Science Score: 54.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
✓CITATION.cff file
Found CITATION.cff file -
✓codemeta.json file
Found codemeta.json file -
✓.zenodo.json file
Found .zenodo.json file -
○DOI references
-
✓Academic publication links
Links to: arxiv.org -
○Academic email domains
-
○Institutional organization owner
-
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (14.5%) to scientific vocabulary
Repository
Tool for Annotating microscopy
Basic Info
- Host: GitHub
- Owner: luckieucas
- License: other
- Language: Python
- Default Branch: main
- Size: 13.8 MB
Statistics
- Stars: 1
- Watchers: 1
- Forks: 0
- Open Issues: 0
- Releases: 0
Metadata Files
README.md

Cellable
Cell Organelle Labeling with Python

Overview
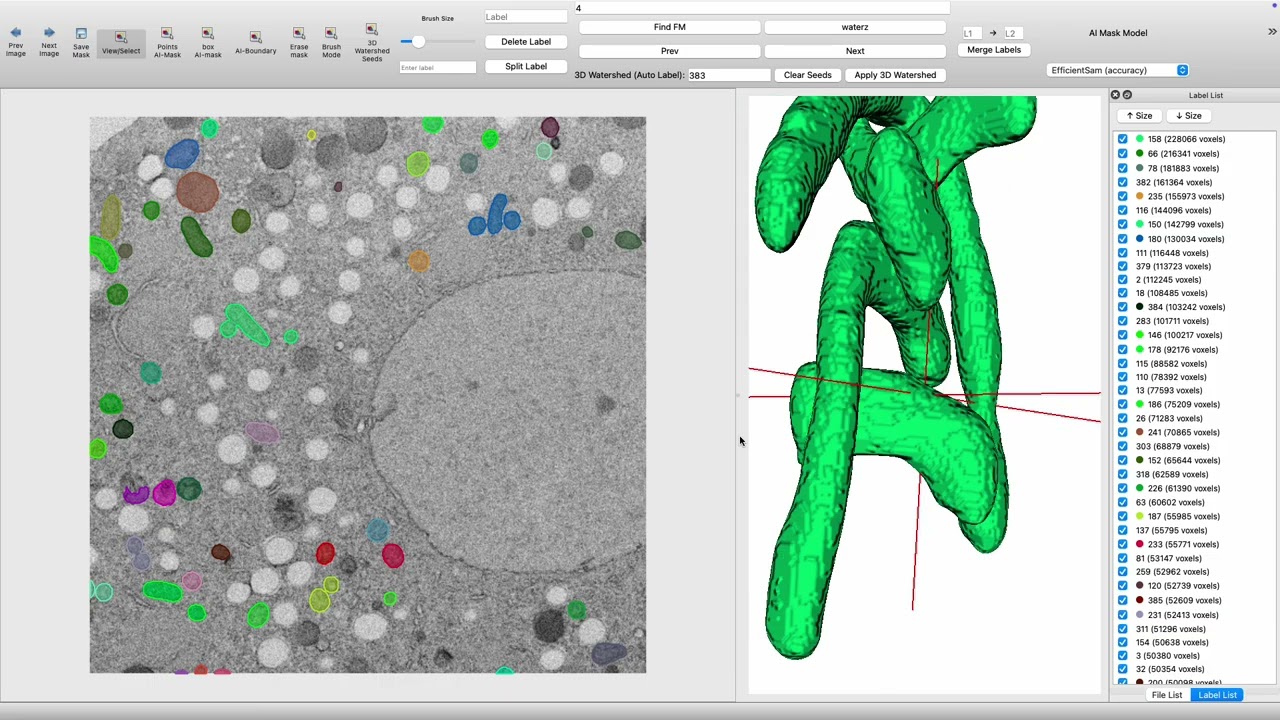
This application is an extended version of Labelme, designed for interactive 2D/3D segmentation and annotation of electron microscopy (EM) and other scientific images. It supports:
- Viewing and annotating 2D slices and 3D volumes
- Loading TIFF stacks for volumetric data
- Automatic AI-assisted segmentation
- Manual mask editing and refinement
- 3D rendering via VTK
Installation
1. Requirements
- Python 3.8+
- GPU recommended for AI-assisted segmentation
- OS: Linux, macOS, or Windows
2. Install Dependencies
Key dependencies include:
PyQt5– GUI frameworkvtk– 3D renderingtifffile– TIFF image I/Occ3d– connected component analysisscikit-image,scipy,numpy– image processingimgviz– visualization utilities
```bash git clone https://github.com/luckieucas/cellable.git cd cellable
Setup conda
conda create --name cellable python=3.9 conda activate cellable
Install dependencies
pip install -r requirements.txt
Install cellable
pip install -e . ```
📚 User Tutorial - Cellable 3D Segmentation Edition
🚀 Getting Started
Launch the Application
bash
conda activate cellable
cellable
🖥️ Interface Overview
Main Window Layout
- Toolbar: File operations, AI segmentation, view adjustments
- Canvas Area: Displays current image or 3D slice
- Label List: Shows all current annotations
- Status Bar: Displays slice index, zoom level, current tool
📁 Data Loading & Supported Formats
Supported File Formats
- Images:
.png,.jpg,.tif,.tiff - Volume Data: Multi-page TIFF stacks
Loading Data Steps
- Open Image/Stack:
File → Open - For 3D TIFF stacks, a slider will appear for slice navigation
️ View Navigation & Operations
Basic Operations
- Mouse Scroll: Change zoom level
- Arrow Keys/Slider: Move between slices
- Drag: Pan the view
✏️ Annotation Tools
1. Polygon Tool - Manual Contour Drawing
- Click on canvas to create vertices
- Double-click to complete drawing
- Right-click to edit vertices
2. Mask Tool - Region Painting
- Select brush size
- Paint mask regions
- Use eraser to remove areas
🤖 AI-Assisted Segmentation
SAM (Segment Anything Model) Segmentation
- Select the AI tool
- Click inside the region of interest
- Automatic segmentation generation
Efficient SAM - Fast Segmentation
- Faster segmentation speed
- Suitable for batch processing
Text-to-Annotation Conversion
- Input descriptive text
- Automatic annotation generation
🔧 Mask Editing & Optimization
Shape Editing
- Move, resize, or delete shapes
- Merge or split regions
- Adjust brightness/contrast
🌊 Watershed Segmentation - Instance Separation
Find False Merge Feature
- Enter the target label ID in the Label ID input field on the right
- Navigate to a slice containing adhered instances
- Click the waterz button
- Automatic boundary computation and view refresh

🎨 3D Rendering & Visualization
VTK 3D Viewer
- View → 3D Viewer
- VTK-based 3D visualization of masks
- Rotate, zoom, and inspect segmented structures
💾 Save & Export
Saving Annotations
File → Savestores as.jsonformat- Mask data can be exported as NumPy arrays
Export Formats
- JSON annotation files
- VOC dataset format
- COCO dataset format
⌨️ Keyboard Shortcuts
| Action | Shortcut |
|--------|----------|
| Open File | Ctrl+O |
| Save Annotation | Ctrl+S |
| Zoom | Hold Cmd + Mouse Scroll |
| Next Slice | D |
| Previous Slice | A |
| Undo | Ctrl+Z |
| Redo | Ctrl+Y |
🚀 Advanced Features
Batch Processing
- Multiple file annotation
- Automatic progress saving
Annotation Quality Control
- Overlap detection
- Completeness checking
- Statistical reports
🎯 Features
Core Features
- ✅ 2D/3D Image Annotation
- ✅ AI-Assisted Segmentation (SAM, Efficient SAM)
- ✅ Text-to-Annotation Conversion
- ✅ Watershed Instance Separation
- ✅ 3D VTK Visualization
- ✅ Multi-format Export Support
Professional Features
- 📊 Volume Data Analysis
- ✏️ Precise Mask Editing
- 📊 Batch Processing Support
❓ Troubleshooting & FAQ
Performance Issues
- Laggy performance: Enable GPU acceleration and close unused windows
- Memory issues: Reduce the number of simultaneously open files
Technical Issues
- Mask misalignment: Check voxel dimensions in TIFF metadata
- VTK viewer not loading: Ensure
vtkandPyQt5versions are compatible
AI Segmentation Issues
- Inaccurate segmentation: Adjust click position, use manual editing for optimization
- Model loading failure: Check network connection and model file integrity
🔗 Advanced Tutorials
Custom Annotation Workflows
- Create annotation templates
- Set annotation rules
- Quality check procedures
Data Preprocessing
- Image enhancement
- Format conversion
- Batch renaming
🤝 Community & Support
- GitHub Issues: Report bugs and feature requests
- Discussions: Share experiences and best practices
- Contributing Guide: Participate in project development
Credits
This version builds upon the original Labelme and integrates:
- VTK for 3D visualization
- cc3d for connected component analysis
- AI models for auto-segmentation
- Efficient SAM for fast segmentation
- Text-to-annotation capabilities
📖 Additional Resources
- Original Labelme Project
- SAM Model Paper
- VTK Documentation
- Electron Microscopy Image Processing Best Practices
🎉 Start using Cellable for professional cell organelle annotation!
For questions, check the tutorial videos or submit a GitHub Issue
🔍 问题分析
GitHub README的限制:
- ❌ 不支持HTML <video> 标签的视频播放
- ❌ 不支持嵌入式YouTube播放器
- ❌ 不支持JavaScript交互
- ✅ 只支持静态图片和链接
** 解决方案**
方案1: 使用YouTube缩略图 + 播放按钮图标 (推荐)
```markdown:README.md
Owner
- Name: luckiepeng
- Login: luckieucas
- Kind: user
- Repositories: 7
- Profile: https://github.com/luckieucas
Citation (CITATION.cff)
cff-version: 1.2.0 message: "If you use this software, please cite it as below." authors: - family-names: "Wada" given-names: "Kentaro" orcid: "https://orcid.org/0000-0002-6347-5156" title: "Labelme: Image Polygonal Annotation with Python" doi: 10.5281/zenodo.5711226 url: "https://github.com/wkentaro/labelme" license: GPL-3
GitHub Events
Total
- Watch event: 1
- Push event: 14
- Public event: 1
Last Year
- Watch event: 1
- Push event: 14
- Public event: 1
Dependencies
- actions/checkout v2 composite
- conda-incubator/setup-miniconda v3 composite
- actions/checkout v2 composite
- actions/download-artifact v1 composite
- actions/upload-artifact v1 composite
- actions/upload-release-asset v1 composite
- conda-incubator/setup-miniconda v3 composite
- mikepenz/action-gh-release v0.2.0-a03 composite
- mikepenz/release-changelog-builder-action v3 composite
- Jinja2 ==3.1.5
- MarkupSafe ==3.0.2
- PyQt5 ==5.15.11
- PyQt5-Qt5 ==5.15.16
- PyQt5_sip ==12.16.1
- PySocks ==1.7.1
- PyYAML ==6.0.2
- QtPy ==2.4.2
- annotated-types ==0.7.0
- beautifulsoup4 ==4.12.3
- cellpose ==3.1.0
- certifi ==2024.12.14
- charset-normalizer ==3.4.1
- click ==8.1.8
- coloredlogs ==15.0.1
- connected-components-3d ==3.22.0
- contourpy ==1.3.0
- cycler ==0.12.1
- fastremap ==1.15.0
- filelock ==3.17.0
- flatbuffers ==25.1.21
- fonttools ==4.55.4
- fsspec ==2024.12.0
- gdown ==5.2.0
- h5py ==3.12.1
- humanfriendly ==10.0
- idna ==3.10
- imagecodecs ==2024.12.30
- imageio ==2.37.0
- imgviz ==1.7.6
- importlib_resources ==6.5.2
- joblib ==1.4.2
- kiwisolver ==1.4.7
- lazy_loader ==0.4
- llvmlite ==0.43.0
- loguru ==0.7.3
- matplotlib ==3.9.4
- mpmath ==1.3.0
- natsort ==8.4.0
- networkx ==3.2.1
- numba ==0.60.0
- numpy ==2.0.2
- onnxruntime ==1.19.2
- opencv-python-headless ==4.11.0.86
- osam ==0.2.2
- packaging ==24.2
- pillow ==11.1.0
- protobuf ==5.29.3
- pydantic ==2.10.5
- pydantic_core ==2.27.2
- pyparsing ==3.2.1
- python-dateutil ==2.9.0.post0
- requests ==2.32.3
- roifile ==2024.9.15
- scikit-image ==0.24.0
- scikit-learn ==1.6.1
- scipy ==1.13.1
- six ==1.17.0
- soupsieve ==2.6
- sympy ==1.13.1
- termcolor ==2.5.0
- threadpoolctl ==3.5.0
- tifffile ==2024.8.30
- torch ==2.5.1
- tqdm ==4.67.1
- typing_extensions ==4.12.2
- urllib3 ==2.3.0
- vtk ==9.4.1
- zipp ==3.21.0
- Pillow >=2.8
- PyYAML *
- gdown *
- imgviz >=1.7.5
- matplotlib *
- natsort >=7.1.0
- numpy *
- onnxruntime >=1.14.1,
- osam >=0.2.2
- qtpy *
- scikit-image *
- termcolor *