Science Score: 57.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
✓CITATION.cff file
Found CITATION.cff file -
✓codemeta.json file
Found codemeta.json file -
✓.zenodo.json file
Found .zenodo.json file -
✓DOI references
Found 3 DOI reference(s) in README -
○Academic publication links
-
○Academic email domains
-
○Institutional organization owner
-
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (17.6%) to scientific vocabulary
Keywords
Repository
A kinetic model of CRISPR-Cas target recognition.
Basic Info
- Host: GitHub
- Owner: hiddeoff
- License: mit
- Language: Python
- Default Branch: main
- Homepage: https://hiddeoff.github.io/crisprzip/
- Size: 8.97 MB
Statistics
- Stars: 2
- Watchers: 2
- Forks: 2
- Open Issues: 0
- Releases: 6
Topics
Metadata Files
README.md
CRISPRzip
Welcome to the codebase of CRISPRzip from the Depken Lab at TU Delft.
About the project
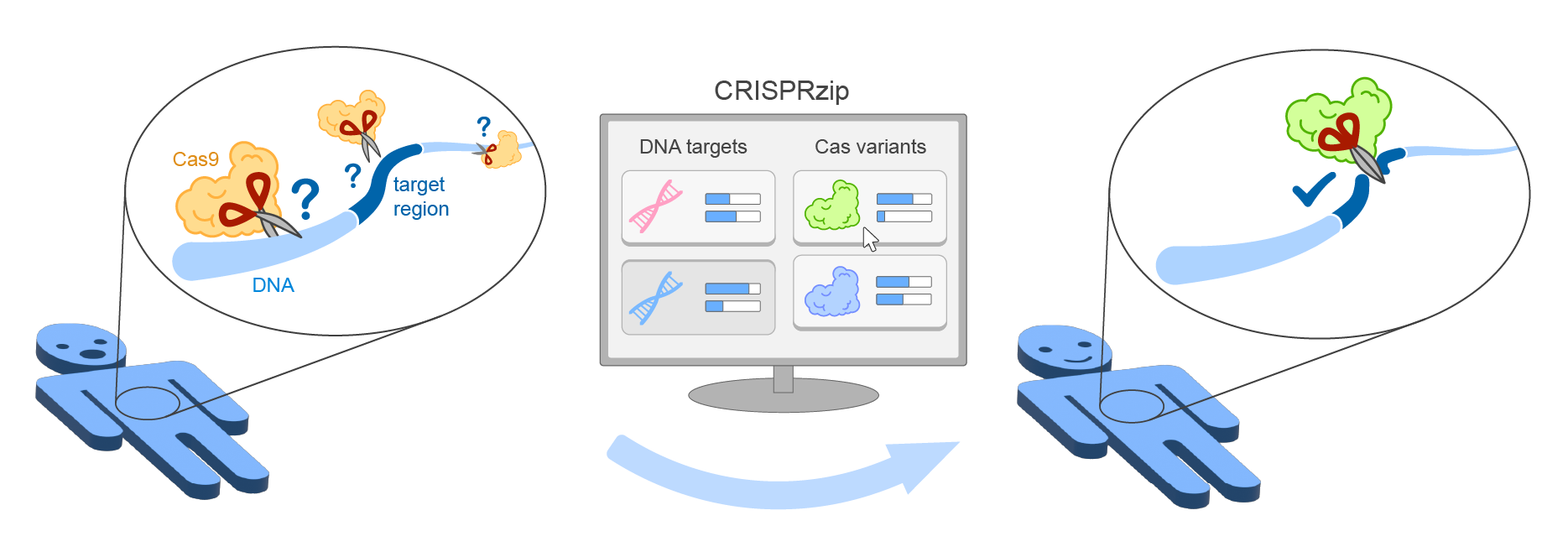
CRISPRzip is a physics-based model to study the target recognition dynamics of CRISPR-associated nucleases like Cas9 (Eslami-Mossalam, 2022). Their interactions with target DNA is represented as an energy landscape, with which you can simulate binding and cleavage kinetics. The parameters have been obtained by machine learning on high-throughput data. CRISPRzip makes quantitative predictions of on-target efficiency and off-target risks of different guide RNAs.
With CRISPRzip, we hope to contribute to assessing the risks that come with particular choices in CRISPR application, and as such contribute to the development of safe gene editing technology.
References
Eslami-Mossallam B et al. (2022) A kinetic model predicts SpCas9 activity, improves off-target classification, and reveals the physical basis of targeting fidelity. Nature Communications. 10.1038/s41467-022-28994-2
Installation
CRISPRzip is available on PyPi and can be installed with pip.
User installation
Although CRISPRzip can be directly installed from pip, creating a virtual environment makes it easier to manage dependencies. Below you can see some intructions on how to generate a virtual environment with venv assuming you already have python installed in a bash-like terminal.
shell
python -m venv crisprzip-venv
source crisprzip-venv/bin/activate
pip install crisprzip
Developer installation
To be able to make changes and contributions to CRISPRzip, you will need to get your own copy of the source code and install software dependencies on your own. Assuming you have a python and git installed in a bash-like terminal, the installation process can be done with the following instructions.
shell
git clone https://github.com/hiddeoff/crisprzip.git
cd crisprzip
python -m venv crisprzip-venv
source crisprzip-venv/bin/activate
pip install -e .
Please use our contributing guidelines if you would like us to consider your developments in a future CRISPRzip release
Verifying installation
CRISPRzip development includes a cross-platform compatibility and test workflow. If you would like to verify your local installation, please follow the Developer Installation instructions, then run the following test.
shell
source crisprzip-venv/bin/activate
pip install -e '.[tests]' # installs pytest and pandas
pytest tests/cleavage_binding_prediction/test_cleavage_binding_prediction.py -v
You can also follow the user installation and execute the code in the Usage section below.
Usage
CRISPRzip makes predictions about cleavage and binding activity on on- and off-targets. First, you define the protospacer and target sequence, and then, you can predict the fraction cleaved or bound.
```python
1. load parameter set
from crisprzip.kinetics import loadlandscape searcher = loadlandscape("sequence_params")
2. define Cas9, gRNA and DNA target
searchertargetcomplex = searcher.probesequence( protospacer = "AGACGCATAAAGATGAGACGCTGG", targetseq = "AGACCCATTAAGATGAGACGCGGG", # A13T G17C )
3. predict activity
fclv = searchertargetcomplex.getcleavedfraction( time=600, # 10 minutes onrate=1E-1 ) fbnd = searchertargetcomplex.getboundfraction( time=600, # 10 minutes onrate=1E-1 )
4. format output
print(f"After 10 minutes, the target (A13T G17C) is ...")
print(f"- cleaved for {100 * fclv:.1f}% by Cas9")
print(f" or ")
print(f"- bound for {100 * fbnd:.1f}% by dCas9")
Output:
After 10 minutes, the target (A13T G17C) is ...
- cleaved for 10.5% by Cas9
or
- bound for 94.2% by dCas9
```
See the tutorial or the docs for more examples how to explore sequence, time and concentration dependency.
Contributing
We encourage contributions in any form - reporting bugs, suggesting features, drafting code changes. Read our Contributing guidelines and our Code of Conduct.
Acknowledgements
Many thanks to Elviss Dvinskis, Raúl Ortiz and Aysun Urhan from the DCC team at TU Delft for their support to get this package released!
Waiver
Technische Universiteit Delft hereby disclaims all copyright interest in the program “CRISPRzip” (a physics-based CRISPR activity predictor) written by the Author(s). Paulien Herder, Dean of Applied Sciences
(c) 2024, Hidde Offerhaus, Delft, The Netherlands.
Owner
- Login: hiddeoff
- Kind: user
- Repositories: 2
- Profile: https://github.com/hiddeoff
Citation (CITATION.cff)
cff-version: 1.2.0 message: "If you are using this software, please cite it as shown below." authors: - family-names: "Offerhaus" given-names: "Hidde" title: "CRISPRzip" version: 1.0.0 date-released: 2025-02-04 license: MIT url: "https://github.com/hiddeoff/crisprzip-model"
GitHub Events
Total
- Create event: 7
- Release event: 6
- Issues event: 10
- Delete event: 5
- Issue comment event: 9
- Public event: 1
- Push event: 55
- Pull request event: 25
- Fork event: 2
Last Year
- Create event: 7
- Release event: 6
- Issues event: 10
- Delete event: 5
- Issue comment event: 9
- Public event: 1
- Push event: 55
- Pull request event: 25
- Fork event: 2
Issues and Pull Requests
Last synced: 9 months ago
All Time
- Total issues: 8
- Total pull requests: 16
- Average time to close issues: about 2 months
- Average time to close pull requests: 5 days
- Total issue authors: 3
- Total pull request authors: 3
- Average comments per issue: 2.13
- Average comments per pull request: 0.13
- Merged pull requests: 13
- Bot issues: 0
- Bot pull requests: 0
Past Year
- Issues: 8
- Pull requests: 16
- Average time to close issues: about 2 months
- Average time to close pull requests: 5 days
- Issue authors: 3
- Pull request authors: 3
- Average comments per issue: 2.13
- Average comments per pull request: 0.13
- Merged pull requests: 13
- Bot issues: 0
- Bot pull requests: 0
Top Authors
Issue Authors
- edvinskis (5)
- rortizmerino (2)
- hiddeoff (1)
Pull Request Authors
- hiddeoff (12)
- rortizmerino (2)
- edvinskis (2)
Top Labels
Issue Labels
Pull Request Labels
Packages
- Total packages: 1
-
Total downloads:
- pypi 83 last-month
- Total dependent packages: 0
- Total dependent repositories: 0
- Total versions: 7
- Total maintainers: 1
pypi.org: crisprzip
A kinetic model of CRISPR-Cas target recognition.
- Homepage: https://github.com/hiddeoff/crisprzip
- Documentation: https://hiddeoff.github.io/crisprzip/
- License: MIT License Copyright (c) 2024 Hidde Offerhaus Permission is hereby granted, free of charge, to any person obtaining a copy of this software and associated documentation files (the "Software"), to deal in the Software without restriction, including without limitation the rights to use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of the Software, and to permit persons to whom the Software is furnished to do so, subject to the following conditions: The above copyright notice and this permission notice shall be included in all copies or substantial portions of the Software. THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE.
-
Latest release: 1.2.2
published 10 months ago
Rankings
Maintainers (1)
Dependencies
- actions/checkout v3 composite
- actions/setup-python v4 composite
- peaceiris/actions-gh-pages v3 composite
- joblib >=1.1
- matplotlib >=3.7
- numba >=0.57
- numpy >=1.24
- scipy >=1.7

