brainglobe-napari-io
Read and write files from the BrainGlobe neuroanatomy suite
Science Score: 54.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
✓CITATION.cff file
Found CITATION.cff file -
✓codemeta.json file
Found codemeta.json file -
✓.zenodo.json file
Found .zenodo.json file -
○DOI references
-
○Academic publication links
-
✓Committers with academic emails
1 of 4 committers (25.0%) from academic institutions -
○Institutional organization owner
-
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (13.3%) to scientific vocabulary
Keywords
Keywords from Contributors
Repository
Read and write files from the BrainGlobe neuroanatomy suite
Basic Info
Statistics
- Stars: 14
- Watchers: 3
- Forks: 6
- Open Issues: 4
- Releases: 7
Topics
Metadata Files
README.md
napari-brainglobe-io
Visualise cellfinder and brainreg results with napari
Installation
This package is likely already installed
(e.g. with cellfinder, brainreg or another napari plugin), but if you want to
install it again, either use the napari plugin install GUI or you can
install brainglobe-napari-io via [pip]:
pip install brainglobe-napari-io
Usage
- Open napari (however you normally do it, but typically just type
napariinto your terminal, or click on your desktop icon)
brainreg
Sample space
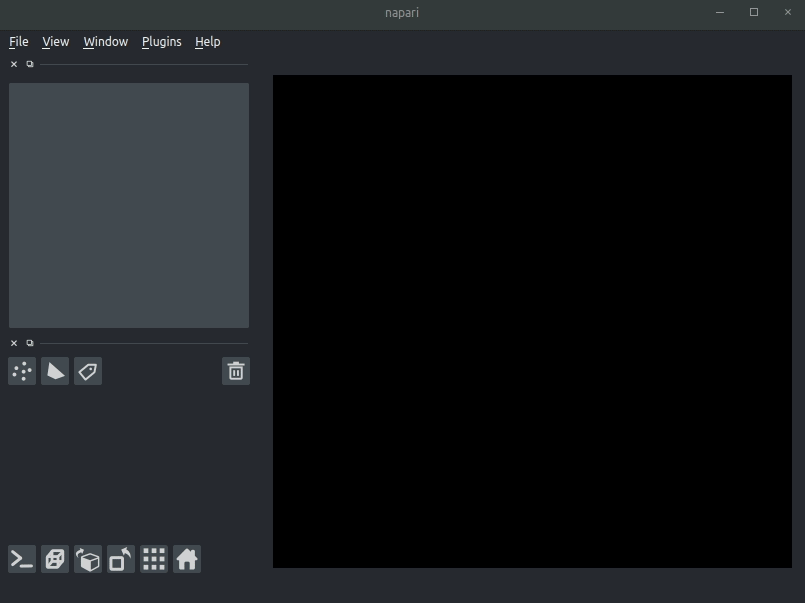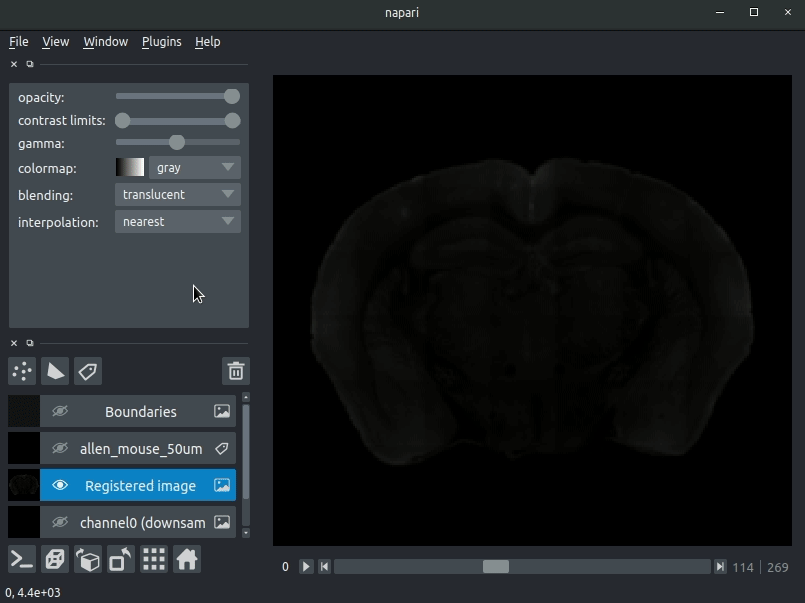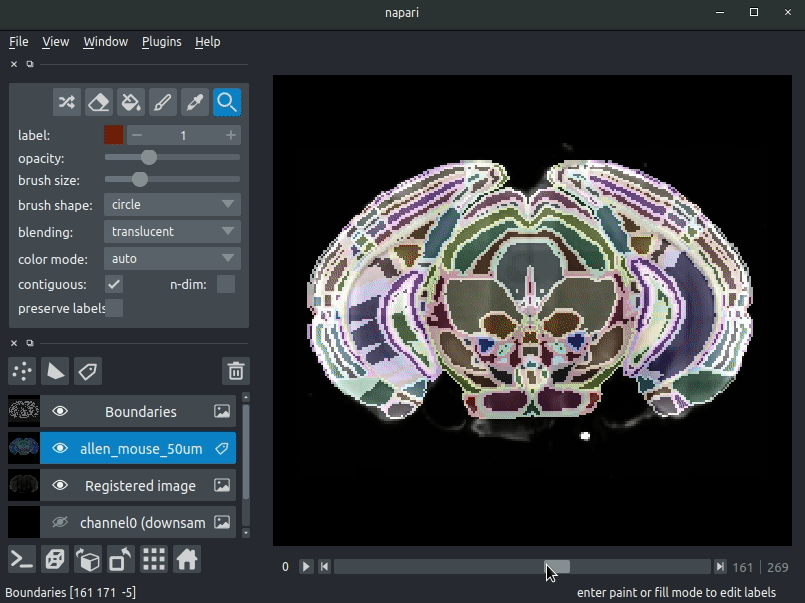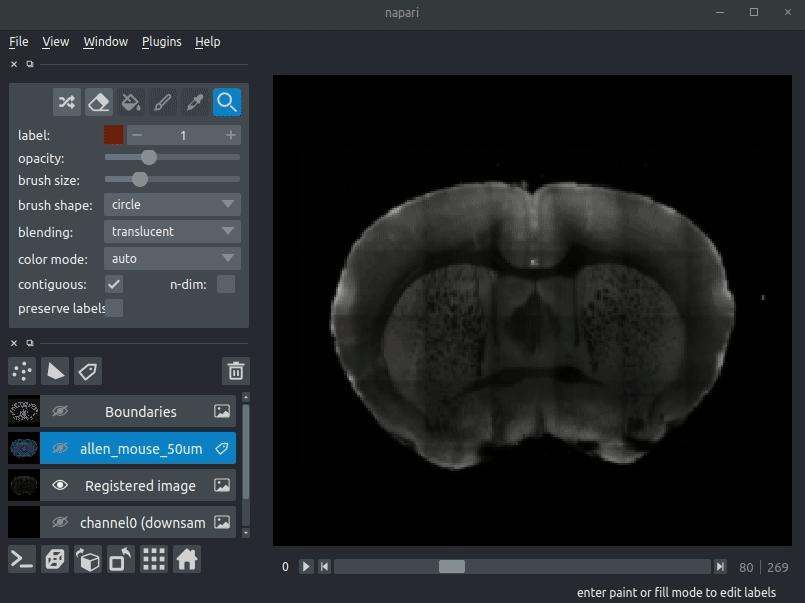
Drag your brainreg output directory (the one with the log file) onto the napari window.
Various images should then open, including:
* Registered image - the image used for registration, downsampled to atlas resolution
* atlas_name - e.g. allen_mouse_25um the atlas labels, warped to your sample brain
* Boundaries - the boundaries of the atlas regions
If you downsampled additional channels, these will also be loaded.
Most of these images will not be visible by default. Click the little eye icon to toggle visibility.
N.B. If you use a high resolution atlas (such as `allenmouse10um`), then the files can take a little while to load.
Atlas space
napari-brainreg also comes with an additional plugin, for visualising your data
in atlas space.
This is typically only used in other software, but you can enable it yourself:
* Open napari
* Navigate to Plugins -> Plugin Call Order
* In the Plugin Sorter window, select napari_get_reader from the select hook... dropdown box
* Drag brainreg_read_dir_atlas_space (the atlas space viewer plugin) above brainreg_read_dir (the normal plugin) to ensure that the atlas space plugin is used preferentially.
cellfinder
Load cellfinder XML file
- Load your raw data (drag and drop the data directories into napari, one at a time)
- Drag and drop your cellfinder XML file (e.g.
cell_classification.xml) into napari.
Load cellfinder directory
- Load your raw data (drag and drop the data directories into napari, one at a time)
- Drag and drop your cellfinder output directory into napari.
The plugin will then load your detected cells (in yellow) and the rejected cell candidates (in blue). If you carried out registration, then these results will be overlaid (similarly to the loading brainreg data, but transformed to the coordinate space of your raw data).
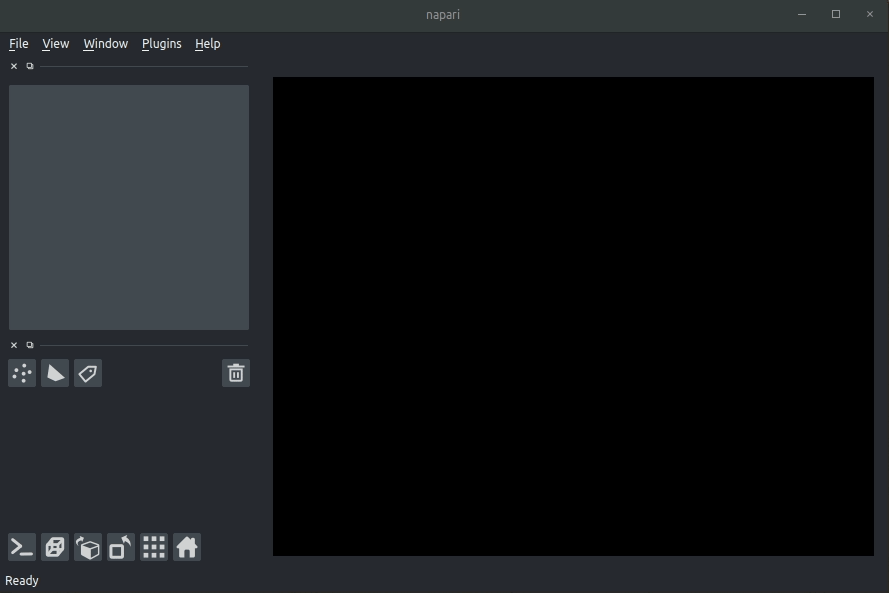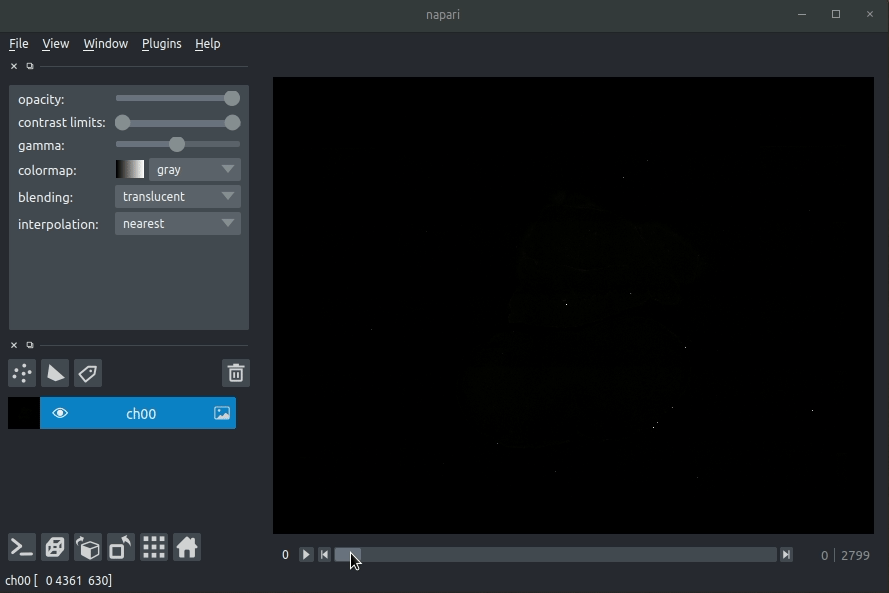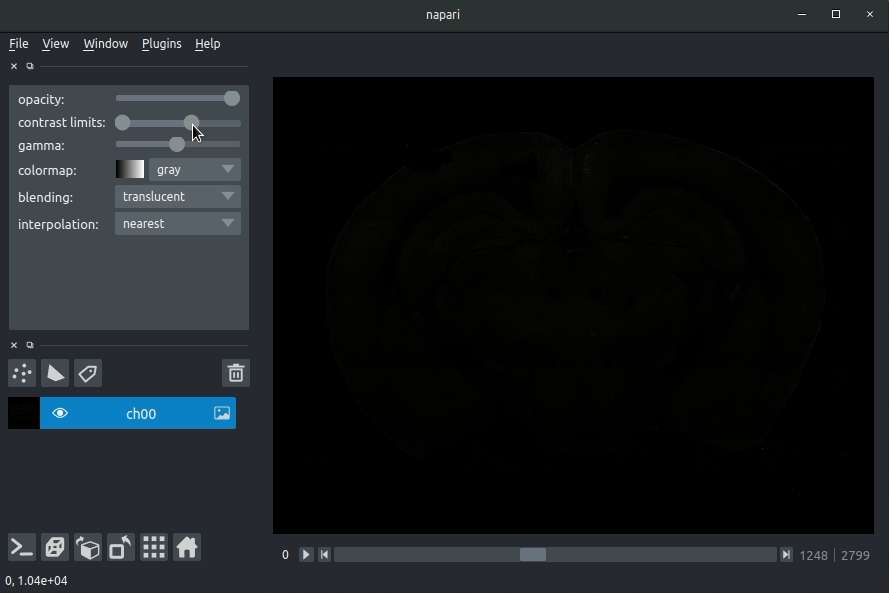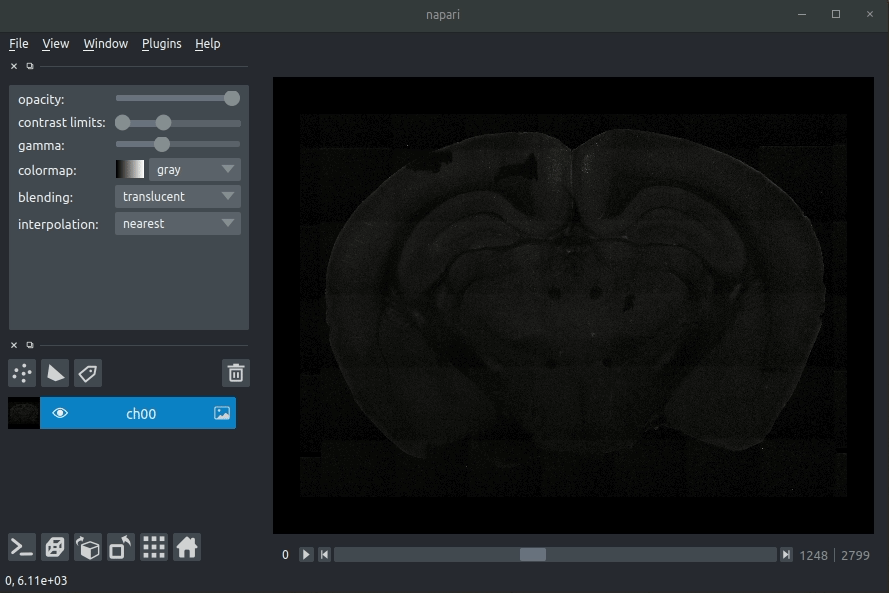 Loading raw data
Loading raw data
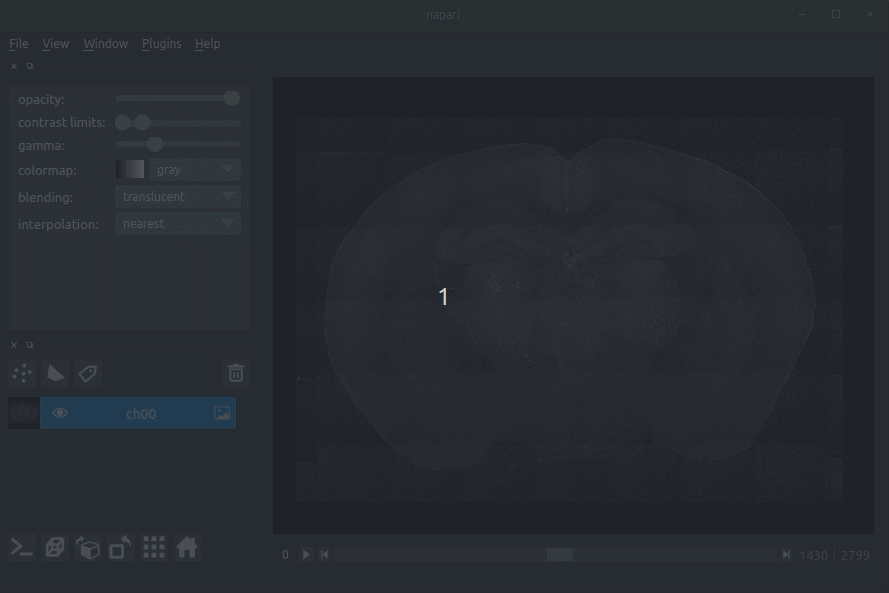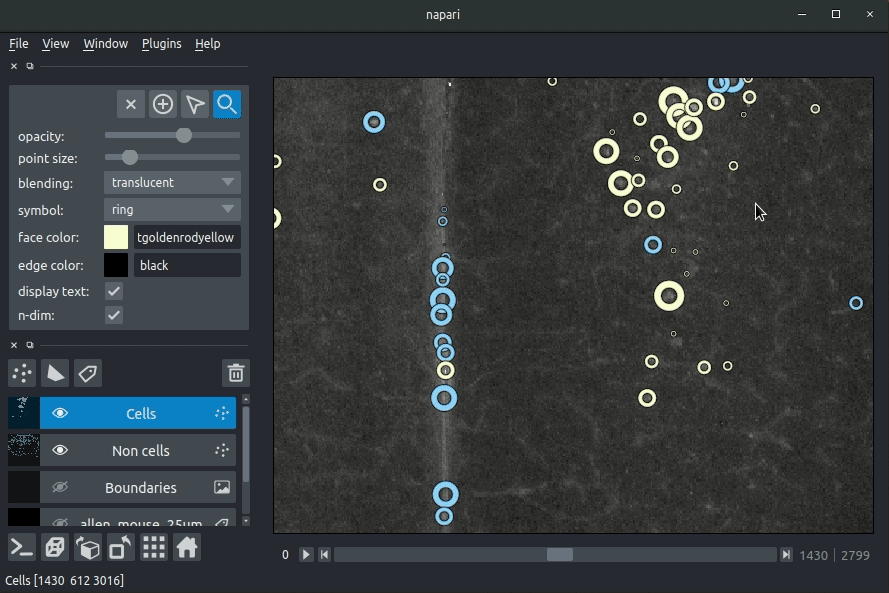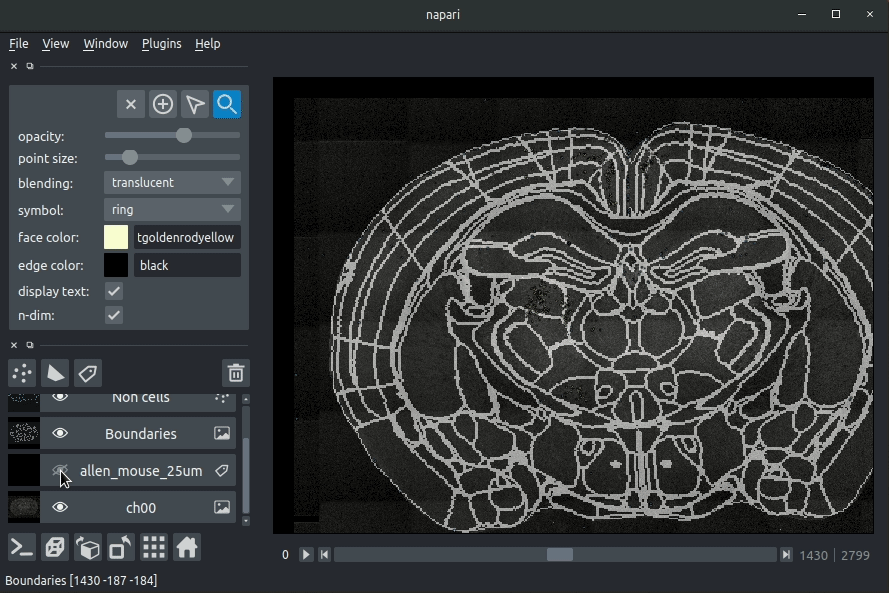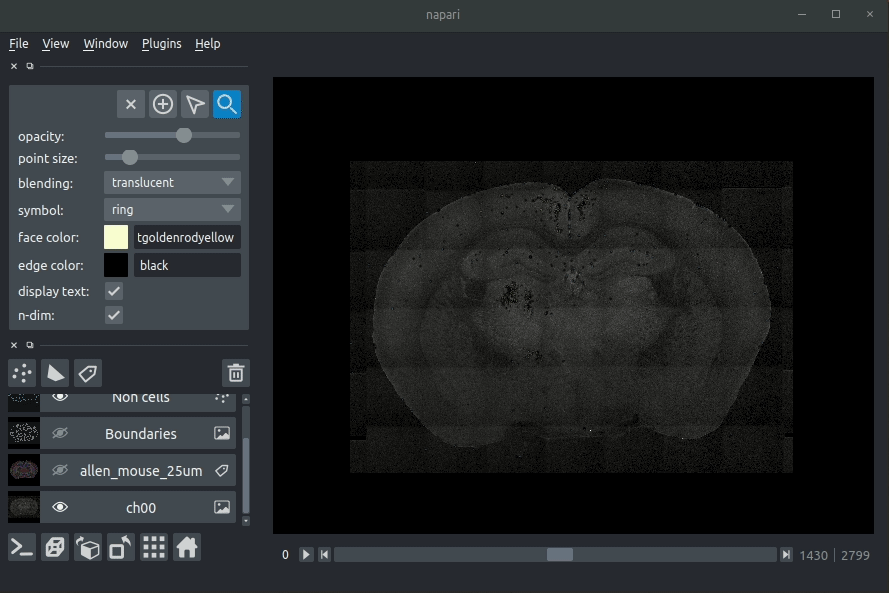 Loading cellfinder results
Loading cellfinder results
Seeking help or contributing
We are always happy to help users of our tools, and welcome any contributions. If you would like to get in contact with us for any reason, please see the contact page of our website.
Owner
- Name: BrainGlobe
- Login: brainglobe
- Kind: organization
- Location: London/Munich
- Website: https://brainglobe.info/
- Twitter: brain_globe
- Repositories: 28
- Profile: https://github.com/brainglobe
Open python tools for morphological analyses in systems neuroscience
Citation (CITATION.cff)
# This CITATION.cff file was generated with cffinit.
# Visit https://bit.ly/cffinit to generate yours today!
cff-version: 1.2.0
title: brainglobe-napari-io
message: >-
If you use this software, please cite it using the
metadata from this file.
type: software
authors:
- given-names: BrainGlobe
family-names: Developers
email: hello@brainglobe.info
- given-names: Adam
family-names: Tyson
email: hello@brainglobe.info
affiliation: 'Sainsbury Wellcome Centre, University College London'
repository-code: 'https://github.com/brainglobe/brainglobe-napari-io'
url: 'https://brainglobe.info'
abstract: Visualise cellfinder and brainreg results with napari.
license: BSD-3-Clause
date-released: '2024-01-25'
year: 2024
GitHub Events
Total
- Create event: 22
- Release event: 2
- Issues event: 6
- Watch event: 1
- Delete event: 15
- Issue comment event: 6
- Push event: 19
- Pull request review event: 18
- Pull request event: 36
- Fork event: 2
Last Year
- Create event: 22
- Release event: 2
- Issues event: 6
- Watch event: 1
- Delete event: 15
- Issue comment event: 6
- Push event: 19
- Pull request review event: 18
- Pull request event: 36
- Fork event: 2
Committers
Last synced: about 3 years ago
Top Committers
| Name | Commits | |
|---|---|---|
| Adam Tyson | c****e@a****m | 24 |
| David Stansby | d****y@g****m | 14 |
| Adam Tyson | a****n@u****k | 7 |
| chili-chiu | 8****u@u****m | 1 |
Committer Domains (Top 20 + Academic)
Issues and Pull Requests
Last synced: 9 months ago
All Time
- Total issues: 15
- Total pull requests: 98
- Average time to close issues: 11 months
- Average time to close pull requests: 1 day
- Total issue authors: 10
- Total pull request authors: 10
- Average comments per issue: 1.6
- Average comments per pull request: 0.65
- Merged pull requests: 95
- Bot issues: 0
- Bot pull requests: 36
Past Year
- Issues: 3
- Pull requests: 37
- Average time to close issues: N/A
- Average time to close pull requests: about 6 hours
- Issue authors: 3
- Pull request authors: 6
- Average comments per issue: 0.67
- Average comments per pull request: 0.0
- Merged pull requests: 34
- Bot issues: 0
- Bot pull requests: 16
Top Authors
Issue Authors
- adamltyson (4)
- dstansby (3)
- matham (1)
- chrytsi (1)
- alessandrofelder (1)
- noisysky (1)
- stephenlenzi (1)
- kailynkfields (1)
- willGraham01 (1)
- marcodallavecchia (1)
Pull Request Authors
- pre-commit-ci[bot] (46)
- adamltyson (24)
- IgorTatarnikov (15)
- dstansby (14)
- willGraham01 (8)
- alessandrofelder (5)
- K-Meech (3)
- lochhh (2)
- marcodallavecchia (2)
- chili-chiu (1)
Top Labels
Issue Labels
Pull Request Labels
Packages
- Total packages: 2
-
Total downloads:
- pypi 1,105 last-month
-
Total dependent packages: 10
(may contain duplicates) -
Total dependent repositories: 1
(may contain duplicates) - Total versions: 24
- Total maintainers: 1
pypi.org: brainglobe-napari-io
Read and write files from the BrainGlobe computational neuroanatomy suite into napari
- Homepage: https://brainglobe.info
- Documentation: https://docs.brainglobe.info
- License: BSD-3-Clause
-
Latest release: 0.4.0
published 9 months ago
Rankings
Maintainers (1)
conda-forge.org: brainglobe-napari-io
- Homepage: https://brainglobe.info
- License: BSD-3-Clause
-
Latest release: 0.1.5
published about 4 years ago
Rankings
Dependencies
- actions/checkout v2 composite
- actions/setup-python v2 composite
- brainglobe/actions/lint v1 composite
- brainglobe/actions/test main composite
- tlambert03/setup-qt-libs v1 composite
- bg-atlasapi *
- bg_space *
- brainglobe-utils *
- napari *
- numpy *
- pandas *
- tifffile >=2020.8.13


