faster-python-using-julia-blogposts
Experiments calling Julia from Python
https://github.com/abelsiqueira/faster-python-using-julia-blogposts
Science Score: 67.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
✓CITATION.cff file
Found CITATION.cff file -
✓codemeta.json file
Found codemeta.json file -
✓.zenodo.json file
Found .zenodo.json file -
✓DOI references
Found 1 DOI reference(s) in README -
✓Academic publication links
Links to: zenodo.org -
○Academic email domains
-
○Institutional organization owner
-
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (11.2%) to scientific vocabulary
Repository
Experiments calling Julia from Python
Basic Info
Statistics
- Stars: 11
- Watchers: 3
- Forks: 6
- Open Issues: 6
- Releases: 5
Metadata Files
README.md
Calling Julia from Python - blog post material
This material is part of a series of blog posts about using Julia from Python (Soon). The idea was initially presented internally at the Netherlands eScience Center(See the slides).
Post links: - https://blog.esciencecenter.nl/how-to-call-julia-code-from-python-8589a56a98f2 - https://blog.esciencecenter.nl/speed-up-your-python-code-using-julia-f97a6c155630 - Soon
Summary
- We read Patrick's blog post about improving the reading of irregular files.
- Patrick has a Python (Pandas) code that is slow.
- Using some packages, he moves the reading and parsing to C++.
- We decided to try to replace C++ with Julia to check:
- How easy/hard it is
- How much improvement can be gained with a basic Julia code;
- How much further improvement can be gained with an optimized Julia code.
The strategies we examined are below, with a plot with the comparison following it:
- Python with Pandas, as seen in Patrick's post. label: "Pure Python".
- Python with reading and parsing in C++, as seen in Patrick's post. label: "C++".
- Python with reading and parsing in Julia, in 4 different versions:
- Basic Julia version with mostly disregard for efficiency, label="Basic Julia".
- Julia version trying to improve memory usage. label: "Prealloc Julia".
- Julia version where the elements are read with
fscanffrom C. label: "Julia + C parsing". - Julia version reading the file as bytes and manually walking through the bytes. label: "Optimized Julia".
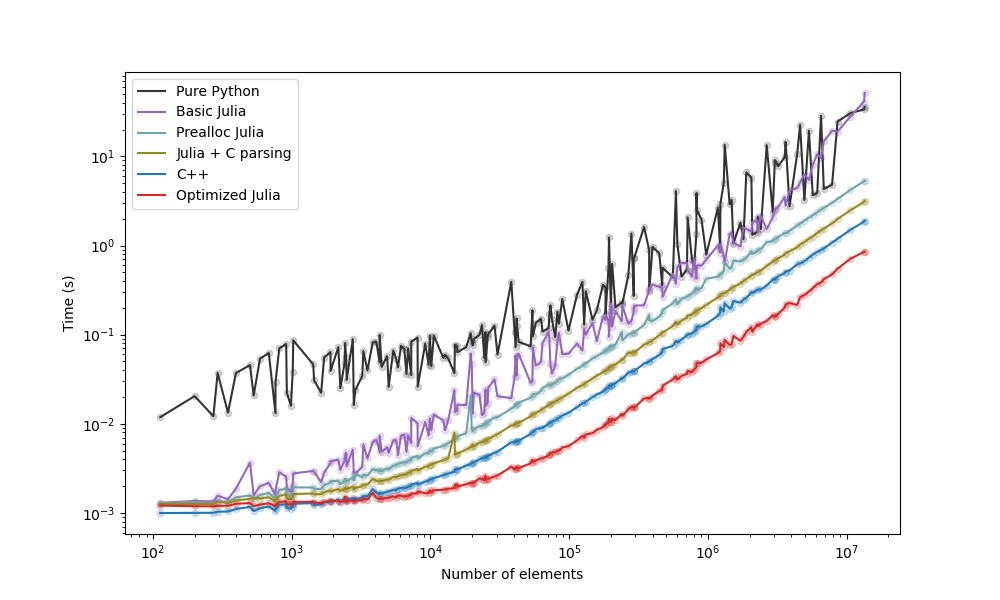
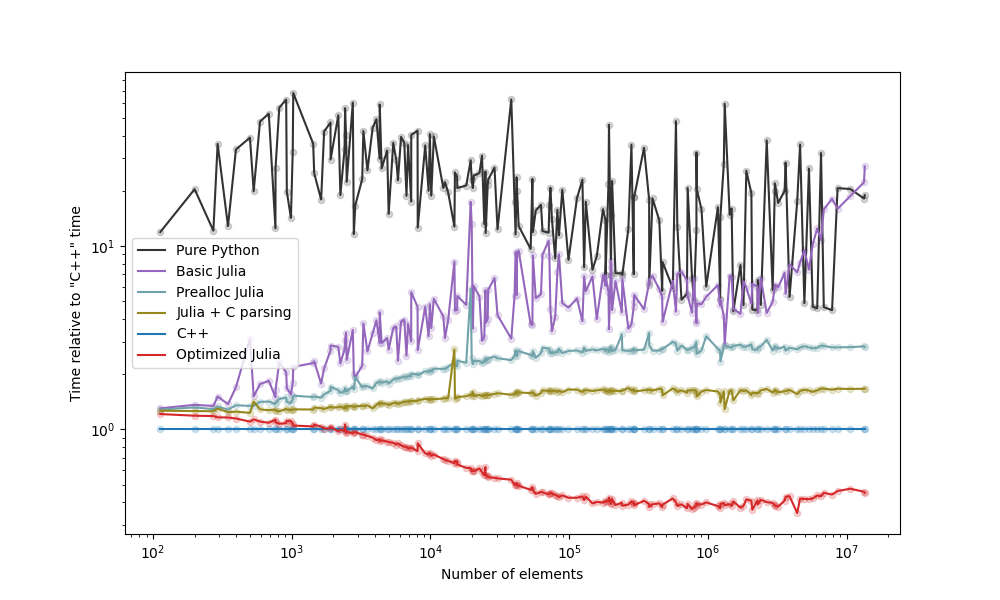
Take-aways (see blog post): - The "Prealloc Julia" strategy is already an improvement over the "Pure Python" strategy. - The "Optimized Julia" strategy is faster than the "C++" strategy. - If you don't know Julia nor C++, moving the slow code to Julia yields benefits faster and with less effort.
The image below shows the speedup gain over the effort to get there:
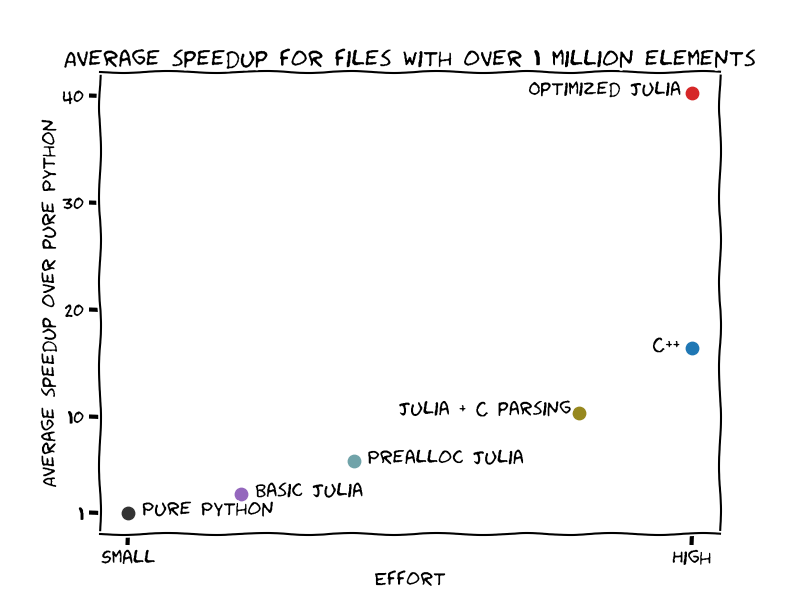
Building the docker images
shell
docker build --tag jl-from-py:<VERSION>
Reproducting the results
- Download dataset and store in a folder called
dataset. - Get the image with
shell docker pull abelsiqueira/faster-python-with-julia-blogpost:post3 - Run it with
shell
docker run --rm --volume "$PWD/dataset:/app/dataset" --volume "$PWD/out:/app/out" abelsiqueira/faster-python-with-julia-blogpost:post3
- You will find the outputs in the out/ folder.
The execution of this script with default options took about 45 minutes on a Dell Precision 5530 with the Intel chip i7-8850H (2.6GHz) and 16GiB of RAM.
The docker runs the script src/main.py that runs run_experiments.py and run_analysis.py.
Arguments
--folder FOLDER: Set the dataset folder. (Default:dataset).--max-num-files N: Maximum number of files to read from can be used to limit the experiment. The files are traversed in sorted name order. Use 0 or a negative number to run all. (Default:0).--skip-after X: Time threshold in seconds to skip the tests of a specific version. If the threshold is reached twice, that version is skipped in the additional tests. (Default:0).--skip VALUE1 [VALUE2 ...]: List of versions to skip. Valid values:python,cpp,julia_basic,julia_c,julia_prealloc,julia_opt.
Owner
- Name: Abel Soares Siqueira
- Login: abelsiqueira
- Kind: user
- Location: Amsterdam - The Netherlands
- Company: Netherlands eScience Center
- Website: https://abelsiqueira.com
- Twitter: abel_siqueira
- Repositories: 331
- Profile: https://github.com/abelsiqueira
Citation (CITATION.cff)
# This CITATION.cff file was generated with cffinit.
# Visit https://bit.ly/cffinit to generate yours today!
cff-version: 1.2.0
title: >-
Calling Julia from Python - an experiment on data
loading
message: >-
If you use this software, please cite it using the
metadata from this file.
type: software
authors:
- given-names: Abel
family-names: Soares Siqueira
orcid: 'https://orcid.org/0000-0003-4451-281X'
- given-names: Faruk
family-names: Diblen
orcid: 'https://orcid.org/0000-0002-0989-929X'
identifiers:
- type: doi
value: 10.5281/zenodo.5708268
description: Zenodo DOI
repository-code: >-
https://github.com/abelsiqueira/call-julia-from-python-experiments
keywords:
- julia
- python
- interoperability
- data-loading
- irregular-data
license: MIT
GitHub Events
Total
Last Year
Issues and Pull Requests
Last synced: over 1 year ago
All Time
- Total issues: 23
- Total pull requests: 16
- Average time to close issues: 4 days
- Average time to close pull requests: 2 days
- Total issue authors: 1
- Total pull request authors: 2
- Average comments per issue: 0.3
- Average comments per pull request: 0.5
- Merged pull requests: 15
- Bot issues: 0
- Bot pull requests: 0
Past Year
- Issues: 0
- Pull requests: 0
- Average time to close issues: N/A
- Average time to close pull requests: N/A
- Issue authors: 0
- Pull request authors: 0
- Average comments per issue: 0
- Average comments per pull request: 0
- Merged pull requests: 0
- Bot issues: 0
- Bot pull requests: 0
Top Authors
Issue Authors
- abelsiqueira (15)
Pull Request Authors
- abelsiqueira (10)
- fdiblen (1)
Top Labels
Issue Labels
Pull Request Labels
Dependencies
- Pillow ==8.4.0
- cycler ==0.11.0
- fonttools ==4.28.3
- julia ==0.5.7
- kiwisolver ==1.3.2
- matplotlib ==3.5.0
- numpy ==1.21.4
- packaging ==21.3
- pandas ==1.3.4
- pybind11 ==2.8.1
- pybind11-global ==2.8.1
- pyparsing ==3.0.6
- python-dateutil ==2.8.2
- pytz ==2021.3
- seaborn ==0.11.2
- setuptools-scm ==6.3.2
- six ==1.16.0
- tomli ==1.2.2
- Pillow ==8.4.0
- cycler ==0.11.0
- fonttools ==4.28.3
- julia ==0.5.7
- kiwisolver ==1.3.2
- matplotlib ==3.5.0
- numpy ==1.21.4
- packaging ==21.3
- pandas ==1.3.4
- pybind11 ==2.8.1
- pybind11-global ==2.8.1
- pyparsing ==3.0.6
- python-dateutil ==2.8.2
- pytz ==2021.3
- seaborn ==0.11.2
- setuptools-scm ==6.3.2
- six ==1.16.0
- tomli ==1.2.2