charlie
Circrnas in Host And viRuses anaLysis pIpEline for Detection Annotation Quantification of circRNAs
Science Score: 49.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
○CITATION.cff file
-
✓codemeta.json file
Found codemeta.json file -
✓.zenodo.json file
Found .zenodo.json file -
✓DOI references
Found 3 DOI reference(s) in README -
✓Academic publication links
Links to: zenodo.org -
○Academic email domains
-
○Institutional organization owner
-
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (9.8%) to scientific vocabulary
Keywords
Repository
Circrnas in Host And viRuses anaLysis pIpEline for Detection Annotation Quantification of circRNAs
Basic Info
- Host: GitHub
- Owner: CCBR
- License: mit
- Language: Python
- Default Branch: main
- Homepage: https://ccbr.github.io/CHARLIE/
- Size: 278 MB
Statistics
- Stars: 2
- Watchers: 2
- Forks: 2
- Open Issues: 12
- Releases: 29
Topics
Metadata Files
README.md
CHARLIE
Circrnas in Host And viRuses anaLysis pIpEline
See the website for detailed information, documentation, and examples: https://ccbr.github.io/CHARLIE/
Table of Contents
1. Introduction
Circrnas in Host And viRuses anaLysis pIpEline
Things to know about CHARLIE:
- Snakemake workflow to detect, annotate and quantify (DAQ) host and viral circular RNAs.
- Primirarily developed to run on BIOWULF
- Reach out to Vishal Koparde for questions/comments/requests.
This circularRNA detection pipeline uses CIRCExplorer2, CIRI2 and many other tools in parallel to detect, quantify and annotate circRNAs. Here is a list of tools that can be run using CHARLIE:
| circRNA Detection Tool | Aligner(s) | Run by default | | ----------------------------------------------------------- | ---------------- | -------------- | | CIRCExplorer2 | STAR1 | Yes | | CIRI2 | BWA1 | Yes | | CIRCExplorer2 | BWA1 | Yes | | CLEAR | STAR1 | Yes | | DCC | STAR2 | Yes | | circRNAFinder | STAR3 | Yes | | find_circ | Bowtie2 | Yes | | MapSplice | BWA2 | No | | NCLScan | NovoAlign | No |
Note: STAR1, STAR2, STAR3 denote 3 different sets of alignment parameters, etc.
Note: BWA1, BWA2 denote 2 different alignment parameters, etc.
2. Flowchart
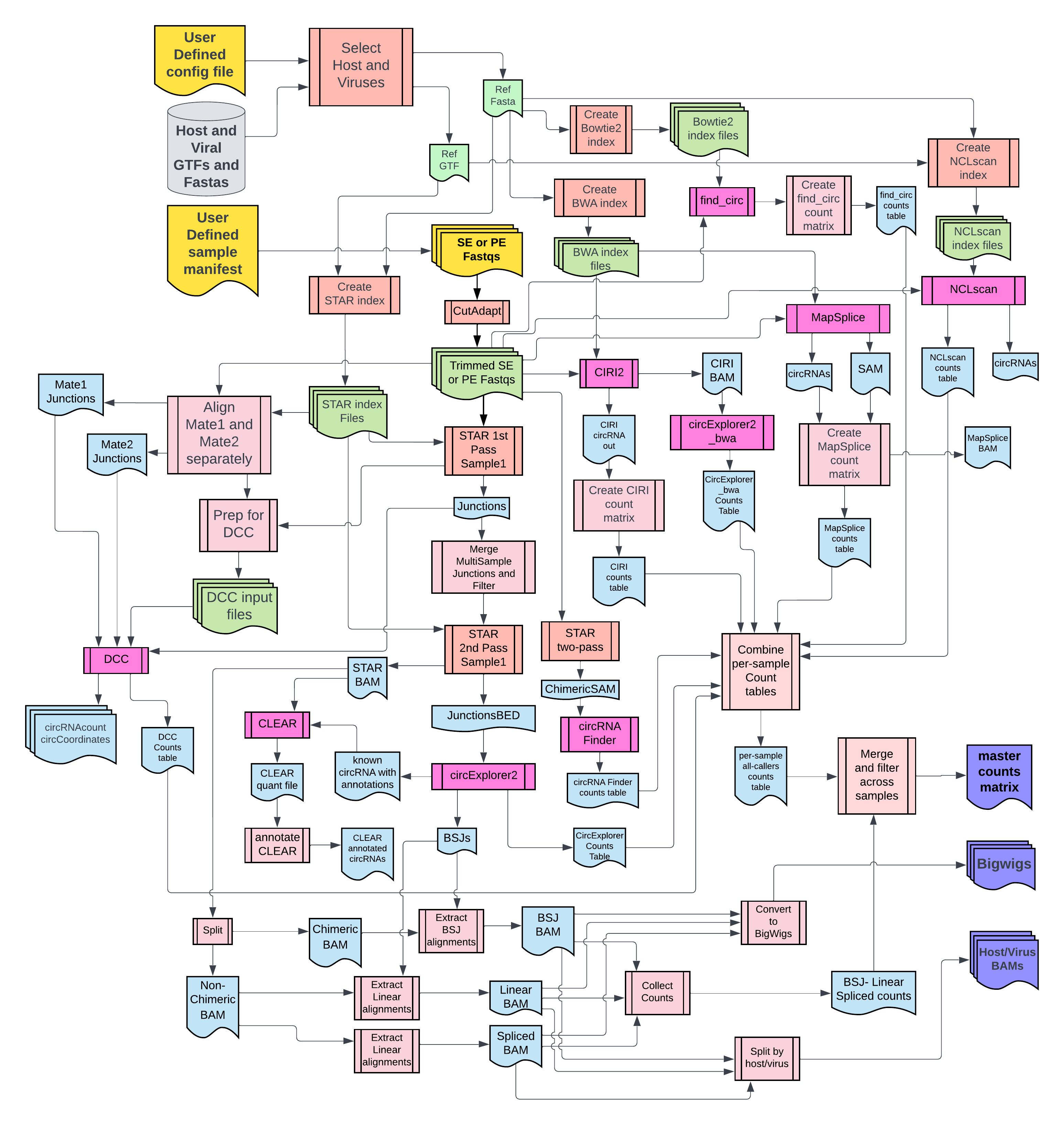
For complete documentation, view the website https://CCBR.github.io/CHARLIE/.
DISCLAIMER: New circRNA tools have been added CHARLIE and the documentation is currently out of date!
3. Software Dependencies
CHARLIE is already installed on biowulf. It is included in the ccbrpipeliner module from release 7 onward. To load the module run:
bash
module load ccbrpipeliner/7
The following version of various bioinformatics tools are using within CHARLIE:
| tool | version | | ------------- | -------- | | blat | 3.5 | | bedtools | 2.30.0 | | bowtie | 2-2.5.1 | | bowtie | 1.3.1 | | bwa | 0.7.17 | | circexplorer2 | 2.3.8 | | cufflinks | 2.2.1 | | cutadapt | 4.4 | | fastqc | 0.11.9 | | hisat | 2.2.2.1 | | java | 18.0.1.1 | | multiqc | 1.9 | | parallel | 20231122 | | perl | 5.34 | | picard | 2.27.3 | | python | 2.7 | | python | 3.8 | | sambamba | 0.8.2 | | samtools | 1.16.1 | | STAR | 2.7.6a | | stringtie | 2.2.1 | | ucsc | 450 | | R | 4.0.5 | | novocraft | 4.03.05 |
4. Usage
```bash charlie
Welcome to
| || | | || _ || _ | | | | | | | | || || || || || | || | | | | | | | || || || |||_ | | | | | |_ | || || || _ || |___ | | | | | | | _ || _ || | | || || | | |_ |______||| |||| |||| |_||__||| |_______|
Circrnas in Host And viRuses anaLysis pIpE_line
This pipeline was built by CCBR (https://bioinformatics.ccr.cancer.gov/ccbr) Please contact Vishal Koparde for comments/questions (vishal.koparde@nih.gov)
CHARLIE can be used to DAQ(Detect/Annotate/Quantify) circRNAs in hosts and viruses.
Here is the list of hosts and viruses that are currently supported:
HOSTS: * hg38 [Human] * mm39 [Mouse]
ADDITIVES: * ERCC [External RNA Control Consortium sequences] * BAC16Insert [insert from rKSHV.219-derived BAC clone of the full-length KSHV genome]
VIRUSES: * NC007605.1 [Human gammaherpesvirus 4 (Epstein-Barr virus)] * NC006273.2 [Human betaherpesvirus 5 (Cytomegalovirus )] * NC001664.4 [Human betaherpesvirus 6A (HHV-6A)] * NC000898.1 [Human betaherpesvirus 6B (HHV-6B)] * NC001716.2 [Human betaherpesvirus 7 (HHV-7)] * NC009333.1 [Human gammaherpesvirus 8 (KSHV)] * NC045512.2 [Severe acute respiratory syndrome(SARS)-related coronavirus] * MN485971.1 [HIV from Belgium] * NC001806.2 Human alphaherpesvirus 1 (Herpes simplex virus type 1) (HSV-1)] * KT899744.1 [HSV-1 strain KOS] * MH636806.1 [MHV68 (Murine herpesvirus 68 strain WUMS)]
USAGE:
charlie -w/--workdir=
Required Arguments: 1. WORKDIR : [Type: String]: Absolute or relative path to the output folder with write permissions.
- RUNMODE : [Type: String] Valid options:
- init : initialize workdir
- dryrun : dry run snakemake to generate DAG
- run : run with slurm
- runlocal : run without submitting to sbatch ADVANCED RUNMODES (use with caution!!)
- unlock : unlock WORKDIR if locked by snakemake NEVER UNLOCK WORKDIR WHERE PIPELINE IS CURRENTLY RUNNING!
- reconfig : recreate config file in WORKDIR (debugging option) EDITS TO config.yaml WILL BE LOST!
- reset : DELETE workdir dir and re-init it (debugging option) EDITS TO ALL FILES IN WORKDIR WILL BE LOST!
- printbinds: print singularity binds (paths)
- local : same as runlocal
Optional Arguments:
--singcache|-c : singularity cache directory. Default is /data/${USER}/.singularity if available, or falls back to ${WORKDIR}/.singularity. Use this flag to specify a different singularity cache directory.
--host|-g : supply host at command line. hg38 or mm39. (--runmode=init only)
--additives|-a : supply comma-separated list of additives at command line. ERCC or BAC16Insert or both (--runmode=init only)
--viruses|-v : supply comma-separated list of viruses at command line (--runmode=init only)
--manifest|-s : absolute path to samples.tsv. This will be copied to output folder (--runmode=init only)
--changegrp|-z : change group to "Ziegelbauer_lab" before running anything. Biowulf-only. Useful for correctly setting permissions.
--help|-h : print this help
Example commands: charlie -w=/my/output/folder -m=init charlie -w=/my/output/folder -m=dryrun charlie -w=/my/output/folder -m=run
VersionInfo: python : 3 snakemake : 7 pipelinehome : /gpfs/gsfs10/users/CCBRPipeliner/Pipelines/CHARLIE/.v0.11.1 git commit/tag : 613fb617f1ed426fb8900f98e599ca0497a67cc0 v0.11.0-49-g613fb61
```
5. License
MIT License
Copyright (c) 2021 Vishal Koparde
Permission is hereby granted, free of charge, to any person obtaining a copy of this software and associated documentation files (the "Software"), to deal in the Software without restriction, including without limitation the rights to use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of the Software, and to permit persons to whom the Software is furnished to do so, subject to the following conditions:
The above copyright notice and this permission notice shall be included in all copies or substantial portions of the Software.
THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE SOFTWARE.
6. Testing
Init
Run init mode:
bash
bash <path to charlie> -w=<path to output dir> -m=init
This will create the folder provided by -w=. The user should have write permission to this folder.
Dry-run
Test data (1 paired-end subsample and 1 single-end subsample) have been including under the /data/CCBR_Pipeliner/testdata/circRNA/human folder. After running in -m=init, samples.tsv should be edited to point the copies of the above mentioned samples with the column headers:
- sampleName
- pathtoR1_fastq
- pathtoR2_fastq
Column path_to_R2_fastq will be blank in case of single-end samples.
After editing samples.tsv, dry run should be run:
bash
bash <path to charlie> -w=<path to output dir> -m=dryrun
This will create the reference fasta and gtf file based on the selections made in the config.yaml.
Run
If -m=dryrun was successful, then simply do -m=run. The output will look something like this
``` ... ... skipping ~1000 lines ... ... Job stats: job count min threads max threads
all 1 1 1 annotateclearoutput 2 1 1 circExplorer 2 2 2 circExplorerbwa 2 2 2 circrnafinder 2 1 1 ciri 2 56 56 clear 2 2 2 createbowtie2index 1 1 1 createbwaindex 1 1 1 createcircExplorerBSJbam 2 4 4 createcircExplorerlinearsplicedbams 2 56 56 createcircExplorermergedfoundcountstable 2 1 1 createhqbams 2 1 1 createindex 1 56 56 createmastercountsfile 1 1 1 cutadapt 2 56 56 dcc 2 4 4 dcccreatesamplesheets 2 1 1 estimateduplication 2 1 1 fastqc 2 4 4 findcirc 2 56 56 findcircalign 2 56 56 mergeSJtabs 1 2 2 mergealignmentstats 1 1 1 mergegenecounts 1 1 1 mergepersample 2 1 1 star1p 2 56 56 star2p 2 56 56 star_circrnafinder 2 56 56 total 52 1 56
Reasons: (check individual jobs above for details) input files updated by another job: alignmentstats, all, annotateclearoutput, circExplorer, circExplorerbwa, circrnafinder, ciri, clear, createcircExplorerBSJbam, createcircExplorerlinearsplicedbams, createcircExplorermergedfoundcountstable, createhqbams, createmastercountsfile, dcc, dcccreatesamplesheets, estimateduplication, fastqc, findcirc, findcircalign, mergeSJtabs, mergealignmentstats, mergegenecounts, mergepersample, star1p, star2p, starcircrnafinder missing output files: alignmentstats, annotateclearoutput, circExplorer, circExplorerbwa, circrnafinder, ciri, clear, createbowtie2index, createbwaindex, createcircExplorerBSJbam, createcircExplorerlinearsplicedbams, createcircExplorermergedfoundcountstable, createhqbams, createindex, createmastercountsfile, cutadapt, dcc, dcccreatesamplesheets, estimateduplication, fastqc, findcirc, findcircalign, mergeSJtabs, mergealignmentstats, mergegenecounts, mergepersample, star1p, star2p, star_circrnafinder
This was a dry-run (flag -n). The order of jobs does not reflect the order of execution. Running... 14743440 ```
6.1 Test Data
The /data/CCBR_Pipeliner/testdata/circRNA/human folder in the repo has test dataset:
bash
tree /data/CCBR_Pipeliner/testdata/circRNA/human
/data/CCBR_Pipeliner/testdata/circRNA/human
GI1_N_ss.R1.fastq.gz
GI1_N_ss.R2.fastq.gz
GI1_T_ss.R1.fastq.gz
samples.tsv
GI1_N is a PE sample while GI1_T is a SE sample.
6.2 Expected Output
Expected output from the sample data is stored under .tests/expected_output.
More details about running test data can be found here.
DISCLAIMER:
CHARLIE is built to be run only on BIOWULF. A newer HPC-agnostic version of CHARLIE is planned for 2024.
Owner
- Name: CCR Collaborative Bioinformatics Resource
- Login: CCBR
- Kind: organization
- Email: nciccbr@mail.nih.gov
- Location: United States of America
- Website: https://bioinformatics.ccr.cancer.gov/ccbr/
- Repositories: 92
- Profile: https://github.com/CCBR
CCR Collaborative Bioinformatics Resource, Center for Cancer Research (NCI), National Institutes of Health
GitHub Events
Total
- Create event: 18
- Release event: 3
- Issues event: 16
- Watch event: 1
- Delete event: 14
- Issue comment event: 28
- Push event: 61
- Pull request event: 27
Last Year
- Create event: 18
- Release event: 3
- Issues event: 16
- Watch event: 1
- Delete event: 14
- Issue comment event: 28
- Push event: 61
- Pull request event: 27




