biobb_wf_pmx_tutorial
This tutorial aims to illustrate how to compute a fast-growth mutation free energy calculation, step by step, using the BioExcel Building Blocks library (biobb).
Science Score: 44.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
✓CITATION.cff file
Found CITATION.cff file -
✓codemeta.json file
Found codemeta.json file -
✓.zenodo.json file
Found .zenodo.json file -
○DOI references
-
○Academic publication links
-
○Academic email domains
-
○Institutional organization owner
-
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (3.4%) to scientific vocabulary
Repository
This tutorial aims to illustrate how to compute a fast-growth mutation free energy calculation, step by step, using the BioExcel Building Blocks library (biobb).
Basic Info
- Host: GitHub
- Owner: bioexcel
- License: apache-2.0
- Language: HTML
- Default Branch: main
- Homepage: https://mmb.irbbarcelona.org/biobb/
- Size: 53.1 MB
Statistics
- Stars: 3
- Watchers: 5
- Forks: 1
- Open Issues: 0
- Releases: 0
Metadata Files
README.md
Mutation Free Energy Calculations using BioExcel Building Blocks (biobb)
Based on the official pmx tutorial.
This tutorial aims to illustrate how to compute a fast-growth mutation free energy calculation, step by step, using the BioExcel Building Blocks library (biobb). The particular example used is the Staphylococcal nuclease protein (PDB code 1STN), a small, minimal protein, appropriate for a short tutorial.
The non-equilibrium free energy calculation protocol performs a fast alchemical transition in the direction WT->Mut and back Mut->WT. The two equilibrium trajectories needed for the tutorial, one for Wild Type (WT) and another for the Mutated (Mut) protein (Isoleucine 10 to Alanine -I10A-), have already been generated and are included in this example. We will name WT as stateA and Mut as stateB.
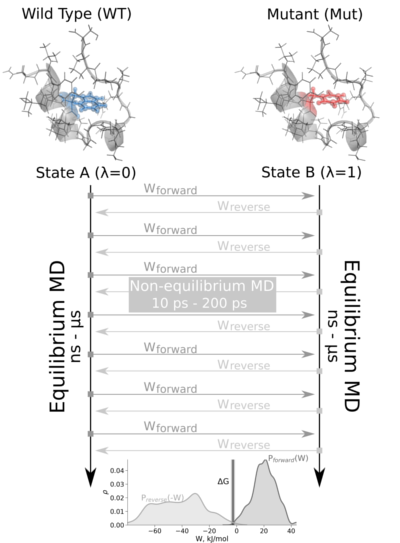
The tutorial calculates the free energy difference in the folded state of a protein. Starting from two 1ns-length independent equilibrium simulations (WT and mutant), snapshots are selected to start fast (50ps) transitions driving the system in the forward (WT to mutant) and reverse (mutant to WT) directions, and the work values required to perform these transitions are collected. With these values, Crooks Gaussian Intersection (CGI), Bennett Acceptance Ratio (BAR) and Jarzynski estimator methods are used to calculate the free energy difference between the two states.
Please note that for the sake of disk space this tutorial is using 1ns-length equilibrium trajectories, whereas in the original example the equilibrium trajectories used were obtained from 10ns-length simulations.
Settings
Biobb modules used
- biobb_pmx: Tools to setup and run Alchemical Free Energy calculations.
- biobb_gromacs: Tools to setup and run Molecular Dynamics simulations.
- biobb_analysis: Tools to analyse Molecular Dynamics trajectories.
Auxiliary libraries used
- jupyter: Free software, open standards, and web services for interactive computing across all programming languages.
- plotly: Python interactive graphing library integrated in Jupyter notebooks.
Conda Installation and Launch
console
git clone https://github.com/bioexcel/biobb_wf_pmx_tutorial.git
cd biobb_wf_pmx_tutorial
conda env create -f conda_env/environment.yml
conda activate biobb_wf_pmx_tutorial
jupyter-notebook biobb_wf_pmx_tutorial/notebooks/biobb_wf_pmx_tutorial.ipynb
Tutorial
Click here to view tutorial in Read the Docs
Click here to execute tutorial in Binder
Version
2024.1
Copyright & Licensing
This software has been developed in the MMB group at the BSC & IRB for the European BioExcel, funded by the European Commission (EU Horizon Europe 101093290, EU H2020 823830, EU H2020 675728).
- (c) 2015-2025 Barcelona Supercomputing Center
- (c) 2015-2025 Institute for Research in Biomedicine
Licensed under the Apache License 2.0, see the file LICENSE for details.

Owner
- Name: BioExcel
- Login: bioexcel
- Kind: organization
- Website: https://bioexcel.eu/
- Repositories: 50
- Profile: https://github.com/bioexcel
Center of Excellence for Computational Biomolecular Research
Citation (CITATION.cff)
abstract: "BioExcel Building Blocks (BioBB) library. BioBB's are built as Python wrappers to provide an interoperable architecture. BioBB's have been integrated in a chain of usual software management tools to generate data ontologies, documentation, installation packages, software containers and ways of integration with workflow managers, that make them usable in most computational environments."
authors:
- affiliation: "Barcelona Supercomputing Center (BSC)"
family-names: "Andrio"
given-names: "Pau"
orcid: "https://orcid.org/0000-0003-2116-3880"
- affiliation: "Institute for Research in Biomedicine (IRB Barcelona)"
family-names: "Hospital"
given-names: "Adam"
orcid: "https://orcid.org/0000-0002-8291-8071"
- affiliation: "Institute for Research in Biomedicine (IRB Barcelona)"
family-names: "Bayarri"
given-names: "Genís"
orcid: "https://orcid.org/0000-0003-0513-0288"
- affiliation: "Institute for Research in Biomedicine (IRB Barcelona)"
family-names: "García"
given-names: "Agustín"
orcid: "https://orcid.org/0009-0002-2159-965X"
- affiliation: "Institute for Research in Biomedicine (IRB Barcelona)"
family-names: "Chaves"
given-names: "Rubén"
- affiliation: "Institute for Research in Biomedicine (IRB Barcelona), University of Barcelona (UB)"
family-names: "Orozco"
given-names: "Modesto"
orcid: "https://orcid.org/0000-0002-8608-3278"
- affiliation: "Barcelona Supercomputing Center (BSC), University of Barcelona (UB)"
family-names: "Gelpí"
given-names: "Josep Ll."
orcid: "https://orcid.org/0000-0002-0566-7723"
email: "gelpi@ub.edu"
cff-version: 1.2.0
date-released: "2019-09-10"
keywords:
- "BioExcel"
- "BioBB"
- "Bioinformatics"
- "Computational Biology"
- "Biomolecular Workflows"
license: "Apache-2.0"
message: "If you use this dataset, please cite it using the metadata from this file."
repository-code: "https://github.com/bioexcel/biobb"
title: "BioExcel Building Blocks, a software library for interoperable biomolecular simulation workflows"
doi: "10.1038/s41597-019-0177-4"
url: "https://mmb.irbbarcelona.org/biobb/"
version: 5.0.0
preferred-citation:
type: "article"
authors:
- affiliation: "Barcelona Supercomputing Center (BSC)"
family-names: "Andrio"
given-names: "Pau"
orcid: "https://orcid.org/0000-0003-2116-3880"
- affiliation: "Institute for Research in Biomedicine (IRB Barcelona)"
family-names: "Hospital"
given-names: "Adam"
orcid: "https://orcid.org/0000-0002-8291-8071"
- affiliation: "Barcelona Supercomputing Center (BSC)"
family-names: "Conejero"
given-names: "Javier"
orcid: "https://orcid.org/0000-0001-6401-6229"
- affiliation: "Barcelona Supercomputing Center (BSC)"
family-names: "Jordà"
given-names: "Luis"
orcid: "https://orcid.org/0000-0002-9407-9703"
- affiliation: "Barcelona Supercomputing Center (BSC)"
family-names: "Del Pino"
given-names: "Marc"
orcid: "https://orcid.org/0000-0001-5565-7577"
- affiliation: "Barcelona Supercomputing Center (BSC)"
family-names: "Laia"
given-names: "Codó"
orcid: "https://orcid.org/0000-0002-6797-8746"
- affiliation: "University of Manchester (UOM)"
family-names: "Soiland-Reyes"
given-names: "Stian"
orcid: "https://orcid.org/0000-0001-9842-9718"
- affiliation: "University of Manchester (UOM)"
family-names: "Goble"
given-names: "Carole"
orcid: "https://orcid.org/0000-0003-1219-2137"
- affiliation: "Barcelona Supercomputing Center (BSC)"
family-names: "Lezzi"
given-names: "Daniele"
orcid: "https://orcid.org/0000-0001-5081-7244"
- affiliation: "Barcelona Supercomputing Center (BSC)"
family-names: "Badia"
given-names: "Rosa M"
orcid: "https://orcid.org/0000-0003-2941-5499"
- affiliation: "Institute for Research in Biomedicine (IRB Barcelona), University of Barcelona (UB)"
family-names: "Orozco"
given-names: "Modesto"
orcid: "https://orcid.org/0000-0002-8608-3278"
- affiliation: "Barcelona Supercomputing Center (BSC), University of Barcelona (UB)"
family-names: "Gelpí"
given-names: "Josep Ll."
orcid: "https://orcid.org/0000-0002-0566-7723"
email: "gelpi@ub.edu"
doi: "10.1038/s41597-019-0177-4"
journal: "Nature Scientific Data"
month: 9
start: 169
title: "BioExcel Building Blocks, a software library for interoperable biomolecular simulation workflows"
issue: 1
volume: 6
year: 2019
GitHub Events
Total
- Watch event: 2
- Push event: 7
Last Year
- Watch event: 2
- Push event: 7
Dependencies
- actions/checkout v2 composite
- actions/stale v5 composite
- recommonmark *
- sphinx_rtd_theme *
