port can be bound to a local port on your host computer. With this parameter you can access your Galaxy instance via `http://localhost:8080` immediately after executing the command above. If you work with the [Docker Toolbox](https://www.docker.com/products/docker-toolbox) on Mac or Windows, you need to connect to the machine generated by 'Docker Quickstart'. You get its IP address from `docker-machine ls` or from the first line in the terminal, e.g.: `docker is configured to use the default machine with IP 192.168.99.100`. - `bgruening/galaxy-stable` is the Image/Container name, that directs docker to the correct path in the [docker index](https://index.docker.io/u/bgruening/galaxy-stable/). - `-d` will start the docker container in daemon mode. For an interactive session, you can execute: ```sh docker run -i -t -p 8080:80 \ bgruening/galaxy-stable \ /bin/bash ``` and run the `startup` script by yourself, to start PostgreSQL, nginx and Galaxy. Docker images are "read-only", all your changes inside one session will be lost after restart. This mode is useful to present Galaxy to your colleagues or to run workshops with it. To install Tool Shed repositories or to save your data you need to export the calculated data to the host computer. Fortunately, this is as easy as: ```sh docker run -d -p 8080:80 \ -v /home/user/galaxy_storage/:/export/ \ bgruening/galaxy-stable ``` With the additional `-v /home/user/galaxy_storage/:/export/` parameter, Docker will mount the local folder `/home/user/galaxy_storage` into the Container under `/export/`. A `startup.sh` script, that is usually starting nginx, PostgreSQL and Galaxy, will recognize the export directory with one of the following outcomes: - In case of an empty `/export/` directory, it will move the [PostgreSQL](http://www.postgresql.org/) database, the Galaxy database directory, Shed Tools and Tool Dependencies and various config scripts to /export/ and symlink back to the original location. - In case of a non-empty `/export/`, for example if you continue a previous session within the same folder, nothing will be moved, but the symlinks will be created. This enables you to have different export folders for different sessions - means real separation of your different projects. You can also collect and store `/export/` data of Galaxy instances in a dedicated docker [Data volume Container](https://docs.docker.com/engine/userguide/dockervolumes/) created by: ```sh docker create -v /export \ --name galaxy-store \ bgruening/galaxy-stable \ /bin/true ``` To mount this data volume in a Galaxy container, use the `--volumes-from` parameter: ```sh docker run -d -p 8080:80 \ --volumes-from galaxy-store \ bgruening/galaxy-stable ``` This also allows for data separation, but keeps everything encapsulated within the docker engine (e.g. on OS X within your `$HOME/.docker` folder - easy to backup, archive and restore. This approach, albeit at the expense of disk space, avoids the problems with permissions [reported](https://github.com/bgruening/docker-galaxy-stable/issues/68) for data export on non-Linux hosts. ## Upgrading images [[toc]](#toc) We will release a new version of this image concurrent with every new Galaxy release. For upgrading an image to a new version we have assembled a few hints for you. Please, take in account that that upgrading may vary depending on your Galaxy installation, and the changes in new versions. Use this example carefully! * Create a test instance with only the database and configuration files. This will allow testing to ensure that things run but won't require copying all of the data. * New unmodified configuration files are always stored in a hidden directory called `.distribution_config`. Use this folder to diff your configurations with the new configuration files shipped with Galaxy. This prevents needing to go through the change log files to find out which new files were added or which new features you can activate. * Note that copying database and datasets can be expensive if you have many GB of data. 1. Download newer version of the Galaxy image ``` $ sudo docker pull bgruening/galaxy-stable ``` 2. Stop and rename the current galaxy container ``` $ sudo docker stop galaxy-instance $ sudo docker rename galaxy-instance galaxy-instance-old ``` 3. Rename the data directory (the one that is mounted to /export in the docker) ``` $ sudo mv /data/galaxy-data /data/galaxy-data-old ``` 4. Run a new Galaxy container using newer image and wait while Galaxy generates the default content for /export ``` $ sudo run -p 8080:80 -v /data/galaxy-data:/export --name galaxy-instance bgruening/galaxy-stable ``` 5. Stop the Galaxy container ``` $ sudo docker stop galaxy-instance ``` 6. Replace the content of the postgres database by the old db data ``` $ sudo rm -r /data/galaxy-data/postgresql/ $ sudo rsync -var /data/galaxy-data-old/postgresql/ /data/galaxy-data/postgresql/ ``` 7. Use diff to find changes in the config files (only if you changed any config file). ``` $ cd /data/galaxy-data/.distribution_config $ for f in *; do echo $f; diff $f ../../galaxy-data-old/galaxy-central/config/$f; read; done ``` 8. Copy all the users' datasets to the new instance ``` $ sudo rsync -var /data/galaxy-data-old/galaxy-central/database/files/* /data/galaxy-data/galaxy-central/da tabase/files/ ``` 9. Copy all the installed tools ``` $ sudo rsync -var /data/galaxy-data-old/tool_deps/* /data/galaxy-data/tool_deps/ $ sudo rsync -var /data/galaxy-data-old/shed_tools/* /data/galaxy-data/shed_tools/ ``` 10. Copy the welcome page and all its files. ``` $ sudo rsync -var /data/galaxy-data-old/welcome* /data/galaxy-data/ ``` 11. Create a auxiliar docker in interactive mode and upgrade the database. ``` $ sudo docker run -it --rm -v /data/galaxy-data:/export bgruening/galaxy-stable /bin/bash # Startup all processes > startup & #Upgrade the database to the most recent version > sh manage_db.sh upgrade #Logout > exit ``` 12. Start the docker and test ``` $ sudo docker start galaxy-instance ``` 13. Clean the old container and image ## Enabling Interactive Environments in Galaxy [[toc]](#toc) Interactive Environments (IE) are sophisticated ways to extend Galaxy with powerful services, like [Jupyter](http://jupyter.org/), in a secure and reproducible way. For this we need to be able to launch Docker containers inside our Galaxy Docker container. At least docker 1.3 is needed on the host system. ```sh docker run -d -p 8080:80 -p 8021:21 -p 8800:8800 \ --privileged=true \ -v /home/user/galaxy_storage/:/export/ \ bgruening/galaxy-stable ``` The port 8800 is the proxy port that is used to handle Interactive Environments. `--privileged` is needed to start docker containers inside docker. If your IE does not open, please make sure you open your Galaxy instance with your hostname or a [FQDN](https://en.wikipedia.org/wiki/Fully_qualified_domain_name), but not with localhost or 127.0.0.1. ## Using passive mode FTP or SFTP [[toc]](#toc) By default, FTP servers running inside of docker containers are not accessible via passive mode FTP, due to not being able to expose extra ports. To circumvent this, you can use the `--net=host` option to allow Docker to directly open ports on the host server: ```sh docker run -d \ --net=host \ -v /home/user/galaxy_storage/:/export/ \ bgruening/galaxy-stable ``` Note that there is no need to specifically bind individual ports (e.g., `-p 80:80`) if you use `--net`. An alternative to FTP and it's shortcomings it to use the SFTP protocol via port 22. Start your Galaxy container with a port binding to 22. ```sh docker run -i -t -p 8080:80 -p 8022:22 \ -v /home/user/galaxy_storage/:/export/ \ bgruening/galaxy-stable ``` And use for example [Filezilla](https://filezilla-project.org/) or the `sftp` program to transfer data: ```sh sftp -v -P 8022 -o User=admin@galaxy.org localhost <<< $'put' ``` ## Using Parent docker [[toc]](#toc) On some linux distributions, Docker-In-Docker can run into issues (such as running out of loopback interfaces). If this is an issue, you can use a 'legacy' mode that use a docker socket for the parent docker installation mounted inside the container. To engage, set the environmental variable `DOCKER_PARENT` ```sh docker run -p 8080:80 -p 8021:21 -p 8800:8800 \ --privileged=true -e DOCKER_PARENT=True \ -v /var/run/docker.sock:/var/run/docker.sock \ -v /home/user/galaxy_storage/:/export/ \ bgruening/galaxy-stable ``` ## Galaxy Report Webapp [[toc]](#toc) For admins wishing to have more information on the status of a galaxy instance, the Galaxy Report Webapp is served on `http://localhost:8080/reports`. As default this site is password protected with `admin:admin`. You can change this by providing a `reports_htpasswd` file in `/home/user/galaxy_storage/`. You can disable the Report Webapp entirely by providing the environment variable `NONUSE` during container startup. ```sh docker run -p 8080:80 \ -e "NONUSE=reports" \ bgruening/galaxy-stable ``` ## Galaxy's config settings [[toc]](#toc) Every Galaxy configuration parameter in `config/galaxy.ini` can be overwritten by passing an environment variable to the `docker run` command during startup. The name of the environment variable has to be: `GALAXY_CONFIG`+ *the_original_parameter_name_in_capital_letters* For example, you can set the Galaxy session timeout to 5 mintues by adding `-e "GALAXY_CONFIG_SESSION_DURATION=5"` to the `docker run command` *by default* the `admin_users`, `master_api_key` and the `brand` variable it set to: ``` GALAXY_CONFIG_ADMIN_USERS=admin@galaxy.org GALAXY_CONFIG_MASTER_API_KEY=HSNiugRFvgT574F43jZ7N9F3 GALAXY_CONFIG_BRAND="Galaxy Docker Build" ``` You can and should overwrite these during launching your container: ```sh docker run -p 8080:80 \ -e "GALAXY_CONFIG_ADMIN_USERS=albert@einstein.gov" \ -e "GALAXY_CONFIG_MASTER_API_KEY=83D4jaba7330aDKHkakjGa937" \ -e "GALAXY_CONFIG_BRAND='My own Galaxy flavour'" \ bgruening/galaxy-stable ``` Note that if you would like to run any of the [cleanup scripts](https://wiki.galaxyproject.org/Admin/Config/Performance/Purge%20Histories%20and%20Datasets), you will need to add the following to `/export/galaxy-central/config/galaxy.ini`: ``` database_connection = postgresql://galaxy:galaxy@localhost:5432/galaxy file_path = /export/galaxy-central/database/files ``` ## Personalize your Galaxy [[toc]](#toc) The Galaxy welcome screen can be changed by providing a `welcome.html` page in `/home/user/galaxy_storage/`. All files starting with `welcome` will be copied during startup and served as introduction page. If you want to include images or other media, name them `welcome_*` and link them relative to your `welcome.html` ([example](`https://github.com/bgruening/docker-galaxy-stable/blob/master/galaxy/welcome.html`)). ## Deactivating services [[toc]](#toc) Non-essential services can be deactivated during startup. Set the environment variable `NONUSE` to a comma separated list of services. Currently, `nodejs`, `postgres`, `proftp`, `reports`, `slurmd` and `slurmctld` are supported. ```sh docker run -d -p 8080:80 -p 9002:9002 \ -e "NONUSE=nodejs,proftp,reports,slurmd,slurmctld" \ bgruening/galaxy-stable ``` A graphical user interface, to start and stop your services, is available on port `9002` if you run your container like above. ## Restarting Galaxy [[toc]](#toc) If you want to restart Galaxy without restarting the entire Galaxy container you can use `docker exec` (docker > 1.3). ```sh docker exec supervisorctl restart galaxy: ``` In addition you start/stop every supervisord process using a webinterface on port `9002`. Start your container with: ```sh docker run -p 9002:9002 bgruening/galaxy-stable ``` ## Advanced Logging [[toc]](#toc) You can set the environment variable $GALAXY_LOGGING to FULL to access all logs from supervisor. For example start your container with: ```sh docker run -d -p 8080:80 -p 8021:21 \ -e "GALAXY_LOGGING=full" \ bgruening/galaxy-stable ``` Then, you can access the supervisord web interface on port `9002` and get access to log files. To do so, start your container with: ```sh docker run -d -p 8080:80 -p 8021:21 -p 9002:9002 \ -e "GALAXY_LOGGING=full" \ bgruening/galaxy-stable ``` Alternatively, you can access the container directly using the following command: ```sh docker exec -it bash ``` Once connected to the container, log files are available in `/home/galaxy/logs`. A volume can also be used to map this directory to one external to the container - for instance if logs need to be persisted for auditing reasons (security, debugging, performance testing, etc...).: ```sh mkdir gx_logs docker run -d -p 8080:80 -p 8021:21 -e "GALAXY_LOGGING=full" -v `pwd`/gx_logs:/home/galaxy/logs bgruening/galaxy-stable ``` ## Using an external Slurm cluster [[toc]](#toc) It is often convenient to configure Galaxy to use a high-performance cluster for running jobs. To do so, two files are required: 1. munge.key 2. slurm.conf These files from the cluster must be copied to the `/export` mount point (i.e., `/data/galaxy` on the host if using below command) accessible to Galaxy before starting the container. This must be done regardless of which Slurm daemons are running within Docker. At start, symbolic links will be created to these files to `/etc` within the container, allowing the various Slurm functions to communicate properly with your cluster. In such cases, there's no reason to run `slurmctld`, the Slurm controller daemon, from within Docker, so specify `-e "NONUSE=slurmctld"`. Unless you would like to also use Slurm (rather than the local job runner) to run jobs within the Docker container, then alternatively specify `-e "NONUSE=slurmctld,slurmd"`. Importantly, Slurm relies on a shared filesystem between the Docker container and the execution nodes. To allow things to function correctly, each of the execution nodes will need `/export` and `/galaxy-central` directories to point to the appropriate places. Suppose you ran the following command to start the Docker image: ```sh docker run -d \ -e "NONUSE=slurmd,slurmctld" \ -p 80:80 \ -v /data/galaxy:/export \ bgruening/galaxy-stable ``` You would then need the following symbolic links on each of the nodes: 1. `/export` `/data/galaxy` 2. `/galaxy-central` `/data/galaxy/galaxy-central` A brief note is in order regarding the version of Slurm installed. This Docker image uses Ubuntu 14.04 as its base image. The version of Slurm in the Ubuntu 14.04 repository is 2.6.5 and that is what is installed in this image. If your cluster is using an incompatible version of Slurm then you will likely need to modify this Docker image. The following is an example for how to specify a destination in `job_conf.xml` that uses a custom partition ("work", rather than "debug") and 4 cores rather than 1: ``` False -p work -n 4 ``` The usage of `-n` can be confusing. Note that it will specify the number of cores, not the number of tasks (i.e., it's not equivalent to `srun -n 4`). ## Using an external Grid Engine cluster [[toc]](#toc) Almost things is as same as Slurm cluster. To use Grid Engine (Sun Grid Engine, Open Grid Scheduler), one configuration file and an environment variable are required: 1. set the environment variable `SGE_ROOT` 2. create `/var/lib/gridengine/default/common/act_qmaster` file By default ``` -e SGE_ROOT=/var/lib/gridengine -v $PWD/act_qmaster:/var/lib/gridengine/default/common/act_qmaster ``` In ***act_qmaster*** is something like this. ``` YOUR_GRIDENGINE_MASTER_HOST ``` Your Grid Engine needs to accept job submissions from inside the container. If Grid Engine accepts job submission from the Docker host, the easiest way to forward all necessary ports is to use the `--net` Docker options in the following way like `--net=host` ## Tips for Running Jobs Outside the Container [[toc]](#toc) In its default state Galaxy assumes both the Galaxy source code and various temporary files are available on shared file systems across the cluster. When using Condor or SLURM (as described above) to run jobs outside of the Docker container one can take steps to mitigate these assumptions. The `embed_metadata_in_job` option on job destinations in `job_conf.xml` forces Galaxy collect metadata inside the container instead of on the cluster: ``` False ``` This has performance implications and may not scale as well as performing these calculations on the remote cluster - but this should not be a problem for most Galaxy instances. Additionally, many framework tools depend on Galaxy's Python virtual environment being available. This should be created outside of the container on a shared filesystem available to your cluster using the instructions [here](https://github.com/galaxyproject/galaxy/blob/dev/doc/source/admin/framework_dependencies.rst#managing-dependencies-manually). Job destinations can then source these virtual environments using the instructions outlined [here](https://github.com/galaxyproject/galaxy/blob/dev/doc/source/admin/framework_dependencies.rst#galaxy-job-handlers). In other words, by adding a line such as this to each job destination: `````` # Enable Galaxy to use BioContainers (Docker) [[toc]](#toc) This is a very cool feature where Galaxy automatically detects that your tool has an associated docker image, pulls it and runs it for you. These images (when available) have been generated using [mulled](https://github.com/mulled). To test, install the [IUC bedtools](https://toolshed.g2.bx.psu.edu/repository?repository_id=8d84903cc667dbe7&changeset_revision=7b3aaff0d78c) from the toolshed. When you try to execute *ClusterBed* for example. You may get a missing dependancy error for *bedtools*. But bedtools has an associated docker image on [quay.io](https://quay.io/). Now configure Galaxy as follows: - Add this environment variable to `docker run`: `-e GALAXY_CONFIG_ENABLE_BETA_MULLED_CONTAINERS=True` - In `job_conf.xml` configure a Docker enabled destination as follows: ```xml true $galaxy_root:ro,$galaxy_root/database/tmp:rw,$tool_directory:ro,$job_directory:ro,$working_directory:rw,$default_file_path:rw false ``` When you execute the tool again, Galaxy will pull the image from Biocontainers ([quay.io/biocontainers](https://quay.io/organization/biocontainers)), run the tool inside of this container to produce the desired output. # Magic Environment variables [[toc]](#toc) | Name | Description | |---|---| | `ENABLE_TTS_INSTALL` | Enables the Test Tool Shed during container startup. This change is not persistent. (`ENABLE_TTS_INSTALL=True`) | | `GALAXY_LOGGING` | Enables for verbose logging at Docker stdout. (`GALAXY_LOGGING=full`) | | `BARE` | Disables all default Galaxy tools. (`BARE=True`) | | `NONUSE` | Disable services during container startup. (`NONUSE=nodejs,proftp,reports,slurmd,slurmctld`) | | `UWSGI_PROCESSES` | Set the number of uwsgi processes (`UWSGI_PROCESSES=2) | | `UWSGI_THREADS` | Set the number of uwsgi threads (`UWSGI_THREADS=4`) | | `GALAXY_DOCKER_ENABLED` | Enable Galaxy to use Docker containers if annotated in tools (`GALAXY_DOCKER_ENABLED=False`) | | `GALAXY_DOCKER_VOLUMES` | Specify volumes that should be mounted into tool containers (`GALAXY_DOCKER_VOLUMES=""`) | | `GALAXY_HANDLER_NUMPROCS` | Set the number of Galaxy handler (`GALAXY_HANDLER_NUMPROCS=2`) | # Lite Mode [[toc]](#toc) The lite mode will only start postgresql and a single Galaxy process, without nginx, uwsgi or any other special feature from the normal mode. In particular there is no support for the export folder or any Magic Environment variables. ```sh docker run -i -t -p 8080:8080 bgruening/galaxy-stable startup_lite ``` This will also use the standard `job_conf.xml.sample_basic` shipped by Galaxy. If you want to use the regular one from the normal mode you can pass `-j` to the `startup_lite` script. # Extending the Docker Image [[toc]](#toc) If the desired tools are already included in the Tool Shed, building your own personalised Galaxy docker Image (Galaxy flavour) can be done using the following steps: 1. Create a file named `Dockerfile` 2. Include `FROM bgruening/galaxy-stable` at the top of the file. This means that you use the Galaxy Docker Image as base Image and build your own extensions on top of it. 3. Supply the list of desired tools in a file (`my_tool_list.yml` below). See [this page](https://github.com/galaxyproject/ansible-galaxy-tools/blob/master/files/tool_list.yaml.sample) for the file format requirements. 4. Execute `docker build -t my-docker-test .` 5. Run your container with `docker run -p 8080:80 my-docker-test` 6. Open your web browser on `http://localhost:8080` For a working example, have a look at the or the Dockerfile's. - [deepTools](http://deeptools.github.io/) [Dockerfile](https://github.com/bgruening/docker-recipes/blob/master/galaxy-deeptools/Dockerfile) - [ChemicalToolBox](https://github.com/bgruening/galaxytools/tree/master/chemicaltoolbox) [Dockerfile](https://github.com/bgruening/docker-recipes/blob/master/galaxy-chemicaltoolbox/Dockerfile) ``` # Galaxy - deepTools # # VERSION 0.2 FROM bgruening/galaxy-stable MAINTAINER Bjrn A. Grning, bjoern.gruening@gmail.com ENV GALAXY_CONFIG_BRAND deepTools WORKDIR /galaxy-central RUN add-tool-shed --url 'http://testtoolshed.g2.bx.psu.edu/' --name 'Test Tool Shed' # Install Visualisation RUN install-biojs msa # Adding the tool definitions to the container ADD my_tool_list.yml $GALAXY_ROOT/my_tool_list.yml # Install deepTools RUN install-tools $GALAXY_ROOT/my_tool_list.yml # Mark folders as imported from the host. VOLUME ["/export/", "/data/", "/var/lib/docker"] # Expose port 80 (webserver), 21 (FTP server), 8800 (Proxy) EXPOSE :80 EXPOSE :21 EXPOSE :8800 # Autostart script that is invoked during container start CMD ["/usr/bin/startup"] ``` If you host your flavor on GitHub consider to test our build with Travis-CI. This project will help you: https://github.com/bgruening/galaxy-flavor-testing ## List of Galaxy flavours [[toc]](#toc) - [NCBI-Blast](https://github.com/bgruening/docker-galaxy-blast) - [ChemicalToolBox](https://github.com/bgruening/docker-recipes/blob/master/galaxy-chemicaltoolbox) - [ballaxy](https://github.com/anhi/docker-scripts/tree/master/ballaxy) - [NGS-deepTools](https://github.com/bgruening/docker-recipes/blob/master/galaxy-deeptools) - [Galaxy ChIP-exo](https://github.com/gregvonkuster/docker-galaxy-ChIP-exo) - [Galaxy Proteomics](https://github.com/bgruening/docker-galaxyp) - [Imaging](https://github.com/bgruening/docker-galaxy-imaging) - [Constructive Solid Geometry](https://github.com/gregvonkuster/docker-galaxy-csg) - [Galaxy for metagenomics](https://github.com/bgruening/galaxy-metagenomics) - [Galaxy with the Language Application Grid tools](https://github.com/lappsgrid-incubator/docker-galaxy-lappsgrid) - [RNAcommender](https://github.com/gianlucacorrado/galaxy-RNAcommender) - [OpenMoleculeGenerator](https://github.com/bgruening/galaxy-open-molecule-generator) - [Workflow4Metabolomics](https://github.com/workflow4metabolomics/w4m-docker) - [HiC-Explorer](https://github.com/maxplanck-ie/docker-galaxy-hicexplorer) - [SNVPhyl](https://github.com/phac-nml/snvphyl-galaxy) - [GraphClust](https://github.com/BackofenLab/docker-galaxy-graphclust) - [RNA workbench](https://github.com/bgruening/galaxy-rna-workbench) - [Cancer Genomics Toolkit](https://github.com/morinlab/tools-morinlab/tree/master/docker) # Integrating non-Tool Shed tools into the container [[toc]](#toc) We recommend to use the [Main Galaxy Tool Shed](https://toolshed.g2.bx.psu.edu/) for all your tools and workflows that you would like to share. In rare situations where you cannot share your tools but still want to include them into your Galaxy Docker instance, please follow the next steps. - Get your tools into the container. Mount your tool directory into the container with a separate `-v /home/user/my_galaxy_tools/:/local_tools`. - Create a `tool_conf.xml` file for your tools. This should look similar to the main [`tool_conf.xml`](https://github.com/galaxyproject/galaxy/blob/dev/config/tool_conf.xml.sample) file, but references your tools from the new directory. In other words a tool entry should look like this ``. Your `tool_conf.xml` should be available from inside of the container. We assume you have it stored under `/local_tools/my_tools.xml`. - Add the new tool config file to the Galaxy configuration. To make Galaxy aware of your new tool configuration file you need to add the path to `tool_config_file`, which is by default `#tool_config_file = config/tool_conf.xml,config/shed_tool_conf.xml`. You can do this during container start by setting the environment variable `-e GALAXY_CONFIG_TOOL_CONFIG_FILE=config/tool_conf.xml.sample,config/shed_tool_conf.xml.sample,/local_tools/my_tools.xml`. # Users & Passwords [[toc]](#toc) The Galaxy Admin User has the username `admin@galaxy.org` and the password `admin`. The PostgreSQL username is `galaxy`, the password is `galaxy` and the database name is `galaxy` (I know I was really creative ;)). If you want to create new users, please make sure to use the `/export/` volume. Otherwise your user will be removed after your docker session is finished. The proftpd server is configured to use the main galaxy PostgreSQL user to access the database and select the username and password. If you want to run the docker container in production, please do not forget to change the user credentials in `/etc/proftpd/proftpd.conf` too. The Galaxy Report Webapp is `htpasswd` protected with username and password st to `admin`. # Development [[toc]](#toc) This repository uses a git submodule to include [Ansible roles](https://github.com/galaxyproject/ansible-galaxy-extras) maintained by the Galaxy project. You can clone this repository and the Ansible submodule with: ```sh git clone --recursive https://github.com/bgruening/docker-galaxy-stable.git ``` Updating already existing submodules is possible with: ```sh git submodule update --init --recursive ``` If you simply want to change the Galaxy repository and/or the Galaxy branch, from which the container is build you can do this with Docker `--build-arg` during the `docker build` step. For example you can use these parameters during container build: ``` --build-arg GALAXY_RELEASE=install_workflow_and_tools --build-arg GALAXY_REPO=https://github.com/manabuishii/galaxy ``` # Requirements [[toc]](#toc) - [Docker](https://www.docker.io/gettingstarted/#h_installation) # History [[toc]](#toc) - 0.1: Initial release! - with Apache2, PostgreSQL and Tool Shed integration - 0.2: complete new Galaxy stack. - with nginx, uwsgi, proftpd, docker, supervisord and SLURM - 0.3: Add Interactive Environments - IPython in docker in Galaxy in docker - advanged logging - 0.4: - base the image on toolshed/requirements with all required Galaxy dependencies - use Ansible roles to build large parts of the image - export the supervisord webinterface on port 9002 - enable Galaxy reports webapp - 15.07: - `install-biojs` can install BioJS visualisations into Galaxy - `add-tool-shed` can be used to activate third party Tool Sheds in child Dockerfiles - many documentation improvements - RStudio is now part of Galaxy and this Image - configurable postgres UID/GID by @chambm - smarter starting of postgres during Tool installations by @shiltemann - 15.10: - new Galaxy 15.10 release - fix https://github.com/bgruening/docker-galaxy-stable/issues/94 - 16.01: - enable Travis testing for all builds and PR - offer new [yaml based tool installations](https://github.com/galaxyproject/ansible-galaxy-tools/blob/master/files/tool_list.yaml.sample) - enable dynamic UWSGI processes and threads with `-e UWSGI_PROCESSES=2` and `-e UWSGI_THREADS=4` - enable dynamic Galaxy handlers `-e GALAXY_HANDLER_NUMPROCS=2` - Addition of a new `lite` mode contributed by @kellrott - first release with Jupyter integration - 16.04: - include a Galaxy-bare mode, enable with `-e BARE=True` - first release with [HTCondor](https://research.cs.wisc.edu/htcondor/) installed and pre-configured - 16.07: - documentation and tests updates for SLURM integration by @mvdbeek - first version with initial Docker compose support (proftpd ) - SFTP support by @zfrenchee - 16.10: - [HTTPS support](https://github.com/bgruening/docker-galaxy-stable/pull/240 ) by @zfrenchee and @mvdbeek - 17.01: - enable Conda dependency resultion by deault - [new Galaxy version](https://docs.galaxyproject.org/en/master/releases/17.01_announce.html) - more compose work (slurm, postgresql) # Support & Bug Reports [[toc]](#toc) You can file an [github issue](https://github.com/bgruening/docker-galaxy-stable/issues) or ask us on the [Galaxy development list](http://lists.bx.psu.edu/listinfo/galaxy-dev).
Owner
- Name: Bérénice Batut
- Login: bebatut
- Kind: user
- Location: Clermont-Ferrand, France
- Company: University of Freiburg
- Website: http://research.bebatut.fr/
- Twitter: bebatut
- Repositories: 86
- Profile: https://github.com/bebatut
@galaxyproject training, @usegalaxy-eu, @open-life-science, @StreetScienceCommunity, @gallantries