Science Score: 67.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
✓CITATION.cff file
Found CITATION.cff file -
✓codemeta.json file
Found codemeta.json file -
✓.zenodo.json file
Found .zenodo.json file -
✓DOI references
Found 6 DOI reference(s) in README -
✓Academic publication links
Links to: biorxiv.org -
○Academic email domains
-
○Institutional organization owner
-
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (11.3%) to scientific vocabulary
Repository
Whole Genome Alignment Tools
Basic Info
- Host: GitHub
- Owner: wjwei-handsome
- License: mit
- Language: Rust
- Default Branch: master
- Size: 2.47 MB
Statistics
- Stars: 196
- Watchers: 2
- Forks: 14
- Open Issues: 10
- Releases: 3
Metadata Files
README.md
Whole Genome Alignment Tools

A Rust library and tools for whole genome alignment files
Table of Contents
- Citation
-
- Usage
- Auto-Completion for easy-use
- Format Conversion
- Dotplot for MAF/PAF file
- Extract regions from MAF file
- View MAF file in terminal
- Call Variants from MAF file
- Chunk MAF file by length
- Statistics for MAF/PAF file
- Validate and fix PAF file
- Filter records for MAF/PAF file
- Rename MAF file
- PAF Coverage for all-to-all alignment
- Generate pseudo MAF from all-to-all PAF
Citation
If you use wgatools in your research, please cite:
BibLaTeX
```bibtex @article{weiWgatoolsUltrafastToolkit2025, title = {Wgatools: An Ultrafast Toolkit for Manipulating Whole Genome Alignments}, shorttitle = {Wgatools}, author = {Wei, Wenjie and Gui, Songtao and Yang, Jian and Garrison, Erik and Yan, Jianbing and Liu, Hai-Jun}, date = {2025-03-27}, journaltitle = {Bioinformatics}, shortjournal = {Bioinformatics}, pages = {btaf132}, issn = {1367-4811}, doi = {10.1093/bioinformatics/btaf132}, url = {https://doi.org/10.1093/bioinformatics/btaf132}, urldate = {2025-03-28}, } ```Install
Conda
shell
conda install wgatools -c bioconda
Build from source
shell
git clone https://github.com/wjwei-handsome/wgatools.git
cd wgatools
cargo build --release
or just install from git:
shell
cargo install --git https://github.com/wjwei-handsome/wgatools.git
Nix
A nix flake is also available. You can build from within the repo like this:
shell
nix build .#wgatools
Or directly install from github:
shell
nix profile install github:wjwei-handsome/wgatools
Docker and Singularity
Using nix, we can derive docker and singularity images:
shell
nix build .#dockerImage
First, we load the docker image into the local daemon:
shell
docker load < result
It's then possible to pack up a singularity image:
shell
singularity build wgatools-$(git log -1 --format=%h --abbrev=8).sif docker-daemon://wgatools:latest
This can be useful when running on HPCs where it might be difficult to build wgatools.
Guix
Clone wgatools repo and create a Guix shell to build and hack on wgatools:
cd wgatools
guix shell --share=/home/wrk/.cargo -C -D -N rust rust-cargo openssl nss-certs nss coreutils-minimal which perl make binutils gcc-toolchain pkg-config cmake zlib
env LD_LIBRARY_PATH=$GUIX_ENVIRONMENT/lib CC=gcc cargo build
The --share switch prevents redownloading cargo packages. -C -D defines a build container that is independent of the underlying
Linux distribution. -N gives network access to the guix shell and -F provides the Linux file system hierarchy standard (FHS).
Note that cargo build --release build does not work (yet). There is some problem with cargo+ctest+static builds.
Tools
Usage
```shell
wgatools wgatools -- a cross-platform and ultrafast toolkit for Whole Genome Alignment Files manipulation
Version: 0.1.0
Authors: Wenjie Wei wjwei9908@gmail.com
Usage: wgatools [OPTIONS]
Commands: maf2paf Convert MAF format to PAF format [aliases: m2p] maf2chain Convert MAF format to Chain format [aliases: m2c] paf2maf Convert PAF format to MAF format [aliases: p2m] paf2chain Convert PAF format to Chain format [aliases: p2c] chain2maf Convert Chain format to MAF format [aliases: c2m] chain2paf Convert Chain format to PAF format [aliases: c2p] maf-index Build index for MAF file [aliases: mi] maf-ext Extract specific region from MAF file with index [aliases: me] chunk Chunk MAF file by length [aliases: ch] call Call Variants from MAF file [aliases: c] tview View MAF file in terminal [aliases: tv] stat Statistics for Alignment file [aliases: st] dotplot Plot dotplot for Alignment file [aliases: dp] filter Filter records for Alignment file [aliases: fl] rename Rename MAF records with prefix [aliases: rn] maf2sam DEV: maf2sam [aliases: m2s] pafcov Calculate coverage for PAF file [aliases: pc] pafpseudo Generate pesudo-maf for divergence analysis from PAF file [aliases: pp] gen-completion Generate completion script for shell [aliases: gc] validate Validate and fix query&target position in PAF file by CIGAR [aliases: vf] help Print this message or the help of the given subcommand(s)
Options: -h, --help Print help (see more with '--help') -V, --version Print version
GLOBAL:
-o, --outfile
Each subcommand could be used with -h or --help to get more information.
Auto-Completion for easy-use
shell
wgatools gen-completion --shell fish > ~/.config/fish/completions/wgatools.fish
Ready to enjoy it!
Format Conversion
Three mainstream formats(PAF, MAF, CHAIN) can be converted to each other.
For example, to convert MAF to PAF:
shell
wgatools maf2paf test.maf > test.paf
or to convert PAF to MAF:
shell
wgatools paf2maf test.paf --target target.fa --query query.fa > test.maf
[!TIP] If you want to convert into MAF format, you should provide target and query genome sequence files in {.fa, .fa.gz}.
stdin and stdout are supported, so you can use pipes to chain commands together🪆:
```shell cat test.maf | wgatools maf2paf | wgatools paf2maf -g target.fa -q query.fa > test.maf
wgatools paf2chain test.paf | wgatools chain2maf -g target.fa -q query.fa | wgatools maf2chain | wgatools chain2paf > funny.paf ```
Dotplot for MAF/PAF file
We provide two modes for plot, for example:
- BaseLevel
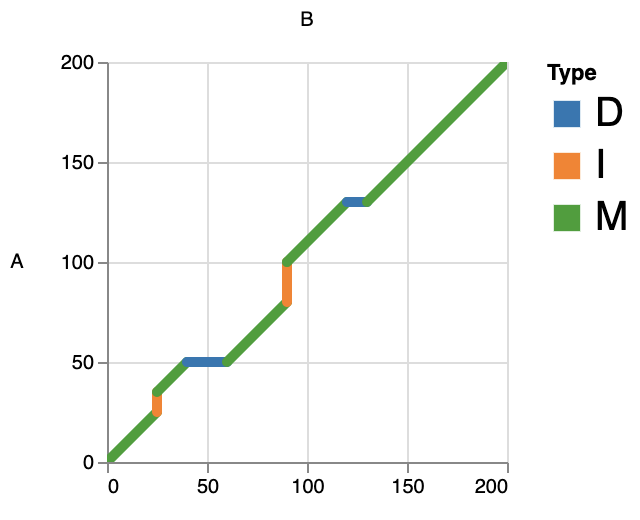
This mode can catch the alignment details in each record, such as matches, insertions and deletions. This can help us to better observe the local alignment.
shell
wgatools dotplot -f paf test/testdotplot.paf > out.html
By default, INDELs smaller than 50bp are merged with adjacent match. You can also use the parameter -l, --length to specify the threshold.
In Interactive html, you can click on the legend to view only the types of interest, for example:
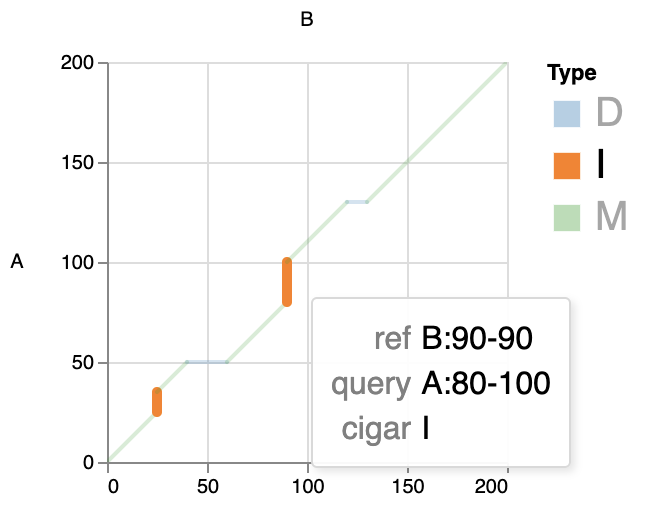
[!WARNING] NOTE: For better interactivity, the
zoomfunction is turned on. However, if there is too much data, the effect may be limited by your browser performance. For Better performance, you can collapse short segments by-lparameter.
This simple example can be found in the test directory.
- Overview
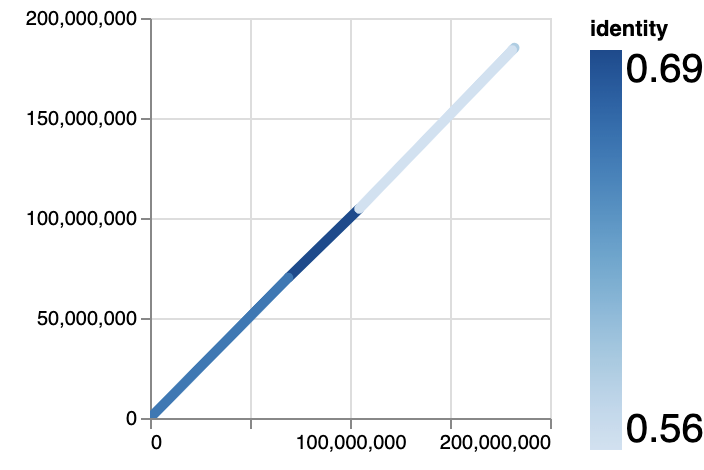
Similar to common dotplot scripts, it will draw each align record and color it according to identity.
shell
wgatools dotplot test.maf -m overview > overview.html
😎 For vega and DIY hackers, we also provide output in json(vega schema) and csv formats.
Official tool vl-converter can convert json files to multiple formats.
Extract regions from MAF file
The line of MAF file is so long that it's hard to read. You can use maf-ext to extract specific region from MAF file with index:
```shell wgatools maf-index test.maf
wgatools maf-extract test.maf -r chr1:1-10,chr2:66-888,chr3:100-50,chr_no:1-10,x:y-z ```
[!TIP] 1. Support multi-interval input, separated by commas 2. Support
bedinput to specify interval 3. Mismatched interval are skipped and warned
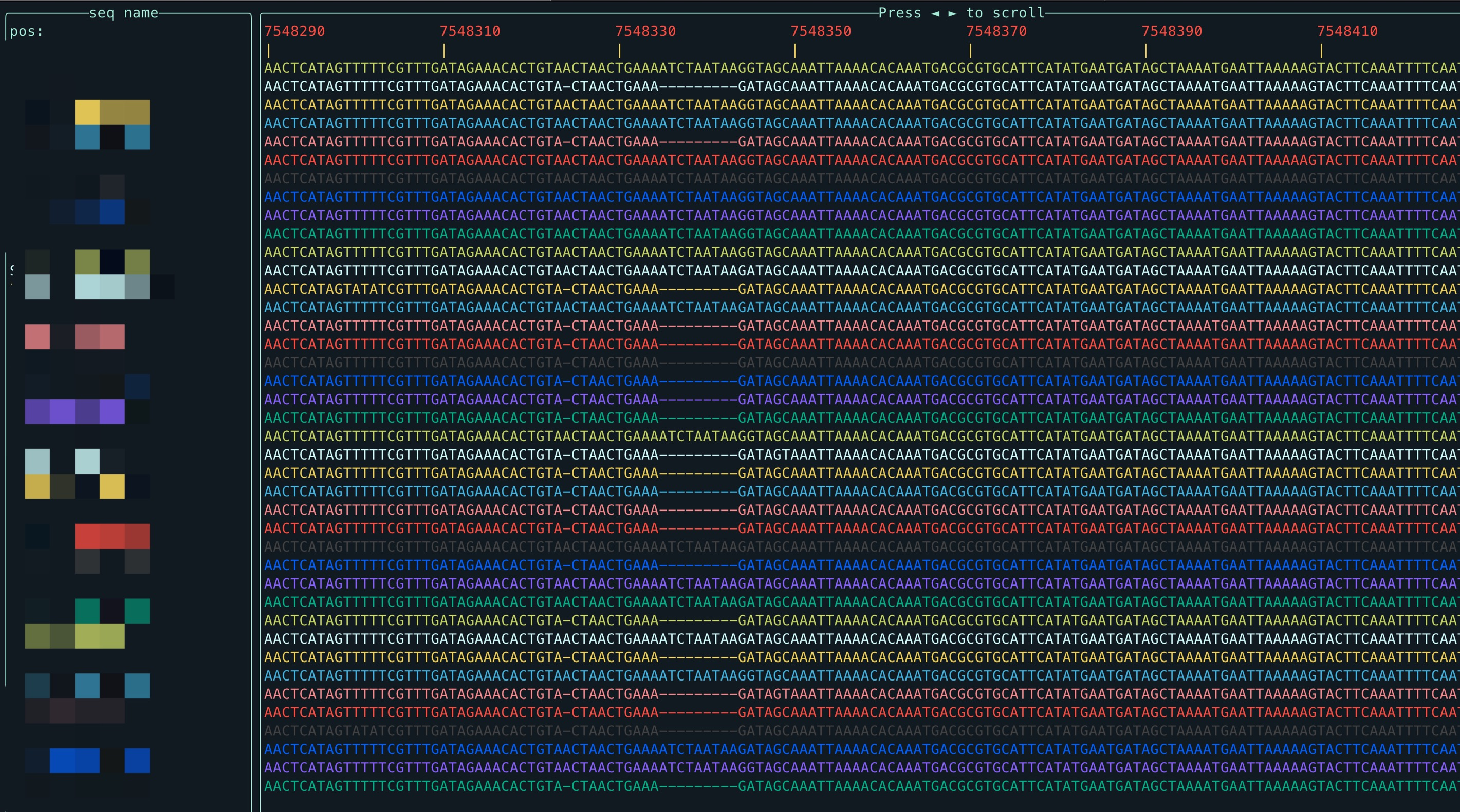
View MAF file in terminal
View the MAF file in the terminal smoothly, and you can also specify the area to view:
shell
wgatools tview test.maf

Press ◄► to slide left and right.
Press q to exit.
Press g to bring up the navigation window, where the left side is the optional sequence name, and the right side is the optional interval of the selected sequence, you can press Tab to switch the left and right selection windows, and you can press ▲▼ to select the sequence and interval
After input a legal interval, you can Press Enter to jump to the Destination. Or press Esc to exit the navigation window.
Call Variants from MAF file
The MAF format completely records the alignment of each base, so it can be used to identify variants.
Supported explicit varaint types: - SNP - INS - DEL - INV
The default parameter does not output SNP and short INS and DEL (<50). The example is as follows:
shell
wgatools call test/test.maf -s -l0
or directly use PAF file with target and query sequence:
shell
wgatools call test/test.paf -s -l0 --target target.fa --query query.fa -f paf
Output vcf: ```
fileformat=VCFv4.4
INFO=
INFO=
INFO=
INFO=
FORMAT=
FORMAT=
CHROM POS ID REF ALT QUAL FILTER INFO FORMAT sample
ref.chr8 181470034 . TG T . . SVTYPE=DEL;SVLEN=1;END=181470035 GT:QI 1|1:query.chr8@181989530@181989530@P ref.chr8 181470279 . G C . . . GT 1|1 ref.chr8 181470292 . A G . . . GT 1|1 ref.chr8 181470431 . C G . . . GT 1|1 ref.chr8 181470609 . C A . . . GT 1|1 ref.chr8 181470641 . C T . . . GT 1|1 ref.chr8 181470774 . A AAACCAAGA . . SVTYPE=INS;SVLEN=8;END=181470774 GT:QI 1|1:query.chr8@181990269@181990277@P ref.chr8 181470793 . G T . . . GT 1|1 ref.chr8 181470894 . C T . . . GT 1|1 ref.chr8 181470895 . A T . . . GT 1|1 ref.chr8 181470903 . G A . . . GT 1|1 ```
[!IMPORTANT] This function does not support the identification of chromosomal rearrangements such as
DUP, as this requires the extraction of sequences for realignment.
Chunk MAF file by length
You can split a huge MAF record into multiple records by length:
shell
wgatools chunk -l 100 test/test.maf -o chunked.maf
Statistics for MAF/PAF file
shell
wgatools stat test.maf
wgatools stat -f paf test.paf
wgatools stat test.maf
Validate and fix PAF file
In some cases, the PAF file may be incorrect, such as the query and target postions are wrong, or CIGAR string is unmatch with sequences. You can use this command to validate and fix the PAF file:
```shell
just validate
wgatools validate wrong.paf
Total records: 2306 Query invalid records: 2283 Target invalid records: 80 Query invalid list:... Target invalid list:...
validate and fix
wgatools validate wrong.paf -f happy.paf
```
Filter records for MAF/PAF file
You can filter some records by block length or query_size.
For example, to filter records that contig vs reference:
shell
wgatools filter test.maf -q 1000000 > filt.maf
For all-to-all alignment paf file which produced by wfmash, you can filter some pairs by align-size:
shell
wgatools filter all2all.paf -a 1000000 > filt.maf
Rename MAF file
In some practices, the chromosome name of ref and query are both called chr1, which is not easy to distinguish.
You can rename the sequence name in MAF file with a prefix:
shell
wgatools rename --prefixs REF.,QUERY. input.maf > rename.maf
PAF Coverage for all-to-all alignment
If you have alignment results for multiple genomes, you can use this command to calculate the alignment coverage on the genomes. It's optimized to use with wfmash output.
shell
wgatools pafcov all.paf > all.cov.beds
Generate pseudo MAF from all-to-all PAF
shell
wgatools pafpseudo -f all.fa.gz all.paf -o out_dir -t 10
[!TIP] Practical processes and profile can refer to this pipleline and this paper
Library
Some simple reader and iterator for PAF, MAF and Chain files:
rust
use wgatools::parser::paf::PafReader;
use wgatools::parser::maf::MAFReader;
use wgatools::parser::chain::ChainReader;
fn main() {
let mut mafreader = MAFReader::from_path("test.maf").unwrap();
for record in mafreader.records() {
let record = record.unwrap();
println!("{:?}", record);
}
/// ...
}
Features
- use
nomto parse CIGAR string - use
rayonto accelerate the speed of conversions - use
ratatuito visualize MAF file in terminal - ...
Benchmark
We use the hyperfine to compare the speed of conversion between wgatools and another Rust-based tool paf2chain. The result is as follows (10 runs):
command|mean(sec)|stddev|median|user|system|min|max :---------------------------------------------------------------------------------------------------------|:-----------------|:-----------------|:-----------------|:----------------|:-----------------|:-----------------|:------------- wgatools p2c Zm-CML333.paf -o foo|3.69|0.36|3.71|3.46|0.14|3.25|4.09 paf2chain --input Zm-CML333.paf > bar|16.28|0.86|16.27|3.80|12.03|15.01|17.67
ROADMAP
- [ ] SAM converter
- [ ] Local improvement of alignment by re-alignment
- [ ] MAF -> GAF -> HAL
- [ ] output gvcf for variants
- [X] call variants from PAF directly
Contributing
Feel free to dive in! Open an issue or submit PRs.
License
MIT License © WenjieWei
Owner
- Name: WeiWenjie
- Login: wjwei-handsome
- Kind: user
- Website: https://www.wjwei.blog
- Repositories: 3
- Profile: https://github.com/wjwei-handsome
Citation (CITATION.bib)
@misc{weiWgatoolsUltrafastToolkit2024,
title = {wgatools: an ultrafast toolkit for manipulating whole genome alignments},
shorttitle = {wgatools},
url = {http://arxiv.org/abs/2409.08569},
doi = {10.48550/arXiv.2409.08569},
abstract = {Summary: With the rapid development of long-read sequencing technologies, the era of individual complete genomes is approaching. We have developed wgatools, a cross-platform, ultrafast toolkit that supports a range of whole genome alignment (WGA) formats, offering practical tools for conversion, processing, statistical evaluation, and visualization of alignments, thereby facilitating population-level genome analysis and advancing functional and evolutionary genomics. Availability and Implementation: wgatools supports diverse formats and can process, filter, and statistically evaluate alignments, perform alignment-based variant calling, and visualize alignments both locally and genome-wide. Built with Rust for efficiency and safe memory usage, it ensures fast performance and can handle large datasets consisting of hundreds of genomes. wgatools is published as free software under the MIT open-source license, and its source code is freely available at https://github.com/wjwei-handsome/wgatools. Contact: weiwenjie@westlake.edu.cn (W.W.) or liuhaijun@yzwlab.cn (H.-J.L.).},
urldate = {2024-09-17},
publisher = {arXiv},
author = {Wei, Wenjie and Gui, Songtao and Yang, Jian and Garrison, Erik and Yan, Jianbing and Liu, Hai-Jun},
month = sep,
year = {2024},
note = {arXiv:2409.08569 [q-bio]},
keywords = {Quantitative Biology - Genomics},
file = {2024_Wei et al_wgatools - an ultrafast toolkit for manipulating whole genome alignments_.pdf:/Users/wjwei/Papers/2024_Wei et al_wgatools - an ultrafast toolkit for manipulating whole genome alignments_.pdf:application/pdf;arXiv.org Snapshot:/Users/wjwei/Zotero/storage/UWYCQI62/2409.html:text/html},
}
GitHub Events
Total
- Create event: 2
- Release event: 2
- Issues event: 14
- Watch event: 85
- Issue comment event: 22
- Push event: 12
- Pull request event: 2
- Fork event: 7
Last Year
- Create event: 2
- Release event: 2
- Issues event: 14
- Watch event: 85
- Issue comment event: 22
- Push event: 12
- Pull request event: 2
- Fork event: 7
Issues and Pull Requests
Last synced: 6 months ago
All Time
- Total issues: 7
- Total pull requests: 1
- Average time to close issues: 2 months
- Average time to close pull requests: about 3 hours
- Total issue authors: 7
- Total pull request authors: 1
- Average comments per issue: 1.43
- Average comments per pull request: 1.0
- Merged pull requests: 1
- Bot issues: 0
- Bot pull requests: 0
Past Year
- Issues: 7
- Pull requests: 1
- Average time to close issues: 2 months
- Average time to close pull requests: about 3 hours
- Issue authors: 7
- Pull request authors: 1
- Average comments per issue: 1.43
- Average comments per pull request: 1.0
- Merged pull requests: 1
- Bot issues: 0
- Bot pull requests: 0
Top Authors
Issue Authors
- baozg (4)
- sdws1983 (1)
- dingyigithub (1)
- ZhaoxuMa (1)
- unavailable-2374 (1)
- Miles-TANGsk (1)
- sivico26 (1)
- socialhang (1)
- sip123a (1)
- rejo27 (1)
- nuriaher (1)
- virag-compbio (1)
Pull Request Authors
- AndreaGuarracino (2)
- sharkLoc (2)
- microfuge (1)
- pjotrp (1)
- ekg (1)

