containers
Containers "distribution" for reproducible neuroimaging
Science Score: 44.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
✓CITATION.cff file
Found CITATION.cff file -
✓codemeta.json file
Found codemeta.json file -
✓.zenodo.json file
Found .zenodo.json file -
○DOI references
-
○Academic publication links
-
○Academic email domains
-
○Institutional organization owner
-
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (12.5%) to scientific vocabulary
Repository
Containers "distribution" for reproducible neuroimaging
Basic Info
- Host: GitHub
- Owner: ReproNim
- License: apache-2.0
- Language: Roff
- Default Branch: master
- Size: 2.9 MB
Statistics
- Stars: 28
- Watchers: 7
- Forks: 16
- Open Issues: 46
- Releases: 5
Metadata Files
README.md
ReproNim/containers - containerized environments for reproducible neuroimaging
This repository provides a DataLad dataset (git/git-annex
repository) with a collection of popular computational tools provided
within ready to use containerized environments. At the moment it
provides only Singularity images. Versions of all images are tracked using
git-annex with content of the images provided from a dedicated
Singularity Hub Collection and http://datasets.datalad.org (AKA /// of
DataLad) or other original collections.
The aims for this project is
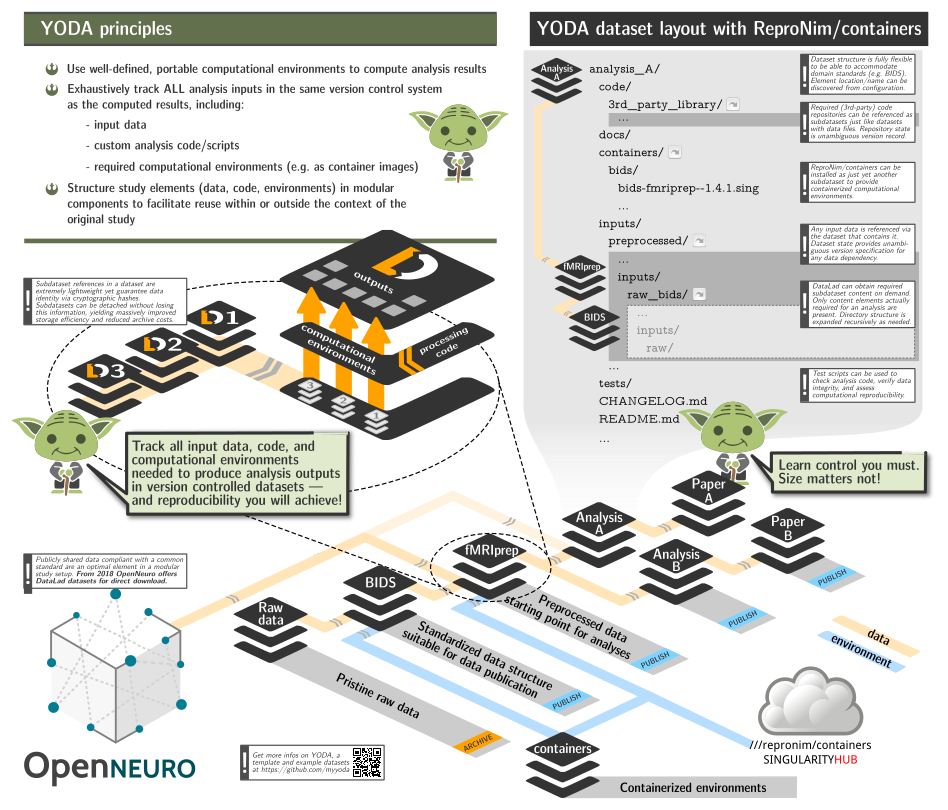
- to be able to include this repository as a subdataset within larger study (super)datasets to facilitate rapid and reproducible computation, while adhering to YODA principles and retaining clear and unambiguous association between data, code, and computing environments using git/git-annex/DataLad;
- to assist with containers execution in "sanitized" environments: no
$HOMEor system-wide/tmpis bind-mounted inside the containers, no environment variables from the host system made available inside; - make Singularity images transparently usable on non-Linux (OSX) systems via Docker.
All images are "registered" within the dataset for execution using datalad containers-run, so it is trivial to list available containers:
shell
$> datalad containers-list
arg-test -> scripts/tests/arg-test.simg
bids-aa -> images/bids/bids-aa--0.2.0.sing
bids-afni-proc -> images/bids/bids-afni-proc--0.0.2.sing
bids-antscorticalthickness -> images/bids/bids-antscorticalthickness--2.2.0-1.sing
bids-baracus -> images/bids/bids-baracus--1.1.2.sing
bids-brainiak-srm -> images/bids/bids-brainiak-srm--latest.sing
... many more to list them all ...
and execute either via datalad containers-run (which would also take care
about getting them first if not present):
```shell
$> datalad containers-run -n bids-validator -- --help
[INFO ] Making sure inputs are available (this may take some time)
[INFO ] == Command start (output follows) =====
Usage: bids-validator
Options: --help, -h Show help [boolean] --version, -v Show version number [boolean] --ignoreWarnings Disregard non-critical issues [boolean] --ignoreNiftiHeaders Disregard NIfTI header content during validation [boolean] --verbose Log more extensive information about issues [boolean] --json Output results as JSON [boolean] --config, -c Optional configuration file. See https://github.com/bids-standard/bids-validator for more info
This tool checks if a dataset in a given directory is compatible with the Brain Imaging Data Structure specification. To learn more about Brain Imaging Data Structure visit http://bids.neuroimaging.io [INFO ] == Command exit (modification check follows) ===== action summary: get (notneeded: 1) save (notneeded: 1) ```
or first getting them using datalad get and then either using
singularity run or exec directly, or (recommended) via
scripts/singularity_cmd. That is the helper which is used by
containers-run (see .datalad/config).
scripts/singularity_cmd
Singularity execution by default is optimized for convenience and not for reproducibility. This helper script assists in making singularity execution reproducible by
- disabling passing environment variables inside your containerized environment
- creating temporary
/tmpdirectory for the environment, so there is no interaction with file paths outside of the current directory (which should ideally be a DataLad dataset) - using custom and nearly empty binds/HOME HOME directory, so there is no possible leakage of locally user-level installed Python and other modules to affect your computation
The binds/HOME also provides a custom minimalistic .bashrc file with e.g. a customized prompt to inform you about which image you are in ATM for use in interactive sessions:
$> scripts/singularity_cmd exec images/repronim/repronim-reproin--0.5.4.sing bash
singularity:repronim-reproin--0.5.4 > yoh@hopa:/home/yoh/proj/repronim/containers$ heudiconv --version
0.5.4
Singularity via Docker
On non-Linux systems, or if REPRONIM_USE_DOCKER environment variable is set to a non-empty value,
scripts/singularity_cmd will use Docker shim image (in privileged mode) to run
singularity within it. All necessary paths will be bind mounted as with a regular direct execution using
singularity.
Interactive sessions
See WiP PR #9 to establish "reproducible interactive sessions" with the help of that script.
Conventions
Container image files
Singularity image files have .sing extension. Since we are providing
a custom filename to store the file at, we cannot guess the format of
the container (e.g., either it is
.sif),
so we just use uniform .sing extension.
A typical YODA workflow
Lets summarize YODA principles as a possible workflow:
- create a new dataset which would contain results and everything needed to obtain them
- install/add subdatasets(code, other datasets, containers)
- perform the analysis using only materials available within the reach of this dataset.
Let's assume that our goal is to do Quality Control of an MRI dataset (which is available as DataLad dataset ds000003). We will create a new dataset with the output of the QC results (as analyzed by mriqc BIDS-App). mriqc is provided by the ReproNim/containers dataset of containers. Below, we execute a simple analysis workflow which adheres to YODA principles and we end up with a dataset that contains all components necessary a history of how it was achieved.
This would help to guarantee reproducibility in the future because all the materials would be reachable within that dataset.
Runnable script
For advanced users who are comfortable with DataLad, the following script may give you everything you need.
The version of the script with all commands explained
```shell #!/bin/sh ( # so it could be just copy pasted or used as a script PS4='> '; set -xeu # to see what we are doing and exit upon error # Work in some temporary directory cd $(mktemp -d ${TMPDIR:-/tmp}/repro-XXXXXXX) # Create a dataset to contain mriqc output datalad create -d ds000003-qc -c text2git cd ds000003-qc # Install our containers collection: datalad install -d . -s ///repronim/containers code/containers # Optionally -- copy container of interest definition to the current (or desired) # version # to facilitate reproducibility while still being able to upgrade containers # subdataset if so desired to get access to newer versions. # We will also use 0.16.0 since newer ones require more memory and # would fail to run on CI. datalad run -m "Downgrade/Freeze mriqc container version" \ code/containers/scripts/freeze_versions --save-dataset=. bids-mriqc=0.16.0 # That version of mriqc does not have an option --no-datalad-get we had to # hardcode for mriqc to workaround an issue. So let's remove it datalad run -m "Remove ad-hoc option for mriqc for older frozen version" sed -i -e 's, --no-datalad-get,,g' .datalad/config # Install input data: datalad install -d . -s https://github.com/ReproNim/ds000003-demo sourcedata/raw # Setup git to ignore workdir to be used by pipelines echo "workdir/" > .gitignore && datalad save -m "Ignore workdir" .gitignore # Execute desired preprocessing while creating a provenance record # in git history datalad containers-run \ -n bids-mriqc \ --input sourcedata/raw \ --output . \ '{inputs}' '{outputs}' participant group -w workdir ) ```Walkthrough
For users who are new to these components, we will walk through how these components are used together in a typical YODA workflow. the steps
bash
mkdir ~/my-experiments
cd ~/my-experiments
datalad create -d ds000003-qc -c text2git
cd ds000003-qc
DataLad has created a new directory for our results, ds000003-qc.
According to YODA principles, this dataset should also contain our input
data, code, and anything else we need to run the analysis.
Install the input dataset:
bash
datalad install -d . -s https://github.com/ReproNim/ds000003-demo sourcedata/raw
Next we install the ReproNim/containers collection.
bash
datalad install -d . -s ///repronim/containers code/containers
Now let's take a look at what we have.
ds000003-qc/ # The root dataset contains everything
|- sourcedata/
| \- raw/ # we call it source, but it is actually ds000003-demo "raw" BIDS dataset
\- code/
\- containers/ # repronim/containers, this is where our non-custom code lives
Freezing Container Image Versions
freeze_versions is an optional step that will record and "freeze" the
version of the container used. Even if the ///repronim/containers dataset is
upgraded with a newer version of our container, we are "pinned" to the
container we explicitly determined. Note: To switch version of the container
(e.g., to upgrade to a new one), rerun freeze_versions script with the version
specified.
The container version can be "frozen" into the clone of the ///repronim/containers
dataset, or the top-level dataset.
Option 1: Top level dataset (recommended)
```bash
Run from ~/my-experiments/ds000003-qc
datalad run -m "Downgrade/Freeze mriqc container version" \ code/containers/scripts/freeze_versions --save-dataset=. bids-mriqc=0.16.0 ```
Option 2: ///repronim/containers
```bash
Run from ~/my-experiments/ds000003-qc/
datalad run -m "Downgrade/Freeze mriqc container version" \ code/containers/scripts/freeze_versions bids-mriqc=0.16.0 ```
Note: It is recommended to freeze a container image version into the
top-level dataset to simplify reuse. If ///repronim/containers is
modified in any way, the author must ensure that their altered fork of
///repronim/containers is publicly available and that its URL
specified in the .gitmodules. By freezing into the top-level dataset
instead, authors do not need to host a modified version of
///reporonim/containers.
Fixup datalad config
The version of mriqc we are using does not have an option --no-datalad-get which is hardcoded
into mriqc config, so we should remove it.
bash
datalad run -m "Remove ad-hoc option for mriqc for older frozen version" sed -i -e 's, --no-datalad-get,,g' .datalad/config
Running the Containers
When we run the bids-mriqc container, it will need a working directory
for intermediate files. These are not helpful to commit, so we will
tell git (and datalad) to ignore the whole directory.
bash
echo "workdir/" > .gitignore && datalad save -m "Ignore workdir" .gitignore
Now we use datalad containers-run to perform the analysis.
bash
datalad containers-run \
-n bids-mriqc \
--input sourcedata/raw \
--output . \
'{inputs}' '{outputs}' participant group -w workdir
If everything worked as expected, we will now see our new analysis, and a commit message of how it was obtained! All of this is contained within a single (nested) dataset with a complete record of how all the data was obtained.
```shell (git) .../ds000003-qc[master] $ git show --quiet Author: Austin austin@dartmouth.edu Date: Wed Jun 5 15:41:59 2024 -0400
[DATALAD RUNCMD] ./code/containers/scripts/singularity_cm...
=== Do not change lines below ===
{
"chain": [],
"cmd": "./code/containers/scripts/singularity_cmd run code/containers/images/bids/bids-mriqc--0.16.0.sing '{inputs}' '{outputs}' participant group -w workdir",
"dsid": "c9c96ab9-f803-43ba-83e2-2eaec7ab4725",
"exit": 0,
"extra_inputs": [
"code/containers/images/bids/bids-mriqc--0.16.0.sing"
],
"inputs": [
"sourcedata/raw"
],
"outputs": [
"."
],
"pwd": "."
}
^^^ Do not change lines above ^^^
```
This record could later be reused (by anyone) using datalad rerun to rerun this computation using exactly the same version(s) of input data and the singularity container. You can even now datalad uninstall sourcedata/raw and even containers sub-datasets to save space - they will be retrievable at those exact versions later on if you need to extend or redo your analysis.
Notes:
- aforementioned example requires DataLad >= 0.11.5 and datalad-containers >= 0.4.0;
- for more eleborate example with use of reproman to parallelize execution on remote resources, see ReproNim/reproman PR#438;
- a copy of the dataset is made available from
///repronim/ds000003-qcand https://github.com/ReproNim/ds000003-qc. - if you would like to create
licenses/folder in your project datasets to e.g. contain license for freesurfer, then you better add them to git-annex. Following commands provide one way to do it:
```shell mkdir licenses
instruct git-annex to add license files to annex, but this added file with instructions to git
echo -e '* annex.largefiles=anything\n.gitattributes annex.largefiles=nothing' > licenses/.gitattributes datalad save -m "Add licenses must go into git-annex so I could avoid sharing them" licenses/.gitattributes cp ~/.freesurfer-license licenses/freesurfer datalad save -m 'added freesurfer license' licenses/freesurfer ```
Installation
It is a DataLad dataset, so you can either just git clone or datalad install it. You will need to have git-annex available to retrieve any images. And you will need DataLad and datalad-container extension installed for datalad containers-run. Since Singularity is Linux-only application, it will be "functional" only on Linux. On OSX (and possibly Windows), if you have Docker installed, singularity images will be executed through the provided docker shim image.
Environment variables
A few environment variables (in addition to those consulted by datalad and datalad-container) are considered in the scripts of this repository:
SINGULARITY_CMD
The default command (as "hardcoded" in .datalad/config) is run
so running the container executes its default "entry point". Setting
SINGULARITY_CMD=exec makes it possible to run an alternative command
in them (e.g. bash for interactive sessions)::
SINGULARITY_CMD=exec datalad containers-run --explicit -n repronim-reproin bash
and then have datalad record any of the introduced changes. Such
runs will not be reproducible but at least clearly annotated in what
environment corresponding actions were taken.
Acknowledgements
Grants
Development of this project and datalad-container extension was supported by the ReproNim project (NIH 1P41EB019936-01A1). DataLad development was supported by a US-German collaboration in computational neuroscience (CRCNS) "DataGit: converging catalogues, warehouses, and deployment logistics into a federated 'data distribution'" (Halchenko/Hanke), co-funded by the US National Science Foundation (NSF 1429999) and the German Federal Ministry of Education and Research (BMBF 01GQ1411). Additional support is provided by the German federal state of Saxony-Anhalt and the European Regional Development Fund, Project: Center for Behavioral Brain Sciences, Imaging Platform.
Copyrighted works
All container images are collections of various projects governed by the
corresponding copyrights/licenses. Some are not completely FOSS and might
require additional license(s) to be obtained and provided (e.g. FreeSurfer
license for fmriprep).
artwork/repronim-containers-yoda_*
Based on the artwork Copyright 2018-2019 Michael Hanke, from myyoda/poster, distributed under CC BY.
Owner
- Name: Center for Reproducible Neuroimaging Computation
- Login: ReproNim
- Kind: organization
- Website: http://repronim.org
- Repositories: 75
- Profile: https://github.com/ReproNim
Citation (CITATION.cff)
# This CITATION.cff file was generated with cffinit.
# Visit https://bit.ly/cffinit to generate yours today!
cff-version: 1.2.0
title: >-
ReproNim/containers - containerized environments for
reproducible neuroimaging
message: >-
If you use this product, please cite it using the metadata
from this file.
type: dataset
authors:
- orcid: 'https://orcid.org/0000-0003-3456-2493'
affiliation: 'Dartmouth College, Hanover, NH, United States'
given-names: Yaroslav
name-particle: O
family-names: Halchenko
- given-names: Christian
family-names: Haselgrove
affiliation: >-
University of Massachusetts Medical School, Worcester,
MA, United States
orcid: 'https://orcid.org/0000-0002-4438-0637'
- family-names: Travers
given-names: Matt
orcid: 'https://orcid.org/0000-0001-5456-5371'
affiliation: 'TCG, Inc, Washington, DC, United States"'
- given-names: John
family-names: Wodder
name-suffix: II
name-particle: T.
affiliation: 'Dartmouth College, Hanover, NH, United States'
- given-names: Austin
family-names: Macdonald
affiliation: 'Dartmouth College, Hanover, NH, United States'
orcid: 'https://orcid.org/0000-0002-8124-807X'
identifiers:
- type: other
value: 'RRID:SCR_018467'
description: 'https://scicrunch.org/resources'
repository-code: 'https://github.com/ReproNim/containers'
repository: 'https://datasets.datalad.org/?dir=/repronim/containers'
abstract: >-
This repository provides a DataLad dataset (git/git-annex
repository) with a collection of popular computational
tools provided within ready to use containerized
environments.
keywords:
- reproducible research
- datalad
- containers
- singularity
version: 0.20240201.0
GitHub Events
Total
- Create event: 6
- Release event: 1
- Issues event: 6
- Watch event: 2
- Delete event: 6
- Member event: 1
- Issue comment event: 27
- Push event: 140
- Pull request review event: 3
- Pull request event: 17
- Fork event: 1
Last Year
- Create event: 6
- Release event: 1
- Issues event: 6
- Watch event: 2
- Delete event: 6
- Member event: 1
- Issue comment event: 27
- Push event: 140
- Pull request review event: 3
- Pull request event: 17
- Fork event: 1
Issues and Pull Requests
Last synced: 9 months ago
All Time
- Total issues: 83
- Total pull requests: 59
- Average time to close issues: about 2 months
- Average time to close pull requests: 16 days
- Total issue authors: 13
- Total pull request authors: 10
- Average comments per issue: 2.31
- Average comments per pull request: 1.66
- Merged pull requests: 51
- Bot issues: 0
- Bot pull requests: 3
Past Year
- Issues: 5
- Pull requests: 10
- Average time to close issues: 10 days
- Average time to close pull requests: 7 days
- Issue authors: 4
- Pull request authors: 4
- Average comments per issue: 2.2
- Average comments per pull request: 1.0
- Merged pull requests: 7
- Bot issues: 0
- Bot pull requests: 0
Top Authors
Issue Authors
- yarikoptic (58)
- jbwexler (5)
- bpinsard (5)
- asmacdo (4)
- chaselgrove (2)
- dnkennedy (2)
- adswa (1)
- effigies (1)
- dorianps (1)
- jsmentch (1)
- Remi-Gau (1)
- mattcieslak (1)
- DVSneuro (1)
Pull Request Authors
- yarikoptic (29)
- asmacdo (10)
- jwodder (9)
- chaselgrove (7)
- mjtravers (4)
- bpinsard (3)
- dependabot[bot] (3)
- candleindark (2)
- mattcieslak (1)
- adswa (1)
Top Labels
Issue Labels
Pull Request Labels
Dependencies
- actions/checkout v3 composite
- actions/checkout v3 composite
- actions/setup-python v4 composite
- actions/checkout v4 composite
- codespell-project/actions-codespell v2 composite