blotr
Rapidly extract band-intensities from Line immunoblots EuroImmun EuroLine Blot
Science Score: 57.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
✓CITATION.cff file
Found CITATION.cff file -
✓codemeta.json file
Found codemeta.json file -
✓.zenodo.json file
Found .zenodo.json file -
✓DOI references
Found 1 DOI reference(s) in README -
○Academic publication links
-
○Academic email domains
-
○Institutional organization owner
-
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (15.2%) to scientific vocabulary
Repository
Rapidly extract band-intensities from Line immunoblots EuroImmun EuroLine Blot
Basic Info
- Host: GitHub
- Owner: atroel01
- License: gpl-3.0
- Language: R
- Default Branch: main
- Size: 37.1 KB
Statistics
- Stars: 0
- Watchers: 1
- Forks: 0
- Open Issues: 0
- Releases: 0
Metadata Files
README.md
BlotR
Rapidly extract band-intensities from Line immunoblots EuroImmun EuroLine Blot
Purpose
This Shiny App is designed to allow you to rapidly extract the band-intensities from EuroImmun Blot results EuroLine LIA results must be generated from EuroImmun (R) scanner software and saved in single page view (one result per pdf), showing the scanned version of the band. - An example of the appearance of the blot below:
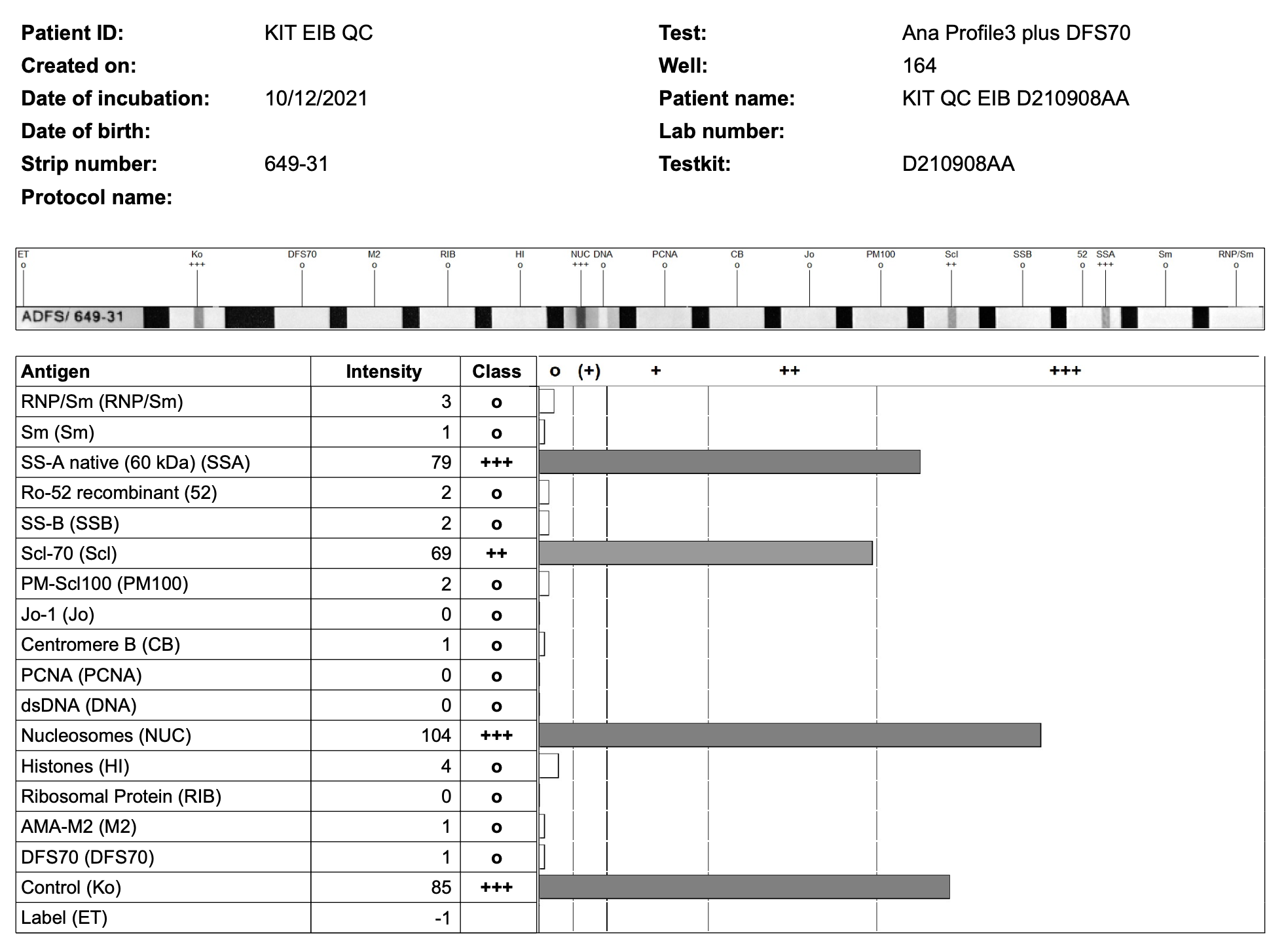
Currently the app will extract details: - Generic: Patient Name, DOB, Lab Number, Created On, Date of Incubation, Patient ID - Specific blots - Myositis 16 Ag LIA - Scleroderma (Nucleoli) profile - ENA profile 3 plus DFS - ENA Profile 5 - Liver Blot - Neuronal Blot 12 Ag
Requirements
Requires R (https://www.r-project.org/) RStudio (now called Posit) is recommended for simpler interfacing and editing of script (https://posit.co/).
Download both R scripts (app.R and fun.R) to chosen folder.
App can be executed through command line
- once R installed open a terminal or command line at app.R file path
- Enter command Rscript app.R and visit the url generated (an example below)
Initial run of application will require download of required dependencies
Usage
Select all of the blots at once that you wish to parse using the browser - any re-selection will replace all existing files selected. Please note as the app must copy the files to its directory there will be some time delay when there are many files to register them all.
Scanner detection
Flatbed and EuroBlotOne scanner results have differing cut-offs. If you have a variety of scanners in use then distinguishing these results maybe important. For example, in contrast to the previous scan which was generated by the EuroBlotOne, the flatbed scanner outputs results on a green paper background.

If you wish to use the scanner detection functionality then you must tick the box before hitting run. Detecting the color of the strip in the PDF is time consuming and will add substantial time to the parsing function. This will determine whether the strip was read by the EuroBlotOne e.g. strip scan is black and white; or green e.g. flatbed scanner.
Bug: Sometimes changing tabs (Summary of Tests/Summary Graph/Repeated) is required to initiate parsing.
Filtering
You can enter names of results you wish to filter out or highlight as qc values in lower case (comma seperated e.g. qc, eib, qap,...). Filtering is based on regular expression detection and so filtering terms must be reasonably specific or it may perform not as intended.
Output
Once the blots have been parsed the app will have outputs of - Table of the number of positives and their overall intensity - Graph of all results (output graph when large data takes a while to generate - this is in plotly format) - Repeated results, intensities and changes where name and DOB match (incorrect entry of these results will make it miss repeated values). - A spreadsheet with one blot result per row and the intensities will be available for download from the download button.
Citation
@software{Troelnikov_BlotR_2022,
author = {Troelnikov, Alexander and Harvey, Isaac},
doi = {10.5281/zenodo.1234},
month = {11},
title = {{BlotR}},
url = {https://github.com/atroel01/BlotR},
version = {1.0.0},
year = {2022}
}
Owner
- Name: Alexander Troelnikov
- Login: atroel01
- Kind: user
- Location: Adelaide, South Australia
- Company: Flinders University
- Twitter: ATroelnikov
- Repositories: 1
- Profile: https://github.com/atroel01
Immunology and Immunopathology Registrar who enjoys tinkering with code. Occassionally some of it is useful.
Citation (CITATION.cff)
cff-version: 1.2.0 message: "If you use this software, please cite it as below." authors: - family-names: "Troelnikov" given-names: "Alexander" orcid: "https://orcid.org/0000-0002-7239-3814" - family-names: "Harvey" given-names: "Isaac" orcid: "https://orcid.org/0000-0000-0000-0000" title: "BlotR" version: 1.0.0 doi: 10.5281/zenodo.1234 date-released: 2022-11-12 url: "https://github.com/atroel01/BlotR"