Recent Releases of bigsdb
bigsdb - Version 1.51.4
This version introduces a new Kaptive plugin for surface polysaccharide typing of Acinetobacter baumannii and Klebsiella. The plugin is a wrapper that runs Kaptive for selected isolates, outputs the results in tabular and SVG format and also stores these within the database so that they can then be included within the isolate information page.
The Kaptive plugin is disabled by default and it needs to be enabled with Kaptive="yes" within the database config.xml file. You also need to select the appropriate Kaptive databases to use as a comma-separated list with the kaptive_dbs attribute - choices are kpsck (Klebsiella_ K locus), kpsco (Klebsiella_ O locus), abk (Acinetobacter baumannii_ K locus), and abo (Acinentobacter baumannii_ OC locus), e.g. ‘kpsck,kpsco’.
Because Kaptive analysis is only valid against certain species, you may wish to restrict which isolates in a database that it can be run against, e.g. the Acinetobacter baumannii analysis is only valid for A. baumannii and not other Acinetobacter species that may be in the database. To restrict this, you can set the kaptive_view attribute, that names a view of the isolate table that is appropriately filtered. An example of such a view can be constructed with the following SQL for an isolate database that contains a species field:
CREATE VIEW abaumannii AS SELECT * FROM isolates WHERE species='Acinetobacter baumannii';
GRANT SELECT ON abaumannii TO apache;
You can also add a description of the view with the kaptive_view_desc which will appear in the interface to explain why some isolates have been filtered prior to running Kaptive. The full Kaptive configuration attributes used for the PubMLST Acinetobacter spp. database are:
Kaptive="yes"
kaptive_dbs="ab_k,ab_o"
kaptive_view="abaumannii"
kaptive_view_desc="Restricted to A. baumannii isolates only."
There is also a script update_kaptive.pl found in scripts/maintenance that can run Kaptive analysis against all valid isolates within a database (or all databases for which Kaptive is enabled), storing the results within the analysis_results table for display within isolate records. Command options are:
``` updatekaptive.pl --help NAME updatekaptive.pl - Perform/update Kaptive analysis
SYNOPSIS update_kaptive.pl [options]
OPTIONS
--database DATABASE CONFIG Database configuration name. If not included then all isolate databases with the Kleborate flag set on their configuration will be checked.
--exclude CONFIG NAMES Comma-separated list of config names to exclude.
--help This help page.
--lastrundays DAYS Only run for a particular isolate when the analysis was last performed at least the specified number of days ago.
--quiet Only show errors.
--refresh_days DAYS Refresh records last analysed longer that the number of days set. By default, only records that have not been analysed will be checked.
--view, -v VIEW Isolate database view (overrides the view set in config.xml). ```
If you use the results of Kaptive, you should cite the following:
Stanton TD, Hetland MAK, Löhr IH, Holt KE, Wyres KL. Fast and Accurate in silico Antigen Typing with Kaptive 3 [Internet]. bioRxiv; 2025 [cited 2025 Feb 27]. p. 2025.02.05.636613. https://www.biorxiv.org/content/10.1101/2025.02.05.636613v1
Documentation for Kaptive can be found at https://kaptive.readthedocs.io/en/latest/. Please consult this for other citations that should be included when using specific Kaptive databases.
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.51.3...v1.51.4
- Perl
Published by kjolley 11 months ago
bigsdb - Version 1.51.3
This version provides the following enhancements:
- Distance matrix calculation is now significantly quicker in Genome Comparator. Various optimisations were made to avoid redundant hash lookups and calculations. The inner loop that performed pairwise comparisons was rewritten in C to provide a major speed boost. Note that this requires a new Perl module dependency: Inline::C.
- There is now an option to disable distance matrix calculation altogether if the user does not require this analysis, e.g. if they're just interested in generating core-genome aligned sequences.
- Finally, an option has been added to the iTOL and Microreact plugins to use allelic profiles, rather than aligned DNA sequences, for construction of neighbour-joining trees. This is much faster and allows for larger datasets but trees will be based on allelic differences rather than nucleotide differences.
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.51.2...v1.51.3
- Perl
Published by kjolley 11 months ago
bigsdb - Version 1.51.2
This version speeds up construction of datasets for Genome Comparator and GrapeTree where allele designations have already been stored in the database.
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.51.1...v1.51.2
- Perl
Published by kjolley 11 months ago
bigsdb - Version 1.51.1
This version adds OAuth authentication to calls to the PubMLST rMLST species identification API service used by the species id plugin and submission system. It is likely that this service will require authentication in the near future.
You will need to obtain a PubMLST API key. You can get this from https://pubmlst.org/bigsdb. Once you have this, use the get_oauth_access_token.pl script with an account that has registered access to the rMLST typing database to obtain an access token and secret. You can request access for the rMLST database also at https://pubmlst.org/bigsdb.
Once you have your API key and secret, and the access token and secret, add these to bigsdb.conf (uncomment the lines beginning with a semi-colon):
```
RMLST SPECIES ID
Register for a PubMLST API key and obtain an access token and secret for an
account with access to the rMLST typing database.
Use the getoauthaccess_token.pl script to obtain the access token with web
address: https://pubmlst.org/bigsdb?db=pubmlstrmlstseqdef and REST API
address: https://rest.pubmlst.org/db/pubmlstrmlstseqdef.
Multiple concurrent connections are restricted so please do not set threads
to >1 unless authorised.
speciesidthreads=1 ;rmlstclientkey=xxxxxxxxxxxx ;rmlstclientsecret=xxxxxxxxxxxx ;rmlstaccesstoken=xxxxxxxxxxxx ;rmlstaccesssecret=xxxxxxxxxxxx ```
- Perl
Published by kjolley about 1 year ago
bigsdb - Version 1.51.0
This version adds support for recording user affilation sector and country to site-wide users databases.
If enabled, when users register for a site account, they will see a compulsory dropdown list of sector options. These are:
- academic/non-profit
- commerical
- healthcare
- public health
- other
If 'other' is selected, then an additional free text field will appear allowing the user to add a different value.
To enable sector and country recording, set the following in bigsdb.conf:
```
Specify whether site users should register affiliation sector and/or country.
siteusersector=1 siteusercountry=1 ```
It is also possible to force users to update their profiles when they log in. There is a new boolean field in the auth database users table update_profile. Set this to TRUE to force a profile update.
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.50.3...v1.51.0
- Perl
Published by kjolley about 1 year ago
bigsdb - Version 1.50.3
This version adds support for Kleborate v3 to the Kleborate plugin. The plugin supports both v2 and v3, automatically detecting which version is used.
If version 3 is used, there is also a new option that can be used to define the preset modules that will be used for analysis. This can be set in the database config.xml file, e.g.
kleborate_preset="kpsc"
Available options are:
- kpsc (Klebsiella pneumoniae species complex) [default]
- kosc (Klebsiella oxytoca species complex)
- escherichia (Escherichia coli)
What's Changed
- add TMPDIR environment variable usage by @braffes in https://github.com/kjolley/BIGSdb/pull/1031
New Contributors
- @braffes made their first contribution in https://github.com/kjolley/BIGSdb/pull/1031
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.50.2...v1.50.3
- Perl
Published by kjolley about 1 year ago
bigsdb - Version 1.50.2
This version provides a fix for API profile downloads where a date restriction is set requiring users to log in to access recent data. Profile downloads are now streamed asynchronously for performance reasons and the user state was not being maintained so it was acting as if a user was not logged in.
It also fixes the set_embargo permission which was not being recognized for non-admin curators.
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.50.1...v1.50.2
- Perl
Published by kjolley over 1 year ago
bigsdb - Version 1.50.1
This version:
- adds supports for Neighbour-joining trees in the GrapeTree plugin
- adds a means to restrict use of particular plugins to specific users or user groups
- adds an option to override system attributes for specific users or user groups to enable different plugin behaviour or limits
- adds the inclusion of registered analysis result fields in API isolate records
NJ trees in GrapeTree plugin There is now support for Neighbour-joining trees in the GrapeTree plugin. Trees are still generated based on allelic profiles rather than allele sequences. Tree-drawing algorithms now include:
- MSTreeV2 - Improved minimum-spanning tree that better handles missing data (default - previously this was the only option)
- MSTree - Classical minimum-spanning tree similar to PhyloViz
- NJ - FastME V2 Neighbour-joining tree
- RapidNJ - RapidNJ Neighbour-joining tree for very large datasets
Restrict use of particular plugins to specific users or user groups A file called restrictions.conf can be placed in the site configuration directory (/etc/bigsdb). This file allows you to define global user groups and then restrict access to particular plugins to either a user group or individual users. This is useful for testing new analysis plugins on a public database. The format for the file is as below:
```
Set restrictions to access for specific plugins to named users or usergroups.
Usergroups are set within specific databases and can also be defined here.
[Usergroups] beta_testers=user1,user2
Specific plugins
Default is to allow access (default=allow) - set default=deny to restrict
access and then allow access to specified users or user groups. Add a new
section for each plugin that you wish to restrict.
[GenomeComparator] default=deny allowedusers=user1,user2 allowedusergroups=beta_testers ```
Override system attributes for specific users or user groups The system.overrides file (described at https://bigsdb.readthedocs.io/en/latest/dbase_setup.html#over-riding-values-set-in-config-xml) has been extended to allow the overriding of configuration parameters for specific users or user groups. This enables different limits to be set for plugins to different sets of users.
An example of a system.overrides file can be seen below. The top section is for global changes, and then different sections can be added for a user or usergroup.
``` view="private" readaccess="authenticatedusers" description="Private view of database"
[user:keith] genomecomparatorlimit=3000
[usergroup:Power users] grapetree_limit=20000 ```
Inclusion of registered analysis result fields in API isolate records Registered analysis fields (released in v1.50.0 - are now included in isolate records retrieved via the API.
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.50.0...v1.50.1
- Perl
Published by kjolley over 1 year ago
bigsdb - Version 1.50.0
This version introduces support for registering fields from the output of third-party analysis tools that store their results within the database. These fields can then be searched, exported, or used by other analysis tools.
BIGSdb can store the results of arbitrary analyses as JSON within the analysisresults table without requiring any understanding of what the data represent. Information from this can be extracted and formatted within an isolate information page using template files stored in /etc/bigsdb/templates or in /etc/bigsdb/dbases/{dbaseconfig}/templates. This is used by the rMLST species id and Kleborate plugins to store their results and make them available to everyone once these analyses have been run by anyone, or by an offline script.
It is now possible to register specific fields from this output and make them available for searching, analysis (currently only in GrapeTree), and export. To do this, you need to be able to describe the location of the field in question within the JSON field. This can be done using JSONPath, e.g. a field called ‘predictedserotype’ at the top-level within the JSON hierarchy can be described with a JSONPath of ‘$.predictedserotype’.
Admins can add fields by expanding the ‘Fields’ selection in the admin interface and clicking the ‘Add’ link for ‘Analysis fields’:

Enter the analysis tool name, which must match the value used within the analysisresults table, the name of the field, and its JSONPath. You also need to indicate the data type (integer, text, float, or date). Optionally you can set the analysisdisplay_name to change how it is listed within the query and analysis interfaces.

This will now be searchable within the isolate query interface. First you will need to select the ‘Analysis results’ form elements from the ‘Modify form’ tab.

Then select the field and the value to search on.

Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.49.1...v1.50.0
- Perl
Published by kjolley over 1 year ago
bigsdb - Version 1.49.1
This version:
- Provides a fix for the LIN code assignment script that was failing when using the latest version of the Perl Data Library (PDL).
- Improves the layout for the site user registration and administration pages. This now utilizes an accordion view to hide sections rather than displaying all form elements on a single page.
- Allows users to now upload their own genomes for analysis in Genome Comparator and related plugins as .tar.gz, .tar, and fasta.gz files rather than just as .zip files as previously.
- Improves the layout of the Export plugin to hide rarely used options. The form can be modified by clicking the 'Modify Form' trigger to display these. An option has also been added to allow export of LIN code prefixes of selected lengths.
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.49.0...v1.49.1
- Perl
Published by kjolley over 1 year ago
bigsdb - Version 1.49.0
- This version enables users with a site-wide account to generate their own API keys. There is now a new section on the user registation page which shows any existing key and allows their revocation or creation of new keys.
[Note the key shown above has been revoked :-)]
There has also been a fix to the verification of OAuth signatures in an API POST query. Previously the signature base string included JSON-encoded parameters, which should not have been included according to the OAuth 1.0A spec.
Finally, curators with appropriate permissions and settings will now receive an E-mail notification if a user requests that private records are made public.
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.48.5...v1.49.0
- Perl
Published by kjolley over 1 year ago
bigsdb - Version 1.48.5
This is a minor release that:
- improves the layout of the user database registration page, making use of the JQuery.multiselect plugin.
- provides an option to select only DNA or only peptide loci when making a sequence query in databases that contain both kinds.
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.48.4...v1.48.5
- Perl
Published by kjolley over 1 year ago
bigsdb - Version 1.48.4
This version speeds up the display of results for an isolate query where scheme fields from large schemes (e.g. cgST for cgMLST schemes) are displayed. The scheme values used to be determined from the isolate allele designations in real time so that they were always up-to-date, but this can take 500ms+ for a large cgMLST scheme per isolate, which is noticeable when displaying a table of 25 records. Now, if 'cache_schemes' is set in config.xml, and the scheme has >100 loci then the scheme cache table is used for the lookup. Smaller schemes (e.g. MLST) are still determined using a live lookup.
If a scheme field has multiple values, which can happen for cgSTs due to missing loci, then these are now collapsed in the results table with only the cgST that has the fewest missing loci shown. Other matching cgSTs can be shown by clicking a hyperlink.
The second record below has the list expanded:
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.48.3...v1.48.4
- Perl
Published by kjolley over 1 year ago
bigsdb - Version 1.48.3
This version introduces a number of performance improvements to sequence queries. The cached BLAST database now also includes sequence length in its headers to avoid a database lookup during the query - to see maximum improvement immediately, mark any BLAST caches as stale (using link in admin interface) so that they can be re-created. The query will still work using the old cache if this isn't done, and the cache will be recreated automatically once a new allele is added or when it is >7 days old.
If LIN codes are assigned for a scheme these are now used to identify the nearest matching genomes following a sequence query.
Please note the new dependency for Perl module Text::CSV.
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.48.2...v1.48.3
- Perl
Published by kjolley over 1 year ago
bigsdb - Version 1.48.2
This release fixes a bug when attempting to download a scheme TSV file from the API when the scheme does not have a primary key field, e.g. https://rest.pubmlst.org/db/pubmlstneisseriaseqdef/schemes/6/profiles_csv. Due to the recent change in the way the results were streamed (introduced in v1.48.1), the check for a valid scheme was not occurring in time, and the process would crash when attempting to access a non-existent table.
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.48.1...v1.48.2
- Perl
Published by kjolley over 1 year ago
bigsdb - Version 1.48.1
This version fixes a couple of issues:
- LIN code prefix lookup of nickname field values was matching multiple values, e.g. the prefix 001051 was also returning values for prefixes 00105, 0010 and 001.
- LIN code assignment was failing when a scheme had a field called 'profile_id'.
In additional a performance issue has been addressed for the API cgMLST profile download in TSV format. This was taking a long time (and potentially timing out).
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.48.0...v1.48.1
- Perl
Published by kjolley over 1 year ago
bigsdb - Version 1.48.0
This version adds optional placeholder text to search fields in isolate queries. You can add placeholder text for any of the fields defined in config.xml by setting the placeholder attribute, e.g.
<field type="text" required="no" length="50" maindisplay="no" comments="region inside country" placeholder="e.g. Oxfordshire">region</field>
Placeholder text can also be set for sparse fields, scheme fields, LIN codes, and LIN code fields by setting the placeholder value in the appropriate table - (in the admin interface):
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.47.3...v1.48.0
- Perl
Published by kjolley over 1 year ago
bigsdb - Version 1.47.3
Potential fix for https://github.com/kjolley/BIGSdb/issues/970.
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.47.2...v1.47.3
- Perl
Published by kjolley almost 2 years ago
bigsdb - Version 1.47.2
This version adds hooks to PluginManager.pm, Plugin.pm and Offline::RunJobs.pm to run plugin tools written entirely in Python using the BIGSdb Python Toolkit.
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.47.1...v1.47.2
- Perl
Published by kjolley almost 2 years ago
bigsdb - Version 1.47.1
This version adds an option to isolate field configurations to enable a different hyperlink to be set depending on the regular expression of the field value. The is the web_regex attribute, demonstrated below:
<field type="text" required="genome_expected" length="20" maindisplay="no" web_regex="^SAM|https://www.ebi.ac.uk/ena/browser/view/[?]?dataType=BIOSAMPLE;^[EDS]RS|https://www.ebi.ac.uk/ena/browser/view/[?]?dataType=SAMPLE" web="https://www.ebi.ac.uk/ena/browser/view/[?]" comments="ENA/SRA biosample accession number" multiple="yes" group="Tracking">biosample_accession</field>
If the value of the biosample_accession field above starts with 'SAM' then the hyperlinked URL will be 'https://www.ebi.ac.uk/ena/browser/view/[?]?dataType=BIOSAMPLE' (where [?] is the field value), whereas if the field value starts with ERS, EDS, or SRS then the hyperlinked URL will be 'https://www.ebi.ac.uk/ena/browser/view/[?]?dataType=SAMPLE'. If none of the regex defined are matched then the value set for the web attribute will be used, e.g. https://www.ebi.ac.uk/ena/browser/view/[?] in the above.
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.47.0...v1.47.1
- Perl
Published by kjolley almost 2 years ago
bigsdb - Version 1.47.0
Version 1.47.0 introduces an embargo system for isolate submissions. This can be enabled in the bigsdb.conf file:
```
ISOLATE EMBARGOES
Set to enable isolate embargoes and limit the time allowed. These can be
overridden in individual isolate database config files.
Values are in months.
embargoenabled=1 defaultembargo=12 maxinitialembargo=24 max_embargo=48 ```
These values can also all be overridden in the config.xml file for a specific database, e.g. embargo_enabled="yes".
If enabled, submitters will now see an option to request an embargo when they submit. They can choose an embargo length up to the value set by maxinitialembargo (default 24 months).
Users will be able to see embargoed isolates when they are logged in and they behave like any other private data in that they can be shared with others by adding them to a shared user project. Curators will also be able to see them, whereas normal private isolates cannot be seen by curators.
It is also possible to search for embargoed isolates and by embargo date and users will see a hyperlinked reminder when they are logged in of how many embargoed isolates they have and the date at which the first of these will become public.
Curators with the appropriate permission set are able to modify embargo dates for both currently public isolates (making them private if they were submitted publicly by mistake) or for currently embargoed isolates.
Finally, you should run the check_embargoes.pl script once a day to automatically publish any isolate that reaches its embargo date.
In addition, quotas for projects has now been introduced. Previously only users could have quotas and projects could be set to be quota-free.
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.46.2...v1.47.0
- Perl
Published by kjolley almost 2 years ago
bigsdb - Version 1.46.2
BingMaps has been removed as an option for mapping as Microsoft will be retiring this service in June 2025. It has been replaced with options to use MapTiler (satellite) and ArcGIS streetmap/satellite imagery, alongside the existing option of OpenStreetMap for street level mapping.
OpenStreetMap
MapTiler
Esri/ArcGIS streetmap
Esri/ArcGIS satellite
You can select the appropriate options in bigsdb.conf (these can all also be overridden in individual database config.xml files if you wish to use different options or API keys):
```
MAPPING
Set the option for displaying location maps.
Note for Esri/ArcGIS services you must register
[https://developers.arcgis.com/en/sign-up/]
and abide by terms of service [https://developers.arcgis.com/en/terms/].
For MapTiler you need to register and get an API key [https://www.maptiler.com/].
0: OpenStreetMaps only
1: OpenStreetMaps + MapTiler satellite imagery
2: OpenStreetMaps + Esri/ArcGIS World Imagery (satellite)
3: Esri/ArcGIS World Street Map + Esri/ArcGIS World Imagery (satellite)
mappingoption=0 ;maptilerapi_key=xxxxxxxxxx ```
Note that use of MapTiler requires an API key - the free tier allows up to 100,000 map tile requests per month. You must also register for an Esri account to use Esri/ArcGIS services (you will need a paid account if using this commercially).
See the database setup documentation if you wish to enable mapping - either using a location field that contains GPS coordinates, or a geographic field such as 'townorcity' which can be linked to a lookup table of GPS coordinates. The latter lookup table can be automatically populated to find the coordinates of towns and cities using the gptownlookups.pl script.
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.46.1...v1.46.2
- Perl
Published by kjolley about 2 years ago
bigsdb - Version 1.46.1
Version 1.46.1 introduces a new SNPsites plugin.
The SNPsites plugin aligns sequences for specified loci for an isolate dataset. The alignment is then passed to snp-sites to identify SNP positions. Output consists of a summary table including the number of alleles and polymorphic sites found for each locus, an interactive D3 chart that displays the summary, and ZIP files containing alignment FASTAs and VCF files for each locus.
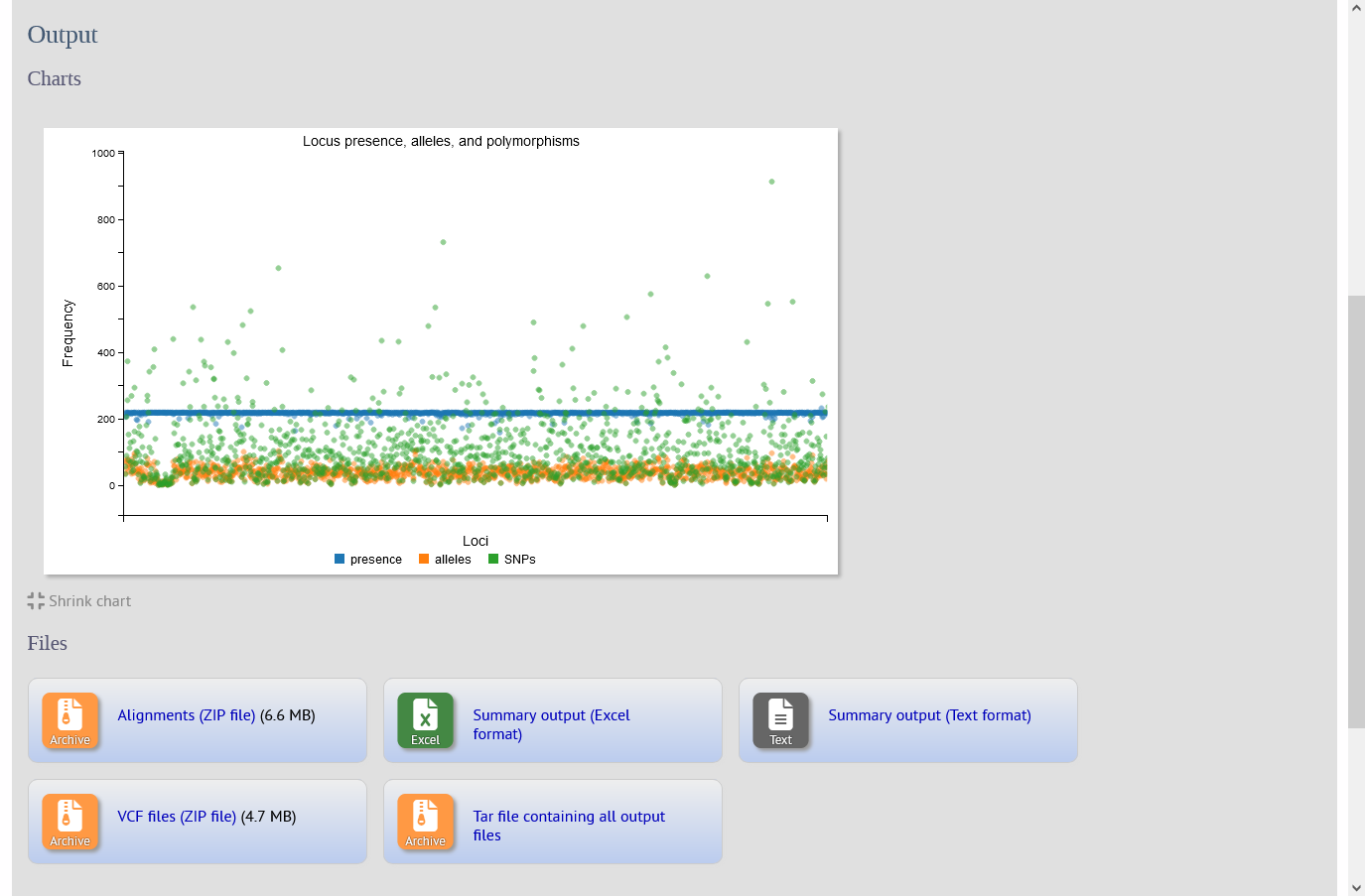
Note that snp-sites needs to be installed and its path set in bigsdb.conf, e.g.
```
snp-sites
snpsitespath=/usr/bin/snp-sites ```
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.46.0...v1.46.1
- Perl
Published by kjolley about 2 years ago
bigsdb - Version 1.46.0
Version 1.46.0 introduces the following changes:
- An option has been added to filter alleles, profiles, and isolates for non-logged in users to records added before a specified date. This requires users to log in if they wish to access more recent data and has been introduced to provide licensing options. This can be activated globally in bigsdb.conf or within individual database config.xml files by setting the logintoshowafterdate attribute.
In bigsdb.conf:
login_to_show_after_date=2024-01-01
In config.xml:
login_to_show_after_date="2024-01-01"
- Fix for missing foreign key on the queryinterfacefields table and missing primary key on scheme completion cache tables.
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.45.0...v1.46.0
- Perl
Published by kjolley about 2 years ago
bigsdb - Version 1.45.0
Version 1.45.0 introduces the following changes:
- An option has been added to highlight private data in the Excel file generated for isolate exports. If a user has access to private data (which may belong to others and been shared with them) there will be a section on the form that allows the user to select how these records are shaded within the output.
- An option has been added for an admin to create customized isolate query forms with pre-selected fields shown. New options will appear in the admin interface (with the Misc toggle selected):
Clicking the Add Query Interface link will allow a user to define new link text that will appear within the Search list on the contents page. Once this has been done, the Query Interface Fields links will become visible and this allows the admin to select any number of provenance, scheme, or LINcode fields to be pre-selected when the link is clicked.
- Finally, separate options have been added to force users to be logged in to download alleles or profiles when using the web interface. This is to encourage users to script downloads using the API. There are global settings in the bigsdb.conf file:
```
DOWNLOADS
Require users to be logged in to download records via web interface
Note that this is likely to prevent automated downloads (which generally
should be done via API). These values can be overridden in the database
config.xml file.
alleledownloadsrequirelogin=1
profiledownloadsrequirelogin=1
These can also be overridden on a per-database basis in the database config.xml file, e.g.
alleledownloadsrequirelogin="yes"
profiledownloadsrequirelogin="no"
```
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.44.1...v1.45.0
- Perl
Published by kjolley about 2 years ago
bigsdb - Version 1.44.1
Version 1.44.1 introduces the following changes:
- An option has been added to show if assembly checks have passed in the isolate results table. This can be enabled by default by setting
display_assembly_checks="yes"in the database config.xml file. It can also be set by end users in the user customization panels (https://bigsdb.readthedocs.io/en/latest/dataquery/0100options.html#main-results-table).
- An option has been added to scannew.pl to only scan records that have passed assembly checks.
- An option has also been added to not scan private records.
See https://bigsdb.readthedocs.io/en/latest/offline_tools.html#automated-offline-allele-definition for all scannew.pl options.
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.44.0...v1.44.1
- Perl
Published by kjolley about 2 years ago
bigsdb - Version 1.44.0
Version 1.44.0 introduces the following changes:
- New database indexes have been added to improve performance (thanks @braffes).
- Some rarely or never used indexes have been dropped.
- The option to set an allele designation with a status of 'ignore' has been removed. This was not really used but added an unnecessary filtering contraint on multiple queries.
- The number of isolates that match the current isolate at different LIN code thresholds are now displayed on an isolate record page (if LIN code has been assigned). The number is hyperlinked and takes you to a query that returns the matching isolates.
An administrator can also decide whether or not to include classification scheme or LIN code scheme matches on the isolate information page. These are enabled by default, if defined, but can be disabled with the following options in config.xml:
show_lincode_matches="no"
show_classification_schemes="no"
In addition, the number of LIN code thresholds to display can be set with the following option (replace the number with the required number of thresholds):
show_lincode_thresholds="5"
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.43.3...v1.44.0
- Perl
Published by kjolley about 2 years ago
bigsdb - Version 1.43.3
Version 1.43.3 brings the following changes:
- An optional message can be added on the allele and profile download pages to indicate that users should use the API for scripted data downloads. It is much easier to rate-limit the API than the web interface and it generally uses less resources to serve data. This message can be enabled by setting the URL for the API in bigsdb.conf, e.g.
rest_url=https://rest.pubmlst.org
- The JQuery.UI.multiselect widget is now used for dropdown boxes with large numbers of options, e.g. locus lists on the isolate query page, or where multiple options can be selected, such as the the 'includes' field list on various plugins. Where appropriate, the list is also searchable to make finding and selecting values quicker and easier.
- Javascript performance has been improved for background loading of isolate query form elements.
- Limits can now be set for the number of offline jobs that any single user can run for individual plugins. See the updated format of the job_limits.conf file found in the conf directory. If the limit is reached then further jobs for that particular plugin and user will be queued.
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.43.2...v1.43.3
- Perl
Published by kjolley about 2 years ago
bigsdb - Version 1.43.2
Version 1.43.2 brings the following changes:
- Additional logging information added. This includes the particular locus or scheme selected in data downloads, or the isolate record being shown.
- The cgST with fewest missing loci is now highlighted on the isolate info page when the isolate has multiple cgSTs.
- The query dashboard is now shown for bookmarked and publication queries.
- There is now an option to force users to log-in if they want to run offline jobs (see
jobs_require_login=1in bigsdb.conf). - There is now an option to automatically check the box to tag incomplete loci at specified identity thresholds (rather than just at 100% identity)
- 'Ambiguous read' flags are now automatically set on tags if sequence contains non-A,G,T,C characters.
- JQuery.UI.multiselect is now used for the flag dropdown box on the tag scan page. Previously we used an unformatted scrolling list box where an allele already had a flag but this was 4-5 lines long and made the table ugly and multiple flags difficult to select. For new tags there was a standard dropdown box which meant only one flag could be selected. As scan tables can be many thousands of rows long, the dropbox is only rendered when it is first visible in the current viewport.
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.43.1...v1.43.2
- Perl
Published by kjolley about 2 years ago
bigsdb - Version 1.43.1
Hotfix release to fix the isolate query modification trigger not appearing on the isolate query page (broken in commit d4b5439).
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.43.0...v1.43.1
- Perl
Published by kjolley over 2 years ago
bigsdb - Version 1.43.0
This version adds optional web logging to record user access. A new table log has been added to the bigsdbauth database that records IP address, username, and page called with a timestamp. In addition, the REST API logging to the bigsdbrest database has been improved by the addition of client and username information for when authenticated access is used.
To enable logging, you need to set the following in bigsdb.conf:
web_log_to_db=1
rest_log_to_db=1
Logging requires writing to the database on each page access so there is a very small performance penalty to enabling this. The tables are however unlogged (i.e. data are not written to the PosgreSQL write-ahead log) which makes them considerably faster than ordinary tables but data in them will be lost in the event of a database crash or unclean shutdown.
As every page access is recorded the log tables will grow in size over time. It is recommended that they are pruned regularly to remove records older than a specified period of time - this may also be required by GDPR! The easiest way to do this is to set up a scheduled CRON job by adding the following to /etc/crontab:
0 * * * * postgres psql -c "DELETE FROM log WHERE timestamp < NOW() - INTERVAL '7 days'" bigsdb_rest > /dev/null
10 * * * * postgres psql -c "DELETE FROM log WHERE timestamp < NOW() - INTERVAL '7 days'" bigsdb_auth > /dev/null
Additionally, there is a fix to the locus stats function in the sequence typing database necessitated by the introduction of schemes that can include locus presence/absence in their definitions. You therefore need to update the bigsdbrest, bigsdbauth, and any sequence definition database using the restv1.43.sql, authv1.43.sql, and seqdefdbv1.43.sql scripts respectively. See https://github.com/kjolley/BIGSdb/blob/develop/UpgradingREADME.txt.
Note that in the unlikely event that you have defined schemes that utilize locus presence/absence (where profiles can include the 'P' designation to indicate 'presence'), rather than just sequence variation, you should update the locus stats within the typing database with the following commands, otherwise the reported minimum allele length may be wrong.
DELETE FROM locus_stats;
INSERT INTO locus_stats(locus,datestamp,allele_count,min_length,max_length)
SELECT loci.id,MAX(sequences.datestamp),COUNT(sequences.allele_id),MIN(LENGTH(sequence)),MAX(LENGTH(sequence))
FROM loci LEFT JOIN sequences ON loci.id=sequences.locus
WHERE allele_id NOT IN ('N','0','P') OR allele_id IS NULL
GROUP BY loci.id;
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.42.7...v1.43.0
- Perl
Published by kjolley over 2 years ago
bigsdb - Version 1.42.7
This version:
- Adds a new Reports plugin - this is used to generate customized genome reports. See https://github.com/kjolley/Klebsiella_reports for examples of how to initiate and modify report templates.
- Adds an option to silently reject alleles that fail similarity check when batch uploading.
- Adds an option to make some fields expected for genome assemblies - set
required="genome_expected"in the field attribute in config.xml. - Makes the 'expand width' view persistent and allow its use for the front-end dashboard. This expands the width of page to that of the web browser and is useful if your results tables are very wide and you have a large monitor.
- Adds an option to return partial sequence matches for scheme queries using the API.
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.42.6...v1.42.7
- Perl
Published by kjolley over 2 years ago
bigsdb - Version 1.42.6
This release adds the following functionality and improvements:
- All tables can now be searched by a list of attributes. This has been possible for isolates, profiles and alleles (when using the specific-locus query) for a long time now, but it is now possible for example, to search for allele designations or tags based on a list of isolates or a list of loci.
- N50 gridlines have been added to the sequence bin cumulative length chart (https://github.com/kjolley/BIGSdb/commit/ff62aca47a4fea345b0852a224e3399f5107a8db).
- Multiple optimizations have been made to improve page load performance. This has largely focussed on the following:
- Reducing the number of database calls required by taking calls out of loops and instead retrieving batched results from the database in a single call and then caching these results (https://github.com/kjolley/BIGSdb/commit/b5998a43953ca259919155f0e1f9cb8b0d94fb72, https://github.com/kjolley/BIGSdb/commit/6131878f2c0dad49b8367c25a65e8591cce973fe, https://github.com/kjolley/BIGSdb/commit/e1ee2b64d2cf5ca3a7d5a0df21e969fef939d659, https://github.com/kjolley/BIGSdb/commit/96cc32127ebb3fe3817ccfb112864b6275dec10e).
- A more efficient SQL query has been made when generating the database view that filters the isolate table to include only records accessible by the current user, depending on their status and access to private records (https://github.com/kjolley/BIGSdb/commit/797a45a1d558b4f834833cb53203fdf4763e5fff).
- Removed the plugin manager isolate count check used to decide whether to list the plugin on the contents page (https://github.com/kjolley/BIGSdb/commit/e399d07055d3d96e33c70657e53124f3a18afd2a)
- Selected fields are now included in the ReporTree partitions summary. Any date field defined in the isolate table can now also be selected and used for the temporal component of cluster definitions.
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.42.5...v1.42.6
- Perl
Published by kjolley over 2 years ago
bigsdb - Version 1.42.5
This release introduces a new plugin that provides a wrapper for ReporTree (Mixão et al. 2023 Genome Med 15:43).
See https://bigsdb.readthedocs.io/en/latest/data_analysis/reportree.html for details.
- Perl
Published by kjolley over 2 years ago
bigsdb - Version 1.42.4
This release improves web-based sequence scanning by:
- showing the position of the first in-frame stop codon for loci defined as 'complete CDS'. The position has a coloured background that changes from red to green based on the position (red when it is at the 5' end, green when at the 3' end). This makes it easier for a curator to identify which matches are likely to be assignable as complete CDS alleles and/or whether they have internal stop codons other than near the 3' end of the sequence.
- adds an option to separate the search for start and stop codons. You can now choose to just look for in-frame stop codons that are within a specified percentage of the length of the best matching allele - either shorter or longer.
- Perl
Published by kjolley almost 3 years ago
bigsdb - Version 1.42.3
This version fixes an issue with exporting LINcode fields with isolate records.
- Perl
Published by kjolley almost 3 years ago
bigsdb - Version 1.41.2
This version includes the following: - Search isolates by SAVs or SNPs defined in typing database. - Use caching for dashboard AJAX calls where possible. - Add option to batch download full allele records via API - see https://bigsdb.readthedocs.io/en/latest/rest.html#db-loci-locus-alleles. - Fix for export of LINcode field values.
Full Changelog: https://github.com/kjolley/BIGSdb/compare/v1.42.1...v1.42.2
- Perl
Published by kjolley almost 3 years ago
bigsdb - Version 1.42.1
This introduces the following changes:
• Update to FontAwesome 6. • Provenance completeness metrics – these work like scheme annotation metrics, but you can define specific fields in config.xml as important for completeness (set annotationmetric=”yes” in the field definition). Note the new updateprovenance_metrics.pl script if you want to cache the metrics table (although it calculates on-the-fly and is pretty quick if not used). The score is simply a percentage of selected fields with a value (null values are excluded – see https://github.com/kjolley/BIGSdb/blob/develop/lib/BIGSdb/Constants.pm#LL296C1-L298C4 for definitions of these). • Isolates can now be searched by allele extended attributes defined in the typing database (https://bigsdb.readthedocs.io/en/latest/administration.html#defining-locus-extended-attributes).
- Perl
Published by kjolley about 3 years ago
bigsdb - Version 1.42.0
This version adds the option to define schemes that can include locus presence in profile definitions. This can be enabled by setting the 'allow_presence' flag in a scheme definition and then using the value 'P' for 'present' within a profile. Locus absence can be assigned with a '0'. Note that scheme caching must be enabled in isolate databases to make use of these gene presence schemes.
There is also a new single amino acid variant (SAAV) and single nucleotide polymorphism (SNP) detection tool. SAAVs and SNPs can be defined for a locus, and then alleles can be annotated by running the new scan_mutations.pl script. SAAV and SNPs will be displayed within allele records, and can be searched in locus-specific queries in the typing database.
Finally, security of cookies has been improved with -secure and -httponly attributes set. Note that to prevent breaking installations running over HTTP, to enable -secure you need to set secure_cookies=1 in bigsdb.conf.
- Perl
Published by kjolley about 3 years ago
bigsdb - Version 1.41.3
Version 1.41.3 adds a plugin to run Kleborate (https://github.com/katholt/Kleborate) to screen genome assemblies of Klebsiella pneumoniae and the Klebsiella pneumoniae species complex (KpSC) for:
- MLST sequence type
- species (e.g. K. pneumoniae, K. quasipneumoniae, K. variicola, etc.)
- ICEKp associated virulence loci: yersiniabactin (ybt), colibactin (clb), salmochelin (iro), hypermucoidy (rmpA)
- virulence plasmid associated loci: salmochelin (iro), aerobactin (iuc), hypermucoidy (rmpA, rmpA2)
- antimicrobial resistance determinants: acquired genes, SNPs, gene truncations and intrinsic β-lactamases
- K (capsule) and O antigen (LPS) serotype prediction, via wzi alleles and Kaptive

The output is a table in HTML, tab-delimited text, and Excel formats.

In addition, if the analysis is run with the --all option then the results are stored as an arbitrary analysis within the isolate database linked to the isolate record (this is the same way that we store the rMLST species identification in the screenshot below). A formatted display of these results is available in the isolate information page.

The analysis can also be run using a standalone script - update_kleborate.pl - found in the scripts/maintenance directory. This will run the analysis against all isolates in a database and populate the arbitrary analysis table for display.
Arbitrary analysis results are stored within the isolate database as a JSON object and can be displayed within isolate information pages using a template file stored in /etc/bigsdb/templates. Currently, this information is not searchable within the BIGSdb search interface, but the plan is to develop this so that specific fields can be registered for searching.
Kleborate needs to be installed on the system and the path to the executable file specifed in the BIGSdb global configuration file (/etc/bigsdb/bigsdb.conf), e.g.
kleborate_path=/usr/local/bin/kleborate
The plugin is not enabled by default, even on databases with allplugins="yes" set in the config.xml file, as it is only appropriate for use against _Klebsiella assemblies. To enable the plugin, set the following attribute within the config.xml <system> section for the database.
Kleborate="yes"
- Perl
Published by kjolley about 3 years ago
bigsdb - Version 1.41.2
This version adds a choice of colour palettes to front-end and query dashboards.
- Perl
Published by kjolley about 3 years ago
bigsdb - Version 1.41.1
Adds support for defining colours for specific field values in dashboard visualisations.
You can define colours for values by field using an additional configuration file called dashboard_colours.toml that can be placed either in /etc/bigsdb (for global use) or within a database configuration directory. The format is as follows:
'eav_Bexsero_reactivity' = {
'exact match' = '#2ca02c',
'cross-reactive' = '#ff7f0e',
'none' = '#d62728',
'insufficient data' = '#888888',
'No value' = '#aaaaaa'
}
'eav_Trumenba_reactivity' = {
'exact match' = '#2ca02c',
'cross-reactive' = '#ff7f0e',
'none' = '#d62728',
'insufficient data' = '#888888',
'No value' = '#aaaaaa'
}
's_1_clonal_complex' = {
'ST-11 complex' = 'yellow',
'ST-41/44 complex' = 'green'
}
Field names are prefixed as follows:
f_ Standard provenance fields, e.g. f_country
e_ Extended attribute fields, e.g. e_country||continent (continent attribute linked to country)
eav_ :ref:
Sparely-populated fields<sparsely_populated_fields>, e.g. eavBexseroreactivitys_ Scheme fields, e.g. s1clonal_complex (clonal complex field in scheme 1)
This works for pie, doughnut, bar, and pie charts. Note that if you define any values for a field then any value not defined will be shown as light grey in the visualisation.
- Perl
Published by kjolley over 3 years ago
bigsdb - Version 1.40.1
Version 1.40.1 adds the following:
- Checks, and adds if necessary, field indexes on scheme field cache tables.
- As the scheme cache tables are not now re-created from scratch, it is necessary to check that the fields and field types match those in the scheme definition in case they have changed. If there are any mismatches, the scheme field cache table is dropped and recreated with a full refresh.
- New index added to the missing_loci field of the scheme cache. This speeds up a query used once for every isolate during scheme cache renewal.
- Sequence export with flanking regions has been fixed.
- --curator option added to autotag.pl and scannew.pl scripts.
- Perl
Published by kjolley over 3 years ago
bigsdb - Version 1.40.0
Cache renewal now uses an embedded database stored procedure for scheme field lookup, improving memory use.
- Perl
Published by kjolley over 3 years ago
bigsdb - Version 1.39.1
This version bring Improvements to scheme caching in isolate databases. This is now more scalable for large databases. Manually refreshing the scheme is now done in a forked process with updates displayed using AJAX calls.
- Perl
Published by kjolley over 3 years ago
bigsdb - Version 1.39.0
This version introduces assembly submissions, allowing users to submit genome assemblies to add to existing records via the submission system. See https://bigsdb.readthedocs.io/en/latest/submissions.html#assembly-submission for details.
- Perl
Published by kjolley over 3 years ago
bigsdb - Version 1.38.1
This version adds front-end dashboards for user and public projects hosted within an isolate database. Previously, front-end dashboards were limited to summarising the complete database, but these now provide a landing page for individual projects. As with other dashboards, these are fully customisable by end users and can be specific for individual projects. Links within the dashboard and menu items lead to database views filtered by the current project.

In addition, the annotation status for any scheme can be displayed within a dashboard.

Custom dashboards can be created for any project by adding a TOML file in the database configuration named dashboardprojectX.toml (where X is the project id). The format for this file is described at https://bigsdb.readthedocs.io/en/latest/dbase_setup.html#defining-default-dashboards.
- Perl
Published by kjolley over 3 years ago
bigsdb - Version 1.38.0
This version adds support to limit submissions and curators to specific database configurations. This can be useful if two distinct datasets, with harmonised isolate fields, are hosted in the same database. Configurations can filter the dataset using database views and sets. Setting:
separate_database="yes"
in config.xml (or in the system.overrides file for a particular config) means that the configuration will be treated as though it was a separate database when it comes to notifying curators of submissions and allowing them to handle them. Curators can be limited to specific configurations by populating the new curator_configs table. Fields can now also be hidden from specific configurations by setting the 'hide' attribute to 'yes' in config.xml (or more usefully in the field.overrides file for a specific config). If a default field value is set, this value is automatically included in the record even if the field is hidden.
An example of where this may be useful is in the recent merging of the PubMLST S. aureus and S. epidermidis databases, so that analysis can be performed across both closely-related species. These appear as distinct when accessed from their respective species pages (https://pubmlst.org/organisms/staphylococcus-aureus and https://pubmlst.org/organisms/staphylococcus-epidermidis) and they each have their own curators. They are, however, also accessible as a merged dataset (https://pubmlst.org/organisms/staphylococcus-spp), where the species field is available to select between the two (this field is hidden in the specific species views).
In addition, the rMLST species id tool now provides text and Excel output files.
- Perl
Published by kjolley over 3 years ago
bigsdb - Version 1.36.5
This version adds new optional query dashboards to isolate databases. These visually summarise the results of a query.

The new query_dashboard attribute needs to be set within bigsdb.conf to enable these.
Support has also been added for GPS maps to be included within dashboards
 .
.
- Perl
Published by kjolley almost 4 years ago
bigsdb - Version 1.36.0
Version 1.36.0 allows isolate fields to be linked to GPS lookup tables to facilitate mapping. This allows, for example, a field for townorcity to contain place names, which can then be mapped within the isolate information page and in the Field Breakdown and Microreact plugins.

In order to enable this, the PostgreSQL PostGIS module needs to be installed and an optional table created in the isolates database using the isolatedbgeocoding.sql script. A field described in config.xml can then be linked to GPS lookups by setting the geographypoint_lookup attribute to 'yes'.
GPS lookup values can then be set in the curator interface by administrators or curators with the 'modify geopoints' permission set.


Alternatively, lookup GPS values can be set in bulk by using the gptownlookups.pl script found in the scripts/maintenance directory. This uses the Geonames database, containing coordinates of town and cities with populations of at least 1000 worldwide, that is distributed with BIGSdb.
- Perl
Published by kjolley almost 4 years ago
bigsdb - Version 1.35.3
Version 1.35.3 improves on the GPS mapping available in the Field Breakdown plugin.
Users can now dynamically change pointer colour and size, and the map is automatically centred and zoomed on the displayed points.

In addition, a race condition was fixed that could occasionally mix up visualisation controls if users rapidly switched between country and GPS fields before the current visualisation was fully loaded.
- Perl
Published by kjolley about 4 years ago
bigsdb - Version 1.35.2
Version 1.35.2 adds support for a new isolate field type to natively support GPS coordinates for use in mapping. A map will appear within the isolate record information page if such a field is populated. By default, OpenStreetMap is used, but Bing maps can be used if a Bing Map API key is set within the bigsdb.conf file. Bing maps allow you to toggle between map and aerial views.


In addition, record locations are mapped within the Field Breakdown plugin.

You can also use the field as the source for mapping information when exporting to Microreact so that sub-country level mapping can now be used.
The field type 'geography_point' needs to be set as the type attribute for the field within the database config.xml file.
In order to create the field within the isolate table you will need to install the PostGIS module for PostgreSQL and enable this within the isolate database with the following:
CREATE EXTENSION postgis;
and then add the field as below, in this case for a field called location:
ALTER TABLE isolates ADD location geography(POINT,4326);
[SRID 4326 represents spatial data using longitude and latitude coordinates on the Earth’s surface.]
Geographic data are entered within the curation interface as a single string in the format '[latitude], [longitude]'. The latitude and longitude are individually queryable when performing an isolate search.
If you already have a database that contains separate latitude and longitude data as float values, you can use these to populate a new location field as follows:
UPDATE isolates SET location = POINT(longitude,latitude)::geometry WHERE latitude IS NOT NULL AND longitude IS NOT NULL;
- Perl
Published by kjolley about 4 years ago
bigsdb - Version 1.35.0
Version 1.35.0 adds support for alternative codon tables. Databases can be set to use a particular codon table, as defined at https://www.ncbi.nlm.nih.gov/Taxonomy/Utils/wprintgc.cgi. This can be done by setting the codontable attribute in config.xml to the appropriate table id number. In addition, if the alternativecodon_tables attribute is set to 'yes', individual isolates can be set with different codon tables. This would be appropriate for databases that contain records of different bacterial species that may use different codon tables.
The selected codon table is used for translations and for checks that involve determining if a sequence is a complete coding sequence (has a start/stop codon in frame, with no internal stop codons).
- Perl
Published by kjolley about 4 years ago
bigsdb - Version 1.34.0
This version provides support for LINcodes based on cgMLST schemes. See https://doi.org/10.1101/2021.07.26.453808 for more information about LINcodes.
Documentation for using setting up LINcodes within BIGSdb can be found at https://bigsdb.readthedocs.io/en/latest/administration.html#setting-up-lincode-definitions-for-cgmlst-schemes.
- Perl
Published by kjolley about 4 years ago
bigsdb - Version 1.33.0
This adds data drill-down functionality linked to the front-end dashboards on isolate databases.
- Perl
Published by kjolley over 4 years ago
bigsdb - Version 1.32.1
This version updates the Microreact plugin to support breaking changes introduced on the Microreact website. API uploads to Microreact now require an account token - this needs to be set in the bigsdb.conf file.
In addition, it is no longer a requirement for databases to have fields named 'country' and 'year' in order for the plugin to be enabled. Fields with different names can be used and set within the config.xml file using the microreactcountryfield and microreactyearfield attributes in the system tag. Note that countries should use the standardised list of names as defined in BIGSdb since these are mapped internally to ISO 3166-2 2 letter country codes used by Microreact. The easiest way to use these is to define a country field in the config.xml as below:
<field type="text" required="yes" maindisplay="yes" comments="country where strain was isolated" optlist="yes" values="COUNTRIES" sort="no">country
<optlist>
<option>Unknown</option>
</optlist>
</field>
If either a country or year field is not used, data are still uploaded to Microreact but either the map or timeline is not shown.
- Perl
Published by kjolley over 4 years ago
bigsdb - Version 1.32.0
This version adds optional front-end dashboards to isolate databases.
Dashboards are fully customisable by end users so you can select the fields and visualisations that you want to see. Modify the layout by dragging elements or change their size. Any changes made are remembered between sessions. A default dashboard can be set for all databases on the system, and overridden for individual database configurations. See https://bigsdb.readthedocs.io/en/latest/dbase_setup.html#setting-up-front-end-dashboards for details on setting up.
In addition, alternative start codons, e.g. CTG, ATT, can now be set on a per-database or a per-locus basis.
- Perl
Published by kjolley over 4 years ago
bigsdb - Version 1.31.5
Includes classification scheme information in profile results returned by the RESTful API.
- Perl
Published by kjolley over 4 years ago
bigsdb - Version 1.31.4
This version changes the way that classification schemes are made available via the API. This is now more consistent with the rest of the interface.
- Perl
Published by kjolley over 4 years ago
bigsdb - Version 1.31.1
In this version:
- Added assembly checks - results of these can be queried and used for filtering of isolate datasets.
- Added announcement banner to database front page. The contents of /etc/bigsdb/announcement.html will be shown on all database front pages - this is useful for service announcements, e.g. scheduled maintenance.
- Upgraded D3 to v6, and changed charting library from c3.js to billboard.js.
- Added means of grouping standard isolate fields into categories. Separate groups are listed in dropdown boxes in queries and plugins, and on the isolate display page.
- Perl
Published by kjolley almost 5 years ago
bigsdb - Version 1.31.0.
This version adds new functionality for assessing and filtering results based on genome assembly metrics.
N50 and L50 values are now stored and can be used in isolate queries. The stats are automatically updated by a database trigger whenever, and however, contigs are added to the sequence bin.
%GC, number of Ns, and gaps in an assembly can be calculated and stored by an external script after assembly uploads. These can be used along with N50, total length and number of contigs in isolate queries.
The results of arbitrary analyses can now be stored in the isolate database as JSON values. These can be displayed within an isolate record using a templating system. rMLST species checks have been implemented as an exemplar.
Checks are now performed on submissions prior to upload to the database. Basic assembly stats are available to submitters and curators with automated warnings shown based on customisable thresholds. Hard limits can also be set so that submissions can be rejected automatically.
We can also run the rMLST species check on submitted genomes so that the results are available to curators shortly after submission and prior to handling.
Batch uploading data is now performed in a separate process, with progress notifications loaded by AJAX polling. This prevents browser timeout when hundreds of genomes are uploaded together.
- Perl
Published by kjolley about 5 years ago
bigsdb - Version 1.30.1.
This version adds an option to limit the number of running jobs for particular plugins.
- Perl
Published by kjolley over 5 years ago
bigsdb - Version 1.30.0.
This version introduces annotation status based on scheme completion.
For a well-annotated genome it can be expected that all the loci, or a specified threshold number of loci, for some schemes will have allele designations. We can use this annotation status metric to help assess genome quality. Individual schemes can be set to be used for quality metrics and threshold numbers of loci that indicate a 'good' or 'bad' annotation based on that scheme can be defined in the schemes table.
The metric can be used in searches to filter datasets that have passed annotation criteria.
- Perl
Published by kjolley over 5 years ago
bigsdb - Version 1.29.5.
This release provides minor bug fixes.
- Perl
Published by kjolley over 5 years ago
bigsdb - Version 1.29.4
This release provides support for uploading compressed files for sequence queries or genome submissions. Sequence files can be in either uncompressed, or gzip/zip compressed FASTA format.
- Perl
Published by kjolley over 5 years ago
bigsdb - Version 1.29.3
This version:
- adds the option to include sequence bin size and the number of contigs within an isolate query.
- improves database connection handling to try to minimise the number of open connections.
- adds the option for a curator to deny a user's request to publish private data records and automatically notifies users of their request outcome.
- Perl
Published by kjolley over 5 years ago
bigsdb - Version 1.29.2
This version sees an upgraded rMLST species id plugin. This is now multi-threaded and works much quicker. In addition to scanning genomes in a local database, it also allows the use of pre-tagged rMLST allele designations which is even quicker.
- Perl
Published by kjolley over 5 years ago
bigsdb - Version 1.29.1
- Isolate fields can be either required, optional and now 'expected'. Expected fields require a submitter to explicitly state if there is no value available by entering 'null'. For use when you want a submitter to always include a value for a field but it is not always possible.
- Fix for tooltip positioning in GenePresence heatmap.
- Perl
Published by kjolley over 5 years ago
bigsdb - Version 1.29.0
- Provides curators with site-wide accounts options to modify how they receive submission notifications.
- Digest mode - E-mail notifications are grouped and sent as a single notification at minimum intervals set by the user. Minimum interval times can be set between 1 and 24 hours.
- Holiday mode - Curators can mark themselves absent for a period of up to 3 months where they will no longer receive notifications.
- Perl
Published by kjolley over 5 years ago
bigsdb - Version 1.28.1
- Update to the ITOL plugin to support the use of an API key which is now necessary due to a change at https://itol.embl.de/ to a subscription model.
- If a new allele is found during a sequence query, the allele sequence is now extracted with the option to start a new submission for assignment.
- Perl
Published by kjolley over 5 years ago
bigsdb - Version 1.28.0
This version has an updated look to support the new design of PubMLST.
- Perl
Published by kjolley over 5 years ago
bigsdb - Version 1.27.1
- Improved kiosk mode support.
- Fixed memory leak in BLAST plugin.
- Perl
Published by kjolley almost 6 years ago
bigsdb - Version 1.27.0
- Support for different locus categories - these show on different tracks on IGV genome viewer.
- Improved formatting of large field values.
- Improved handling of sparse field categories.
- Perl
Published by kjolley almost 6 years ago
bigsdb - BIGSdb version 1.20.1
- Improved exception handling - please note new Perl module dependencies: Try::Tiny and Exception::Class.
- Updated GrapeTree plugin reflects upstream filename and data formatting changes (see attributes in bigsdb.conf to enable).
- Perl
Published by kjolley over 7 years ago
bigsdb - BIGSdb version 1.19.1
This repository contains the BIGSdb platform described in Open-access bacterial population genomics: BIGSdb software, the PubMLST.org website and their applications, Jolley et al. 2018, submitted to Wellcome Open Research.
- Perl
Published by kjolley over 7 years ago