mbgdml
Create, use, and analyze machine learning potentials within the many-body expansion framework.
Science Score: 75.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
✓CITATION.cff file
Found CITATION.cff file -
✓codemeta.json file
Found codemeta.json file -
✓.zenodo.json file
Found .zenodo.json file -
✓DOI references
Found 7 DOI reference(s) in README -
✓Academic publication links
Links to: zenodo.org -
○Committers with academic emails
-
✓Institutional organization owner
Organization keithgroup has institutional domain (www.klic.pitt.edu) -
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (15.6%) to scientific vocabulary
Keywords
Repository
Create, use, and analyze machine learning potentials within the many-body expansion framework.
Basic Info
- Host: GitHub
- Owner: keithgroup
- License: mit
- Language: Python
- Default Branch: main
- Homepage: https://keithgroup.github.io/mbGDML/
- Size: 47.4 MB
Statistics
- Stars: 10
- Watchers: 3
- Forks: 1
- Open Issues: 0
- Releases: 6
Topics
Metadata Files
README.md
mbGDML
Create, use, and analyze machine learning potentials within the many-body expansion framework.
Documentation
Motivation • Approach • Features • Installation • License
Motivation
Machine learning potentials (i.e., force fields) often rely on local descriptors for size transferability. These descriptors partition total properties into atomic contributions; however, they inherently neglect complicated long-range interactions by enforcing atomic radial cutoffs. Global descriptors encode the entire structure with no cutoffs and can capture interactions at all scales. However, they are restricted to systems with the same number of atoms.
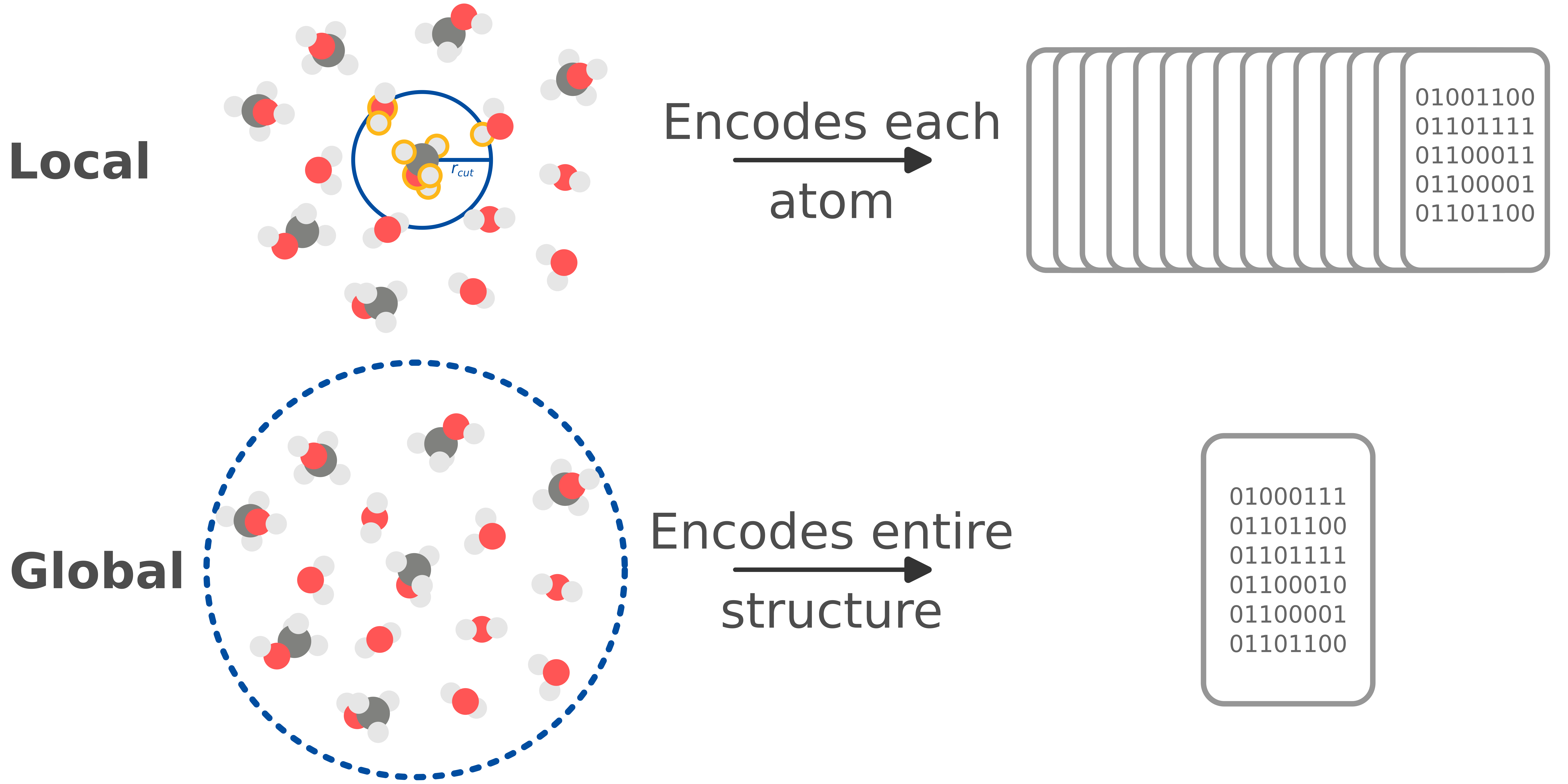
Gradient-domain machine learning (GDML) is one example of a ML potential with a global descriptor. GDML is unique because it trains directly on forces and recovers total energy through analytical integration. This provides substantially more information about the potential energy surface (PES) and allows for better interpolation between training data. As a result, GDML typically only needs 1000 structures to accurately learn energies and forces.

To date, GDML has been limited to the exact system it was trained on. This makes simulations on arbitrarily size systems, like solvents, futile.
Approach
Many-body expansions (MBEs) rigorously decomposes total (i.e., supersystem) energies into fundamental n-body interactions. This expansion is formally exact when all N-body interactions are accounted for. In practice, however, it is typically truncated to the third order. One can then model any system by summing up 1-, 2-, and 3-body contributions.
MBEs driven by GDML potentials trained on n-body interactions is a promising approach for size-transferable potentials. Furthermore, GDML model's remarkable data efficiency enables training on highly accurate quantum chemical methods.
Features
Train
- Train GDML models using grid searches, Bayesian optimization, or both on CPUs.
- Custom loss functions.
- Iterative training procedure for automated curation of optimal training sets.
Predict
- Many-body predictions with GDML, SchNet and GAP potentials.
- Parallel GDML predictions with ray from a laptop to multiple nodes.
- Periodic structures with the minimum-image convention.
- Alchemical predictions by tuning out 2- or 3-body contributions of specific entities.
Analysis
- Prediction sets that store decomposed predictions for further analysis.
- Radial distribution functions.
- Cluster and identify problematic (i.e., high error) structures using sklearn.
Interfaces
- Atomic Simulation Environment (ASE) for geometry optimizations, molecular dynamics simulations, and more.
Installation
You can install mbGDML from PyPI by using pip install mbGDML.
Or, the latest development version can be installed directly from the GitHub repository or from TestPyPI.
text
git clone https://github.com/keithgroup/mbGDML
cd mbGDML
pip install .
Citing this work
If you find this code helpful in your research or project, please consider citing the following paper:
Maldonado, A. M.; Poltavsky, I.; Vassilev-Galindo, V.; Tkatchenko, A.; Keith, J. A. Modeling molecular ensembles with gradient-domain machine learning force fields. Digital Discovery 2023, 2 (3), 871-880. DOI: 10.1039/D3DD00011G.
bibtex
@article{maldonado2023modeling,
title={Modeling molecular ensembles with gradient-domain machine learning force fields},
author={Maldonado, Alex M and Poltavsky, Igor and Vassilev-Galindo, Valentin and Tkatchenko, Alexandre and Keith, John A},
journal={Digital Discovery},
volume={2},
number={3},
pages={871--880},
year={2023},
publisher={Royal Society of Chemistry},
doi={10.1039/D3DD00011G}
}
Citing the paper helps acknowledge the effort put into developing and maintaining this codebase, and it provides a way to support further research and development. Thank you for your support!
License
Distributed under the MIT License. See LICENSE for more information.
Owner
- Name: Keith Lab in Computational Chemistry
- Login: keithgroup
- Kind: organization
- Location: Pittsburgh, PA
- Website: https://www.klic.pitt.edu/
- Twitter: jakeith01
- Repositories: 3
- Profile: https://github.com/keithgroup
Citation (CITATION.cff)
message: If you use this software, please cite it as below.
title: Many-body gradient-domain machine learning (mbGDML)
authors:
- family-names: Maldonado
given-names: Alex M.
orcid: "https://orcid.org/0000-0003-3280-062X"
- family-names: Keith
given-names: John A.
orcid: "https://orcid.org/0000-0002-6583-6322"
type: software
version: v0.1.1
license: MIT
preferred-citation:
type: article
title: "Modeling molecular ensembles with gradient-domain machine learning force fields"
authors:
- family-names: "Maldonado"
given-names: "Alex M."
orcid: "https://orcid.org/0000-0003-3280-062X"
- family-names: "Poltavsky"
given-names: "Igor"
orcid: "https://orcid.org/0000-0002-3188-7017"
- family-names: "Vassilev-Galindo"
given-names: "Valentin"
orcid: "https://orcid.org/0000-0001-7532-3590"
- family-names: "Tkatchenko"
given-names: "Alexandre"
email: "alexandre.tkatchenko@uni.lu"
orcid: "https://orcid.org/0000-0002-1012-4854"
- family-names: "Keith"
given-names: "John A."
email: "jakeith@pitt.edu"
orcid: "https://orcid.org/0000-0002-6583-6322"
journal: "Digital Discovery"
volume: 2
issue: 3
pages: "871-880"
year: 2023
doi: "10.1039/D3DD00011G"
contact:
- email: "alex.maldonado@pitt.edu"
family-names: "Maldonado"
given-names: "Alex"
cff-version: 1.2.0
GitHub Events
Total
- Pull request event: 1
- Create event: 1
Last Year
- Pull request event: 1
- Create event: 1
Committers
Last synced: almost 3 years ago
All Time
- Total Commits: 572
- Total Committers: 2
- Avg Commits per committer: 286.0
- Development Distribution Score (DDS): 0.301
Top Committers
| Name | Commits | |
|---|---|---|
| Alex Maldonado | a****o@g****m | 400 |
| Alex Maldonado | a****3@g****m | 172 |
Issues and Pull Requests
Last synced: 8 months ago
All Time
- Total issues: 0
- Total pull requests: 14
- Average time to close issues: N/A
- Average time to close pull requests: 2 days
- Total issue authors: 0
- Total pull request authors: 1
- Average comments per issue: 0
- Average comments per pull request: 0.86
- Merged pull requests: 14
- Bot issues: 0
- Bot pull requests: 0
Past Year
- Issues: 0
- Pull requests: 0
- Average time to close issues: N/A
- Average time to close pull requests: N/A
- Issue authors: 0
- Pull request authors: 0
- Average comments per issue: 0
- Average comments per pull request: 0
- Merged pull requests: 0
- Bot issues: 0
- Bot pull requests: 0
Top Authors
Issue Authors
Pull Request Authors
- aalexmmaldonado (14)
Top Labels
Issue Labels
Pull Request Labels
Dependencies
- actions/checkout v2 composite
- actions/setup-python v2 composite
- codecov/codecov-action v3 composite
- actions/checkout v3 composite
- actions/configure-pages v2 composite
- actions/deploy-pages v1 composite
- actions/upload-pages-artifact v1 composite
- actions/checkout v3 composite
- actions/setup-python v2 composite
- github/super-linter v4 composite
- actions/checkout v3 composite
- actions/setup-python v3 composite
- actions/checkout v3 composite
- actions/setup-python v3 composite
- pypa/gh-action-pypi-publish v1.5.2 composite
- actions/checkout v3 composite
- actions/setup-python v3 composite
- pypa/gh-action-pypi-publish v1.5.2 composite




