qpcr-pipeline
Auto-analysis for Real-Time PCR (qPCR) data
Science Score: 44.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
✓CITATION.cff file
Found CITATION.cff file -
✓codemeta.json file
Found codemeta.json file -
✓.zenodo.json file
Found .zenodo.json file -
○DOI references
-
○Academic publication links
-
○Committers with academic emails
-
○Institutional organization owner
-
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (8.2%) to scientific vocabulary
Keywords
python
qpcr
r
Last synced: 7 months ago
·
JSON representation
·
Repository
Auto-analysis for Real-Time PCR (qPCR) data
Basic Info
- Host: GitHub
- Owner: tiramisutes
- License: apache-2.0
- Language: R
- Default Branch: master
- Homepage: https://ihope.shinyapps.io/qRT-PCR-Pipeline/
- Size: 101 KB
Statistics
- Stars: 37
- Watchers: 1
- Forks: 16
- Open Issues: 0
- Releases: 0
Topics
python
qpcr
r
Created over 7 years ago
· Last pushed over 2 years ago
Metadata Files
Readme
License
Citation
README.md
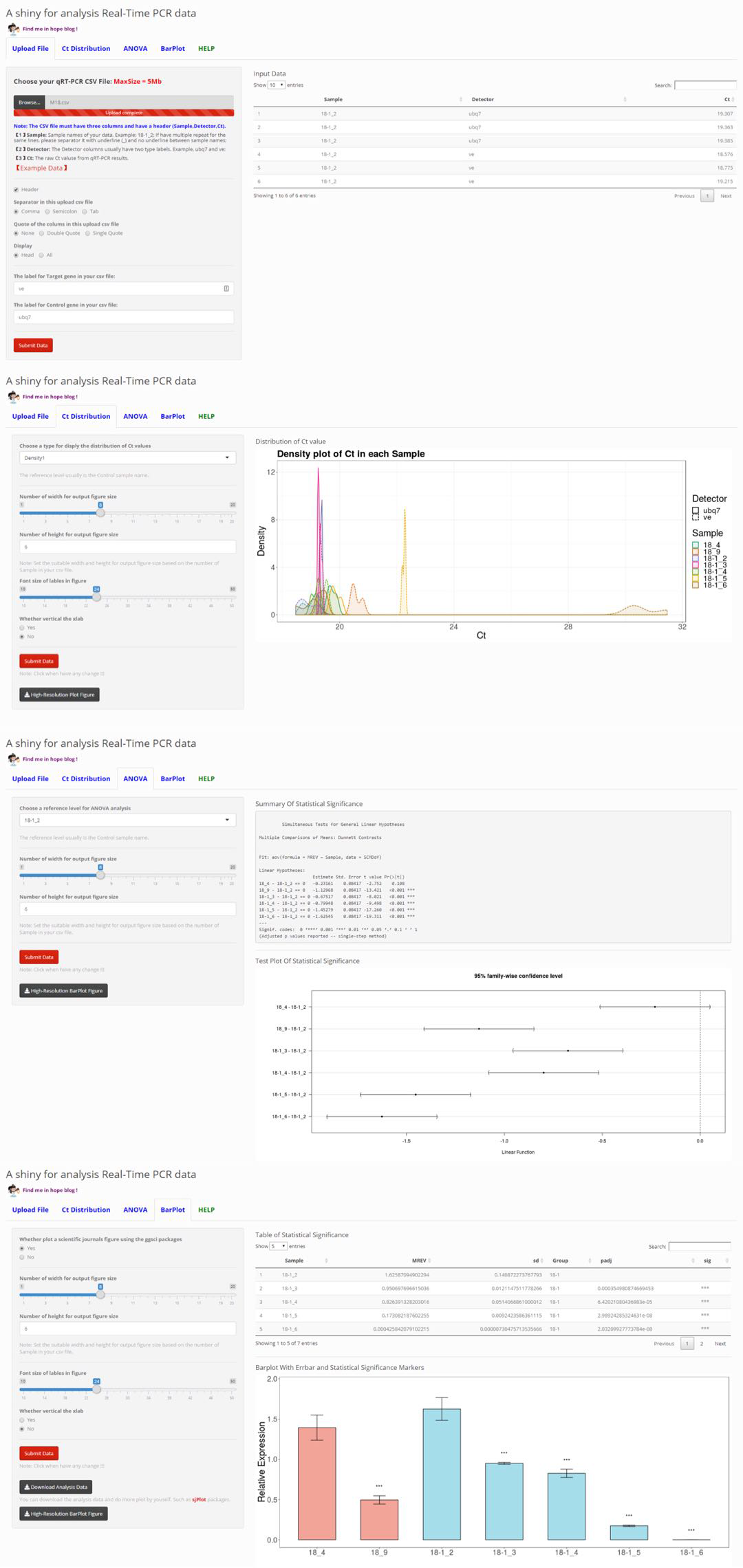
Real-Time PCR (qPCR) Pipeline
Auto-analysis for Real-Time PCR (qPCR) data.
This qPCR experiment usually have two group: Control (CK) and sample, and at least two type genes: Housekeeping Gene (HG) and Target Gene (TG). In our lab this usually label with ub7 and ve.
Congratulations! A alternative way to analysis of qPCR data can be find in : A shiny for analysis qRT-PCR data
- Dependencies
- INPUT
- USAGE
- OUTPUT ## 1. Dependencies
- python
- collections
- itertools
- csv
- pandas
- numpy
- sys
- R
- ggplot2
- gridExtra
- ggthemes
- ggsci
- reshape2
- dplyr
## 2. INPUT
There is a example input data in
Example_data/. ## 3. USAGE./qPCR.sh A B C D E FArgument:
- A: absolute path of Input data (e,
/public/home/hope/qPCR) - B: Input Data (e, rawinputqPCR.csv)
- C: Width of output pdf file
- D: Height of output pdf file
- E: Whether plot a scientific journals figure using the ggsci packages (yes or no)
- F: The sample name used as reference level in ANOVA analysis
Running on the example data:
./qPCR.sh /public/home/hope/qPCR raw_input_qPCR.csv 18 8 no CK
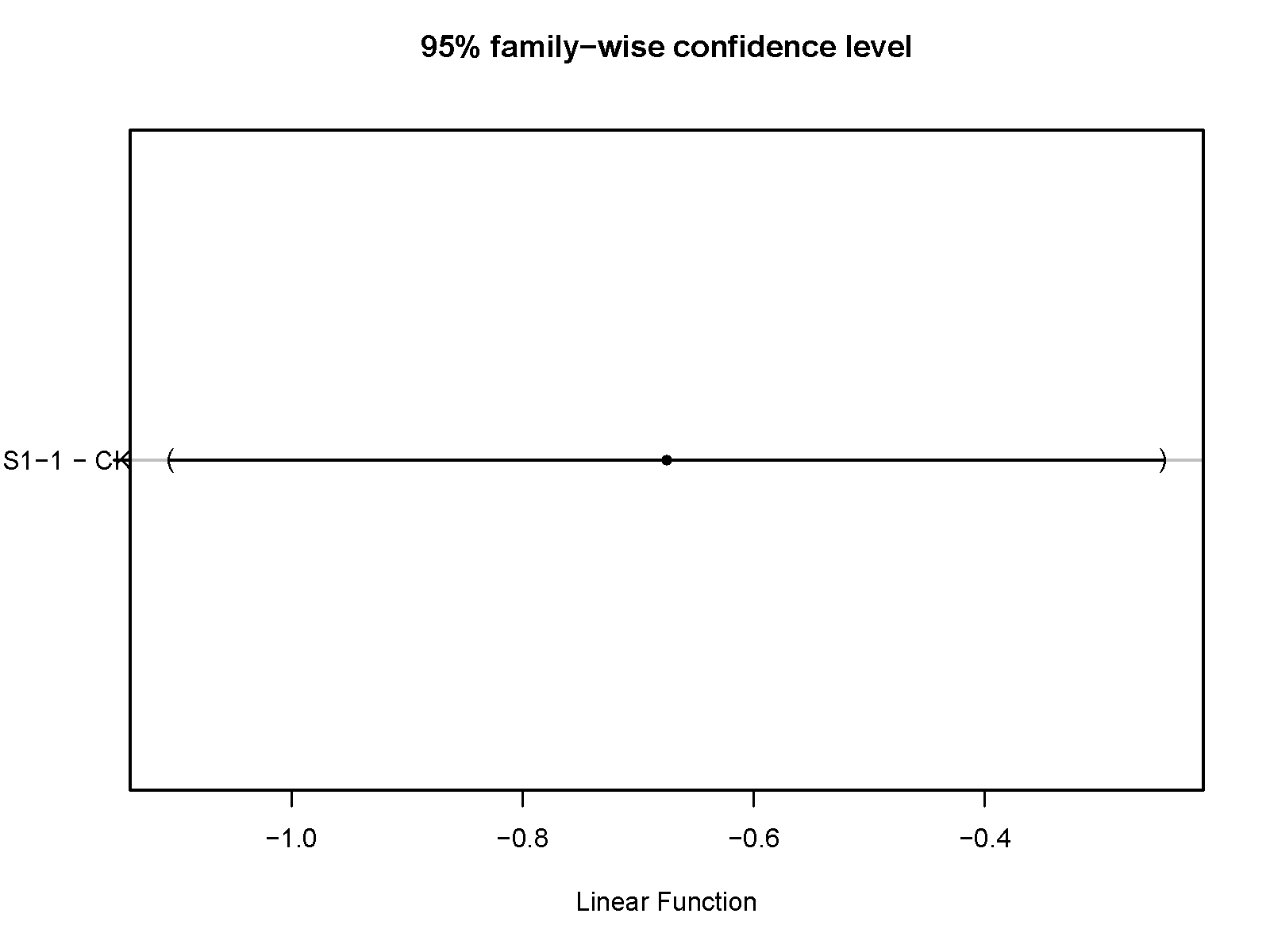
4. OUTPUT
- rawinputqPCR_ubq7.csv
- rawinputqPCR_ve.csv
- RQrawinputqPCR_ve.csv
- RQrawinputqPCR_ubq7.csv
- rawinputqPCR_R.csv
- rawinputqPCR_ANOVA.csv
- ProrawinputqPCRR.csv
- Prorawinput_qPCR.pdf
- rawinputqPCR_T-test.pdf

Owner
- Name: hope
- Login: tiramisutes
- Kind: user
- Website: http://tiramisutes.github.io/
- Twitter: hopetogy
- Repositories: 1
- Profile: https://github.com/tiramisutes
Bioinformatician, interested in genomics.
Citation (CITATION.cff)
cff-version: 1.2.0
message: "If you use this software, please cite it as below."
authors:
- family-names: "Lisa"
given-names: "Mona"
orcid: "https://orcid.org/0000-0000-0000-0000"
- family-names: "Bot"
given-names: "Hew"
orcid: "https://orcid.org/0000-0000-0000-0000"
title: "My Research Software"
version: 2.0.4
doi: 10.5281/zenodo.1234
date-released: 2017-12-18
url: "https://github.com/github/linguist"
preferred-citation:
type: article
authors:
- family-names: "Lisa"
given-names: "Mona"
orcid: "https://orcid.org/0000-0000-0000-0000"
- family-names: "Bot"
given-names: "Hew"
orcid: "https://orcid.org/0000-0000-0000-0000"
doi: "10.0000/00000"
journal: "Journal Title"
month: 9
start: 1 # First page number
end: 10 # Last page number
title: "My awesome research software"
issue: 1
volume: 1
year: 2021
GitHub Events
Total
- Watch event: 2
- Fork event: 1
Last Year
- Watch event: 2
- Fork event: 1
Committers
Last synced: over 2 years ago
Top Committers
| Name | Commits | |
|---|---|---|
| hope | 1****8@q****m | 70 |
Committer Domains (Top 20 + Academic)
qq.com: 1