renal_segmentor
An application to segment kidneys from renal MRI data using a convolutional neural network.
Science Score: 77.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
✓CITATION.cff file
Found CITATION.cff file -
✓codemeta.json file
Found codemeta.json file -
✓.zenodo.json file
Found .zenodo.json file -
✓DOI references
Found 14 DOI reference(s) in README -
✓Academic publication links
Links to: zenodo.org -
✓Committers with academic emails
1 of 3 committers (33.3%) from academic institutions -
○Institutional organization owner
-
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (17.3%) to scientific vocabulary
Repository
An application to segment kidneys from renal MRI data using a convolutional neural network.
Basic Info
Statistics
- Stars: 2
- Watchers: 1
- Forks: 0
- Open Issues: 6
- Releases: 28
Metadata Files
README.md
Renal Segmentor
An application and Python package to segment kidneys from renal MRI data using a convolutional neural network (CNN).
Using the segmentor
The easiest way to make use of the segmentor is to download the windows executable, this allows you to mask data with a stand-alone application. The executable can either be run as a GUI or a command line application allowing it to be integrated into bash scripts.
Alternatively, the methods used by the segmentor are available as a Python package and can be integrated into existing Python pipelines. Instructions for making use of the segmentor via each method are given below.
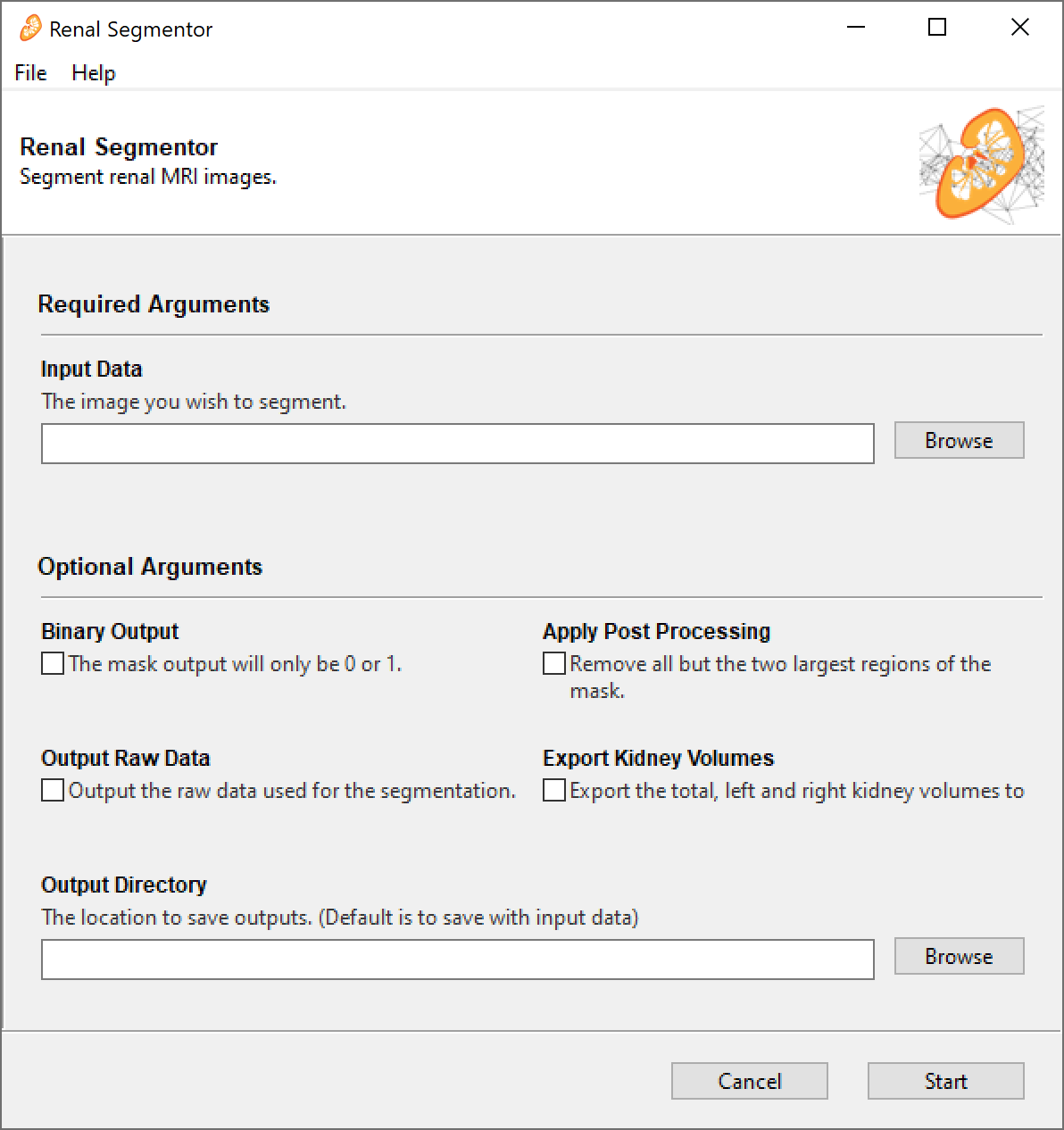
As a Graphical User Interface (GUI)
- Download the latest release
- Double click
renal_segmentor.exe. The GUI takes quite a long time to load (~30 sec) and doesn't have a splash screen so be patient. - Once the GUI has loaded, click
Browseand select all the raw data you want to segment. You can select multiple files at once. Supported file types are.PAR/.REC,.nii,.nii.gzand.img/.hdr, other file types supported by nibable may work but are untested. - If you want the mask to be just 0s and 1s tick the
binarycheck box, if you want the CNNs probability that the voxel is a kidney, leave it unchecked. - Post-processing can be applied, this discards all but the two largest connected regions in the mask in theory retaining only the two kidneys and removing any erroneous unconnected regions. It should be noted that because post-processing keeps only two regions, care should be taken if using this application with transplant patients as they may have three kidneys.
- Tick the
rawcheckbox if you want the raw image data to be saved as a.nii.gzfile in the same location as the mask (can be useful if you're using.PAR/RECas your input). - If you would like a
.csvfile containing the Total Kidney Volume (TKV), Left Kidney Volume (LKV) and Right Kidney Volume (RKV) for each image that was segmented tick theExport Kidney Volumesbox. - The masks are output from the segmentor as
.nii.gzwith_maskadded to the original file name e.g. the mask ofsub_01.PARissub_01_mask.nii.gz. By default, the mask is output to the same place as the raw data, if you would like the masks to be output to a different directory clickBrowseunderOutput Directoryand select the folder you wish the masks to go to. - Click start.
- The application will run and a few seconds later a box will appear saying the program completed successfully. The first time you run a segmentation will take a little longer as the latest algorithm weights are downloaded from the internet at this point.
- If you want to segment some more data click the
editbutton on the bottom of the finished screen, if you're done, clickclose.
Via a Command Line Interface (CLI)
- Download the latest release
- Run the
renal_segmentor.exe -hto generate a list of available parameters. The application runs via a command line if any input arguments are specified, if not, it opens as a GUI.
As a Python package
- Activate the python environment you want to install the package on and run
pip install renalsegmentor. If you want to install the additional dependencies required for the GUI, runpip install renalsegmentor[gui]however to use the segmentor as a python package, these are not required. - The example code snippet below will generate a mask of
T2w.nii.gzas a numpy array and print the TKV to the terminal.
python
from segment import Tkv
segmentation = Tkv('T2w.nii.gz')
mask = segmentation.get_mask()
print(f'Total Kidney Volume = {segmentation.tkv:.2f} ml')
Citing Renal Segmentor
If you have made use of renal segmentor for your work, please cite Daniel AJ, et al. Automated renal segmentation in healthy and chronic kidney disease subjects using a convolutional neural network. Magnetic Resonance in Medicine 2021;86:1125–1136 doi: https://doi.org/10.1002/mrm.28768. Alternatively if you wish to cite a specific software version, each release has an individual DOI on Zenodo, the DOI for the latest release can be found here.
How it works
The methods used in this software are outlined in Daniel AJ, et al. Automated renal segmentation in healthy and chronic kidney disease subjects using a convolutional neural network. Magnetic Resonance in Medicine 2021;86:1125–1136 doi: https://doi.org/10.1002/mrm.28768.
The dataset used to train this network is freely available on Zenodo, doi: https://doi.org/10.5281/zenodo.5153567.
Contributing
Feel free to open a pull request if you have a feature you want to develop or drop me an email to discuss things further.
Owner
- Name: Alex Daniel
- Login: alexdaniel654
- Kind: user
- Location: Nottingham
- Company: The University of Nottingham @SPMIC-UoN
- Website: alexdaniel.info
- Repositories: 21
- Profile: https://github.com/alexdaniel654
Computational physicist working on renal MRI at The University of Nottingham.
Citation (CITATION.cff)
cff-version: 1.2.0 message: "If you use this software, please cite it as below." authors: - family-names: "Daniel" given-names: "Alexander J" orcid: "https://orcid.org/0000-0003-2353-3283" - family-names: "Francis" given-names: "Susan T" orcid: "https://orcid.org/0000-0003-0903-7507" title: "Renal Segmentor" version: 1.3.9 doi: 10.5281/zenodo.5796277 date-released: 2024-03-20 url: "https://github.com/alexdaniel654/Renal_Segmentor"
GitHub Events
Total
Last Year
Committers
Last synced: almost 3 years ago
All Time
- Total Commits: 324
- Total Committers: 3
- Avg Commits per committer: 108.0
- Development Distribution Score (DDS): 0.448
Top Committers
| Name | Commits | |
|---|---|---|
| Alex Daniel | a****l@n****k | 179 |
| dependabot-preview[bot] | 2****]@u****m | 97 |
| dependabot[bot] | 4****]@u****m | 48 |
Committer Domains (Top 20 + Academic)
Issues and Pull Requests
Last synced: 7 months ago
All Time
- Total issues: 8
- Total pull requests: 111
- Average time to close issues: about 2 months
- Average time to close pull requests: 23 days
- Total issue authors: 1
- Total pull request authors: 2
- Average comments per issue: 0.0
- Average comments per pull request: 1.44
- Merged pull requests: 59
- Bot issues: 0
- Bot pull requests: 95
Past Year
- Issues: 0
- Pull requests: 0
- Average time to close issues: N/A
- Average time to close pull requests: N/A
- Issue authors: 0
- Pull request authors: 0
- Average comments per issue: 0
- Average comments per pull request: 0
- Merged pull requests: 0
- Bot issues: 0
- Bot pull requests: 0
Top Authors
Issue Authors
- alexdaniel654 (8)
Pull Request Authors
- dependabot[bot] (94)
- alexdaniel654 (18)
Top Labels
Issue Labels
Pull Request Labels
Dependencies
- nibabel ==3.2.2
- numpy ==1.22.0
- pandas ==1.4.2
- pytest ==7.1.2
- scikit-image ==0.19.2
- tensorflow ==2.9.1
- wget ==3.2
- Gooey ==1.0.8.1
- pyinstaller ==5.1
- actions/checkout v2 composite
- actions/download-artifact v2 composite
- actions/setup-python v2 composite
- actions/upload-artifact v2 composite
- docker://antonyurchenko/git-release v4 composite
- pypa/gh-action-pypi-publish release/v1 composite
- actions/checkout v2 composite
- actions/setup-python v2 composite
- codecov/codecov-action v2 composite