MSMExplorer
MSMExplorer: Data Visualizations for Biomolecular Dynamics - Published in JOSS (2017)
Science Score: 95.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
○CITATION.cff file
-
✓codemeta.json file
Found codemeta.json file -
✓.zenodo.json file
Found .zenodo.json file -
✓DOI references
Found 3 DOI reference(s) in README and JOSS metadata -
✓Academic publication links
Links to: joss.theoj.org -
✓Committers with academic emails
4 of 7 committers (57.1%) from academic institutions -
○Institutional organization owner
-
✓JOSS paper metadata
Published in Journal of Open Source Software
Keywords from Contributors
Repository
Data visualizations for biomolecular dynamics
Basic Info
- Host: GitHub
- Owner: msmexplorer
- License: mit
- Language: Python
- Default Branch: master
- Homepage: http://msmbuilder.org/msmexplorer/
- Size: 559 KB
Statistics
- Stars: 5
- Watchers: 1
- Forks: 3
- Open Issues: 0
- Releases: 0
Metadata Files
README.md
MSMExplorer: data visualizations for biomolecular dynamics
MSMExplorer is a Python visualization library for statistical models of biomolecular dynamics. It provides a high-level interface for drawing attractive statistical graphics with MSMBuilder.
Documentation
Online documentation is available here. It includes IPython notebooks, detailed API documentation, and other useful info.
There are docs for the development version here. These should correspond with the github master branch.
Examples
```python from msmbuilder.example_datasets import FsPeptide from msmbuilder.featurizer import RMSDFeaturizer
import msmexplorer as msme
Load Fs Peptide Data
traj = FsPeptide().get().trajectories[0]
Calculate RMSD
featurizer = RMSDFeaturizer(referencetraj=traj[0]) rmsd = featurizer.partialtransform(traj).flatten()
Plot Trace
msme.plot_trace(rmsd, label='traj0', xlabel='Timestep', ylabel='RMSD (nm)') ```
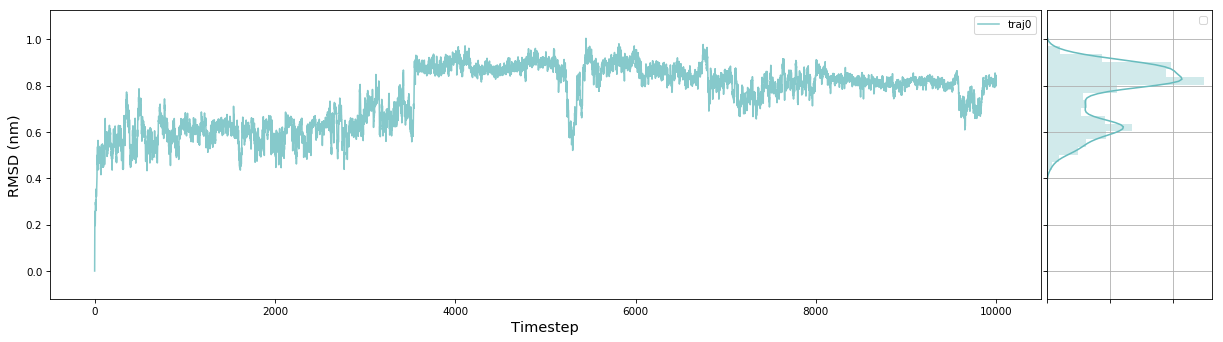
The documentation has an example gallery with short scripts showing how to use different parts of the package.
Dependencies
- Python 3.4+
Mandatory
The latest versions of the following packages are required:
Installation
The preferred installation mechanism for msmexplorer is with conda:
bash
$ conda install -c omnia msmexplorer
If you don't have conda, or are new to scientific python, we recommend that you download the Anaconda scientific python distribution.
To install from PyPI, just do:
pip install msmexplorer
You may instead want to use the development version from Github, by running
pip install git+git://github.com/msmexplorer/msmexplorer.git#egg=msmexplorer
Development
All development happens here, on Github.
If you're interested in contributing to MSMExplorer, please refer to our Contributing guide.
Support
Please submit any bugs or questions to the Github issue tracker.
License
Released under a MIT license
Citing
bibtex
@article{msmexplorer,
doi = {10.21105/joss.00188},
url = {https://doi.org/10.21105%2Fjoss.00188},
year = {2017},
month = {apr},
publisher = {The Open Journal},
volume = {2},
number = {12},
author = {Carlos X. Hern{\'{a}}ndez and Matthew P. Harrigan and Mohammad M. Sultan and Vijay S. Pande},
title = {{MSMExplorer}: Data Visualizations for Biomolecular Dynamics},
journal = {The Journal of Open Source Software}
}
Owner
- Name: MSMExplorer
- Login: msmexplorer
- Kind: organization
- Location: Stanford University
- Website: http://msmexplorer.github.io/msmexplorer-d3/
- Repositories: 2
- Profile: https://github.com/msmexplorer
Analytics for Markov state models
JOSS Publication
MSMExplorer: Data Visualizations for Biomolecular Dynamics
Authors
Stanford University
Tags
plotting molecular dynamics markov modelGitHub Events
Total
- Watch event: 1
Last Year
- Watch event: 1
Committers
Last synced: 7 months ago
Top Committers
| Name | Commits | |
|---|---|---|
| Carlos Hernandez | c****h@s****u | 133 |
| Carlos Hernández | c****z | 43 |
| Juan Eiros | j****z@g****m | 27 |
| Matthew Harrigan | h****n@s****u | 9 |
| msultan | m****n@s****u | 4 |
| Juan Eiros | j****4@i****k | 3 |
| Shyam Saladi | s****i | 2 |
Committer Domains (Top 20 + Academic)
Issues and Pull Requests
Last synced: 6 months ago
All Time
- Total issues: 0
- Total pull requests: 0
- Average time to close issues: N/A
- Average time to close pull requests: N/A
- Total issue authors: 0
- Total pull request authors: 0
- Average comments per issue: 0
- Average comments per pull request: 0
- Merged pull requests: 0
- Bot issues: 0
- Bot pull requests: 0
Past Year
- Issues: 0
- Pull requests: 0
- Average time to close issues: N/A
- Average time to close pull requests: N/A
- Issue authors: 0
- Pull request authors: 0
- Average comments per issue: 0
- Average comments per pull request: 0
- Merged pull requests: 0
- Bot issues: 0
- Bot pull requests: 0
Top Authors
Issue Authors
Pull Request Authors
Top Labels
Issue Labels
Pull Request Labels
Dependencies
- jinja2 *
- jupyter *
- msmb_data *
- msmbuilder *
- nbconvert *
- numpydoc *
- pyparsing *
- python *
- traitlets *
- corner *
- matplotlib *
- mdtraj *
- mpld3 *
- msmbuilder *
- nglview *
- numpy *
- pandas *
- seaborn *