Science Score: 36.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
○CITATION.cff file
-
✓codemeta.json file
Found codemeta.json file -
✓.zenodo.json file
Found .zenodo.json file -
○DOI references
-
○Academic publication links
-
✓Committers with academic emails
2 of 21 committers (9.5%) from academic institutions -
○Institutional organization owner
-
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (12.3%) to scientific vocabulary
Keywords
Keywords from Contributors
Repository
Library for Digital Pathology Image Processing
Basic Info
- Host: GitHub
- Owner: histolab
- License: apache-2.0
- Language: Python
- Default Branch: main
- Homepage: http://histolab.readthedocs.io
- Size: 365 MB
Statistics
- Stars: 422
- Watchers: 6
- Forks: 62
- Open Issues: 39
- Releases: 17
Topics
Metadata Files
README.md


| Test Status |
|
| Code Quality |


|
| Version Info |



|
| License |

|
| Documentation |
Table of Contents
- Motivation
- Quickstart
- Versioning
- Authors
- License
- Roadmap
- Acknowledgements
- References
- Contribution guidelines
Motivation
The histo-pathological analysis of tissue sections is the gold standard to assess the presence of many complex diseases, such as tumors, and understand their nature. In daily practice, pathologists usually perform microscopy examination of tissue slides considering a limited number of regions and the clinical evaluation relies on several factors such as nuclei morphology, cell distribution, and color (staining): this process is time consuming, could lead to information loss, and suffers from inter-observer variability.
The advent of digital pathology is changing the way pathologists work and collaborate, and has opened the way to a new era in computational pathology. In particular, histopathology is expected to be at the center of the AI revolution in medicine [1], prevision supported by the increasing success of deep learning applications to digital pathology.
Whole Slide Images (WSIs), namely the translation of tissue slides from glass to digital format, are a great source of information from both a medical and a computational point of view. WSIs can be coloured with different staining techniques (e.g. H&E or IHC), and are usually very large in size (up to several GB per slide). Because of WSIs typical pyramidal structure, images can be retrieved at different magnification factors, providing a further layer of information beyond color.
However, processing WSIs is far from being trivial. First of all, WSIs can be stored in different proprietary formats, according to the scanner used to digitalize the slides, and a standard protocol is still missing. WSIs can also present artifacts, such as shadows, mold, or annotations (pen marks) that are not useful. Moreover, giving their dimensions, it is not possible to process a WSI all at once, or, for example, to feed a neural network: it is necessary to crop smaller regions of tissues (tiles), which in turns require a tissue detection step.
The aim of this project is to provide a tool for WSI processing in a reproducible environment to support clinical and scientific research. histolab is designed to handle WSIs, automatically detect the tissue, and retrieve informative tiles, and it can thus be integrated in a deep learning pipeline.
Getting Started
Prerequisites
Please see installation instructions.
Documentation
Read the full documentation here https://histolab.readthedocs.io/en/latest/.
Communication
Join our user group on  Slack
Slack
5 minutes introduction
Quickstart
Here we present a step-by-step tutorial on the use of histolab to
extract a tile dataset from example WSIs. The corresponding Jupyter
Notebook is available at https://github.com/histolab/histolab-box:
this repository contains a complete histolab environment that can be
used through Docker on all platforms.
Thus, the user can decide either to use histolab through
histolab-box or installing it in his/her python virtual environment
(using conda, pipenv, pyenv, virtualenv, etc...). In the latter case, as
the histolab package has been published on (PyPi),
it can be easily installed via the command:
shell
pip install histolab
alternatively, it can be installed via conda:
shell
conda install -c conda-forge histolab
TCGA data
First things first, lets import some data to work with, for example the
prostate tissue slide and the ovarian tissue slide available in the
data module:
python
from histolab.data import prostate_tissue, ovarian_tissue
Note: To use the data module, you need to install pooch, also
available on PyPI (https://pypi.org/project/pooch/). This step is
needless if we are using the Vagrant/Docker virtual environment.
The calling to a data function will automatically download the WSI
from the corresponding repository and save the slide in a cached
directory:
python
prostate_svs, prostate_path = prostate_tissue()
ovarian_svs, ovarian_path = ovarian_tissue()
Notice that each data function outputs the corresponding slide, as an
OpenSlide object, and the path where the slide has been saved.
Slide initialization
histolab maps a WSI file into a Slide object. Each usage of a WSI
requires a 1-o-1 association with a Slide object contained in the
slide module:
python
from histolab.slide import Slide
To initialize a Slide it is necessary to specify the WSI path, and the
processed_path where the tiles will be saved. In our
example, we want the processed_path of each slide to be a subfolder of
the current working directory:
```python import os
BASE_PATH = os.getcwd()
PROCESSPATHPROSTATE = os.path.join(BASEPATH, 'prostate', 'processed') PROCESSPATHOVARIAN = os.path.join(BASEPATH, 'ovarian', 'processed')
prostateslide = Slide(prostatepath, processedpath=PROCESSPATHPROSTATE) ovarianslide = Slide(ovarianpath, processedpath=PROCESSPATHOVARIAN) ```
Note: If the slides were stored in the same folder, this can be done
directly on the whole dataset by using the SlideSet object of the
slide module.
With a Slide object we can easily retrieve information about the
slide, such as the slide name, the number of available levels, the
dimensions at native magnification or at a specified level:
python
print(f"Slide name: {prostate_slide.name}")
print(f"Levels: {prostate_slide.levels}")
print(f"Dimensions at level 0: {prostate_slide.dimensions}")
print(f"Dimensions at level 1: {prostate_slide.level_dimensions(level=1)}")
print(f"Dimensions at level 2: {prostate_slide.level_dimensions(level=2)}")
shell
Slide name: 6b725022-f1d5-4672-8c6c-de8140345210
Levels: [0, 1, 2]
Dimensions at level 0: (16000, 15316)
Dimensions at level 1: (4000, 3829)
Dimensions at level 2: (2000, 1914)
python
print(f"Slide name: {ovarian_slide.name}")
print(f"Levels: {ovarian_slide.levels}")
print(f"Dimensions at level 0: {ovarian_slide.dimensions}")
print(f"Dimensions at level 1: {ovarian_slide.level_dimensions(level=1)}")
print(f"Dimensions at level 2: {ovarian_slide.level_dimensions(level=2)}")
shell
Slide name: b777ec99-2811-4aa4-9568-13f68e380c86
Levels: [0, 1, 2]
Dimensions at level 0: (30001, 33987)
Dimensions at level 1: (7500, 8496)
Dimensions at level 2: (1875, 2124)
Note:
If the native magnification, i.e., the magnification factor used to scan the slide, is provided in the slide properties, it is also possible
to convert the desired level to its corresponding magnification factor with the level_magnification_factor property.
```python print( "Native magnification factor:", prostateslide.levelmagnification_factor() )
print(
"Magnification factor corresponding to level 1:",
prostate_slide.level_magnification_factor(level=1),
)
```
shell
Native magnification factor: 20X
Magnification factor corresponding to level 1: 5.0X
Moreover, we can retrieve or show the slide thumbnail in a separate window:
python
prostate_slide.thumbnail
prostate_slide.show()
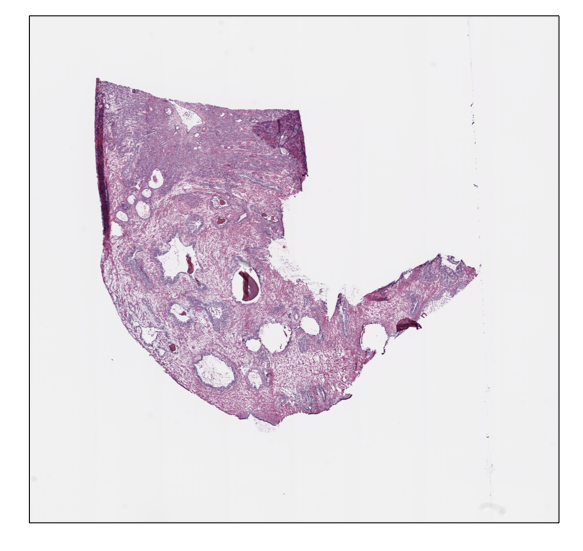
python
ovarian_slide.thumbnail
ovarian_slide.show()
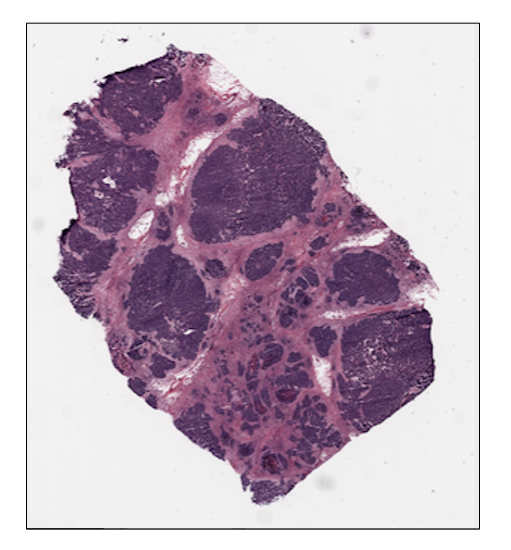
Tile extraction
Once that the Slide objects are defined, we can proceed to extract the
tiles. To speed up the extraction process, histolab automatically
detects the tissue region with the largest connected area and crops the
tiles within this field. The tiler module implements different
strategies for the tiles extraction and provides an intuitive interface
to easily retrieve a tile dataset suitable for our task. In particular,
each extraction method is customizable with several common parameters:
tile_size: the tile size;level: the extraction level (from 0 to the number of available levels);check_tissue: if a minimum percentage of tissue is required to save the tiles;tissue_percent: number between 0.0 and 100.0 representing the minimum required percentage of tissue over the total area of the image (default is 80.0);prefix: a prefix to be added at the beginning of the tiles filename (default is the empty string);suffix: a suffix to be added to the end of the tiles filename (default is.png).
Random Extraction
The simplest approach we may adopt is to randomly crop a fixed number of
tiles from our slides; in this case, we need the RandomTiler
extractor:
python
from histolab.tiler import RandomTiler
Let us suppose that we want to randomly extract 30 squared tiles at level
2 of size 128 from our prostate slide, and that we want to save them only
if they have at least 80% of tissue inside. We then initialize our
RandomTiler extractor as follows:
python
random_tiles_extractor = RandomTiler(
tile_size=(128, 128),
n_tiles=30,
level=2,
seed=42,
check_tissue=True, # default
tissue_percent=80.0, # default
prefix="random/", # save tiles in the "random" subdirectory of slide's processed_path
suffix=".png" # default
)
Notice that we also specify the random seed to ensure the reproducibility of the extraction process.
We may want to check which tiles have been selected by the tiler, before starting the extraction procedure and saving them;
the locate_tiles method of RandomTiler returns a scaled version of the slide with the corresponding tiles outlined. It is also possible to specify
the transparency of the background slide, and the color used for the border of each tile:
python
random_tiles_extractor.locate_tiles(
slide=prostate_slide,
scale_factor=24, # default
alpha=128, # default
outline="red", # default
)

Starting the extraction is then as simple as calling the extract method on the extractor, passing the
slide as parameter:
python
random_tiles_extractor.extract(prostate_slide)

Random tiles extracted from the prostate slide at level 2.
Grid Extraction
Instead of picking tiles at random, we may want to retrieve all the tiles available. The Grid Tiler extractor crops the tiles following a grid structure on the largest tissue region detected in the WSI:
python
from histolab.tiler import GridTiler
In our example, we want to extract squared tiles at level 0 of size 512
from our ovarian slide, independently of the amount of tissue detected.
By default, tiles will not overlap, namely the parameter defining the
number of overlapping pixels between two adjacent tiles,
pixel_overlap, is set to zero:
python
grid_tiles_extractor = GridTiler(
tile_size=(512, 512),
level=0,
check_tissue=False,
pixel_overlap=0, # default
prefix="grid/", # save tiles in the "grid" subdirectory of slide's processed_path
suffix=".png" # default
)
Again, we can exploit the locate_tiles method to visualize the selected tiles on a scaled version of the slide:
python
grid_tiles_extractor.locate_tiles(
slide=ovarian_slide,
scale_factor=64,
alpha=64,
outline="#046C4C",
)

python
grid_tiles_extractor.extract(ovarian_slide)
and the extraction process starts when the extract method is called on our extractor:

Examples of non-overlapping grid tiles extracted from the ovarian slide at level 0.
Score-based extraction
Depending on the task we will use our tile dataset for, the extracted
tiles may not be equally informative. The ScoreTiler allows us to save
only the "best" tiles, among all the ones extracted with a grid
structure, based on a specific scoring function. For example, let us
suppose that our goal is the detection of mitotic activity on our
ovarian slide. In this case, tiles with a higher presence of nuclei are
preferable over tiles with few or no nuclei. We can leverage the
NucleiScorer function of the scorer module to order the extracted
tiles based on the proportion of the tissue and of the hematoxylin
staining. In particular, the score is computed as where
is the percentage of nuclei and
the percentage of tissue in the tile t
First, we need the extractor and the scorer:
python
from histolab.tiler import ScoreTiler
from histolab.scorer import NucleiScorer
As the ScoreTiler extends the GridTiler extractor, we also set the
pixel_overlap as additional parameter. Moreover, we can specify the
number of the top tiles we want to save with the n_tile parameter:
python
scored_tiles_extractor = ScoreTiler(
scorer = NucleiScorer(),
tile_size=(512, 512),
n_tiles=100,
level=0,
check_tissue=True,
tissue_percent=80.0,
pixel_overlap=0, # default
prefix="scored/", # save tiles in the "scored" subdirectory of slide's processed_path
suffix=".png" # default
)
Notice that also the ScoreTiler implements the locate_tiles method, which visualizes (on a scaled version of the slide) the first n_tiles with the highest scores:
python
grid_tiles_extractor.locate_tiles(slide=ovarian_slide)

Finally, when we extract our cropped images, we can also write a report of the saved tiles and their scores in a CSV file:
```python summaryfilename = "summaryovariantiles.csv" SUMMARYPATH = os.path.join(ovarianslide.processedpath, summary_filename)
scoredtilesextractor.extract(ovarianslide, reportpath=SUMMARY_PATH) ```

Representation of the score assigned to each extracted tile by the
NucleiScorer, based on the amount of nuclei detected.
Versioning
We use PEP 440 for versioning.
Authors
License
This project is licensed under Apache License Version 2.0 - see the LICENSE.txt file for details
Roadmap
Acknowledgements
References
[1] Colling, Richard, et al. "Artificial intelligence in digital pathology: A roadmap to routine use in clinical practice." The Journal of pathology 249.2 (2019)
Contribution guidelines
If you want to contribute to histolab, be sure to review the contribution guidelines
Owner
- Name: histolab
- Login: histolab
- Kind: organization
- Repositories: 2
- Profile: https://github.com/histolab
GitHub Events
Total
- Issues event: 3
- Watch event: 47
- Delete event: 67
- Issue comment event: 83
- Push event: 254
- Pull request review comment event: 5
- Pull request review event: 45
- Pull request event: 132
- Fork event: 4
- Create event: 67
Last Year
- Issues event: 3
- Watch event: 47
- Delete event: 67
- Issue comment event: 83
- Push event: 254
- Pull request review comment event: 5
- Pull request review event: 45
- Pull request event: 132
- Fork event: 4
- Create event: 67
Committers
Last synced: over 1 year ago
Top Committers
| Name | Commits | |
|---|---|---|
| ernestoarbitrio | e****o@g****m | 457 |
| alessiamarcolini | 9****i@g****m | 451 |
| dependabot[bot] | 4****] | 182 |
| Alessia Marcolini | a****i@f****u | 86 |
| Nicole Bussola | n****i@g****m | 86 |
| kheffah | m****d@e****u | 65 |
| Nicole Bussola | n****i@g****m | 45 |
| pre-commit-ci[bot] | 6****] | 28 |
| Nicole Bussola | b****a@f****u | 17 |
| Etty | b****r@g****m | 5 |
| nicolebussola | 3****a | 4 |
| Marco Burro | m****8@g****m | 3 |
| Nicole Bussola | n****e@N****l | 3 |
| Patrick Arminio | p****o@g****m | 3 |
| Christopher Gundler | c****r@g****e | 2 |
| Nicole Bussola | n****e@n****t | 2 |
| leriomaggio | v****o@g****m | 2 |
| BilGuet | b****i@l****t | 1 |
| dependabot-preview[bot] | 2****] | 1 |
| Christopher Gundler | c****r@u****e | 1 |
| nipeone | o****d@1****m | 1 |
Committer Domains (Top 20 + Academic)
Issues and Pull Requests
Last synced: 8 months ago
All Time
- Total issues: 31
- Total pull requests: 299
- Average time to close issues: 8 months
- Average time to close pull requests: 21 days
- Total issue authors: 22
- Total pull request authors: 9
- Average comments per issue: 2.35
- Average comments per pull request: 1.31
- Merged pull requests: 179
- Bot issues: 0
- Bot pull requests: 252
Past Year
- Issues: 2
- Pull requests: 73
- Average time to close issues: about 5 hours
- Average time to close pull requests: 4 days
- Issue authors: 2
- Pull request authors: 4
- Average comments per issue: 0.5
- Average comments per pull request: 0.68
- Merged pull requests: 39
- Bot issues: 0
- Bot pull requests: 66
Top Authors
Issue Authors
- alessiamarcolini (5)
- neoMerz (2)
- Neuro-nerd-scientist (2)
- bguetarni (2)
- CaiYitao (2)
- qasimgilani (2)
- realHongYuZhou (1)
- rahit (1)
- delta2golf (1)
- suke18 (1)
- xuanblo (1)
- rongyua (1)
- explainable-ai (1)
- a1ecbennington (1)
- gakabani (1)
Pull Request Authors
- dependabot[bot] (201)
- pre-commit-ci[bot] (51)
- alessiamarcolini (21)
- ernestoarbitrio (18)
- nicolebussola (4)
- erich-r (1)
- bguetarni (1)
- patrick91 (1)
- ajinkya-kulkarni (1)
Top Labels
Issue Labels
Pull Request Labels
Packages
- Total packages: 2
-
Total downloads:
- pypi 1,272 last-month
-
Total dependent packages: 1
(may contain duplicates) -
Total dependent repositories: 2
(may contain duplicates) - Total versions: 21
- Total maintainers: 3
pypi.org: histolab
Python library for Digital Pathology Image Processing
- Homepage: https://github.com/histolab/histolab
- Documentation: https://histolab.readthedocs.io
- License: Apache-2.0
-
Latest release: 0.7.0
published about 2 years ago
Rankings
Maintainers (3)
conda-forge.org: histolab
- Homepage: https://github.com/histolab/histolab
- License: Apache-2.0
-
Latest release: 0.5.1
published almost 4 years ago
Rankings
Dependencies
- IPython *
- histolab *
- sphinx ==4.5.0
- sphinx-prompt *
- sphinx-rtd-theme *
- sphinxcontrib-katex *
- sphinxemoji *
- 122 dependencies
- Sphinx ^5.1.1 develop
- bandit ^1.7.1 develop
- black ^22.6.0 develop
- coverage ^6.4.4 develop
- flake8 * develop
- ipdb ^0.13.9 develop
- isort ^5.10.1 develop
- large-image >=1.8.11,<1.8.12 develop
- large-image-source-openslide >=1.8.11,<1.13.1 develop
- large-image-source-pil >=1.8.11,<1.13.1 develop
- pooch ^1.5.2 develop
- pre-commit ^2.15.0 develop
- pycodestyle ^2.9.1 develop
- pyflakes ^2.5.0 develop
- pytest ^7.1.2 develop
- pytest-benchmark ^3.4.1 develop
- pytest-cov ^3.0.0 develop
- pytest-html ^3.1.1 develop
- pytest-xdist ^2.4.0 develop
- sphinx-prompt ^1.5.0 develop
- sphinx-rtd-theme ^1.0.0 develop
- sphinxcontrib-katex >=0.8.6,<0.10.0 develop
- sphinxemoji ^0.2.0 develop
- toml ^0.10.2 develop
- twine ^4.0.1 develop
- Pillow >=9.1.0,<10.0.0
- Sphinx ^5.1.1
- importlib-metadata ^4.12.0
- numpy >=1.18.4,<1.23.1
- openslide-python >=1.1.2, <1.2.1
- python >=3.7,<3.11
- scikit-image >=0.19.0,<0.19.3
- scipy >=1.5.0,<1.8.2
- sphinx-prompt ^1.5.0
- sphinx-rtd-theme ^1.0.0
- sphinxcontrib-katex >=0.8.6,<0.10.0
- sphinxemoji ^0.2.0
- typing-extensions ^4.0.0
- actions/cache v1 composite
- actions/checkout v2 composite
- actions/setup-python v2 composite
- rhysd/github-action-benchmark v1 composite
- actions/checkout v3 composite
- github/codeql-action/analyze v2 composite
- github/codeql-action/autobuild v2 composite
- github/codeql-action/init v2 composite
- actions/checkout v3 composite
- actions/setup-python v3 composite
- act10ns/slack v1 composite
- actions/cache v3 composite
- actions/checkout v3 composite
- actions/download-artifact v2 composite
- actions/setup-python v4 composite
- actions/upload-artifact v3 composite
- codecov/codecov-action v3 composite
- geekyeggo/delete-artifact v1 composite
- msys2/setup-msys2 v2 composite
