https://github.com/cpanse/tartare
raw file collection recorded on Thermo Fisher Scientific mass spectrometers for extented unit testing
Science Score: 26.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
○CITATION.cff file
-
✓codemeta.json file
Found codemeta.json file -
○.zenodo.json file
-
✓DOI references
Found 3 DOI reference(s) in README -
○Academic publication links
-
○Academic email domains
-
○Institutional organization owner
-
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (7.6%) to scientific vocabulary
Keywords
Repository
raw file collection recorded on Thermo Fisher Scientific mass spectrometers for extented unit testing
Basic Info
- Host: GitHub
- Owner: cpanse
- Language: R
- Default Branch: master
- Homepage: https://bioconductor.org/packages/devel/data/experiment/html/tartare.html
- Size: 63.5 KB
Statistics
- Stars: 1
- Watchers: 2
- Forks: 0
- Open Issues: 0
- Releases: 0
Topics
Metadata Files
README.md
tartare
raw file collection recorded on Thermo Fisher Scientific mass spectrometers for extented unit testing
https://github.com/Bioconductor/Contributions/issues/1286
install
```
Release (3.10)
if (!requireNamespace("BiocManager", quietly = TRUE)) install.packages("BiocManager")
BiocManager::install("tartare", version="devel") ```
see also https://bioconductor.org/packages/tartare/
requires
the following usage example requires the following packages:
- https://github.com/rformassspectrometry/Spectra
- https://github.com/cpanse/MsBackendRawFileReader
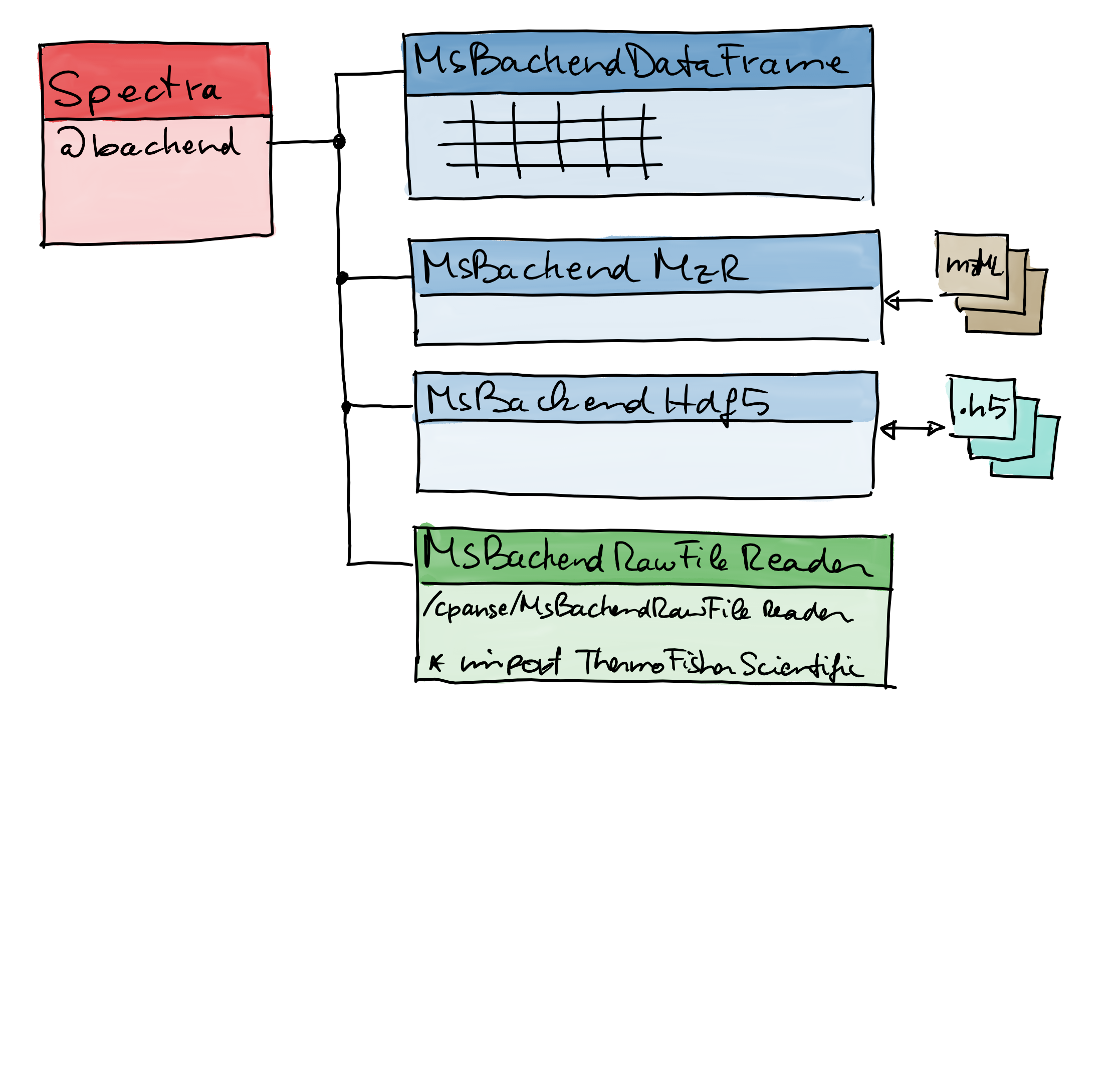 Courtesy of @jorainer; SWEMSA 2019 conference in Erding, Germany 10.5281/zenodo.3499649.
Courtesy of @jorainer; SWEMSA 2019 conference in Erding, Germany 10.5281/zenodo.3499649.
usage
taken from the help pages ?tartare.
``` library(tartare) library(ExperimentHub)
eh <- ExperimentHub() (files <- getFilename(eh))
library(MsBackendRawFileReader)
be <- lapply(files, function(f){ if (grepl("mzXML$", f)) backendInitialize(MsBackendMzR(), files = f) else backendInitialize(MsBackendRawFileReader(), files = f, extra=FALSE) })
be ```
```
R> be
$EH3219
MsBackendMzR with 1764 spectra
msLevel rtime scanIndex
file(s): 203c4d41aa69_3235
$EH3220
filename: /Users/cp/Library/Caches/ExperimentHub/203c576b7cbf_3236.raw creation date: 7/11/2019 11:04:01 AM first scan: 1 last scan: 1877 model: Q Exactive HF-X Orbitrap name: Q Exactive HF-X Orbitrap SerialNumber: Exactive Series slot #6114
$EH3221
MsBackendMzR with 8742 spectra
msLevel rtime scanIndex
file(s): 203c51cb0c6f_3237
$EH3222
filename: /Users/cp/Library/Caches/ExperimentHub/24ab678291f6_3238.raw creation date: 7/16/2019 5:56:24 PM first scan: 1 last scan: 8742 model: Orbitrap Fusion Lumos name: Orbitrap Fusion Lumos SerialNumber: FSN20583
R> ```
hfx.filename <- getFilename(eh, c('tartar', '20190710_003_PierceHeLaProteinDigestStd.raw'));
x <- .cnew ("Rawfile", hfx.filename);
x$GetInfoValues()
see ?tartare and browseVignettes('tartare') for documentation
downloading 0 resources
loading from cache
[1] "/Users/cp/Library/Caches/ExperimentHub/203c576b7cbf_3236.raw"
[2] "7/11/2019 11:04:01 AM"
[3] "1"
[4] "1877"
[5] "Q Exactive HF-X Orbitrap"
[6] "Q Exactive HF-X Orbitrap"
[7] "Exactive Series slot #6114"
Owner
- Name: Christian Panse
- Login: cpanse
- Kind: user
- Location: 47N 008E
- Company: Swiss federal institute of technology in Zurich
- Website: https://fgcz.ch/compms
- Repositories: 72
- Profile: https://github.com/cpanse
proteome informatics; ms comp; visualization; maps; code; reproducible research @hb9feb@fosstodon.org
GitHub Events
Total
Last Year
Issues and Pull Requests
Last synced: about 1 year ago
All Time
- Total issues: 2
- Total pull requests: 0
- Average time to close issues: 22 days
- Average time to close pull requests: N/A
- Total issue authors: 1
- Total pull request authors: 0
- Average comments per issue: 1.5
- Average comments per pull request: 0
- Merged pull requests: 0
- Bot issues: 0
- Bot pull requests: 0
Past Year
- Issues: 0
- Pull requests: 0
- Average time to close issues: N/A
- Average time to close pull requests: N/A
- Issue authors: 0
- Pull request authors: 0
- Average comments per issue: 0
- Average comments per pull request: 0
- Merged pull requests: 0
- Bot issues: 0
- Bot pull requests: 0
Top Authors
Issue Authors
- cpanse (2)
Pull Request Authors
Top Labels
Issue Labels
Pull Request Labels
Dependencies
- AnnotationHub >= 2.16 depends
- ExperimentHub >= 1.0 depends
- R >= 3.6 depends
- utils * imports
- BiocStyle * suggests
- knitr * suggests
- testthat * suggests
- tools * suggests