instanovo
De novo peptide sequencing with InstaNovo: Accurate, database-free peptide identification for large scale proteomics experiments
Science Score: 67.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
✓CITATION.cff file
Found CITATION.cff file -
✓codemeta.json file
Found codemeta.json file -
✓.zenodo.json file
Found .zenodo.json file -
✓DOI references
Found 6 DOI reference(s) in README -
✓Academic publication links
Links to: nature.com, zenodo.org -
○Committers with academic emails
-
○Institutional organization owner
-
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (12.0%) to scientific vocabulary
Keywords from Contributors
Repository
De novo peptide sequencing with InstaNovo: Accurate, database-free peptide identification for large scale proteomics experiments
Basic Info
Statistics
- Stars: 97
- Watchers: 8
- Forks: 23
- Open Issues: 9
- Releases: 15
Metadata Files
README.md
De novo peptide sequencing with InstaNovo
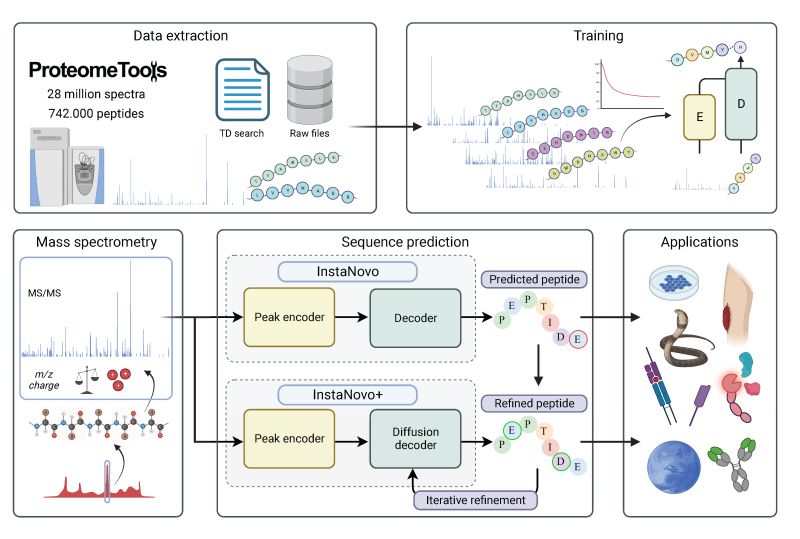
The official code repository for InstaNovo. This repo contains the code for training and inference of InstaNovo and InstaNovo+. InstaNovo is a transformer neural network with the ability to translate fragment ion peaks into the sequence of amino acids that make up the studied peptide(s). InstaNovo+, inspired by human intuition, is a multinomial diffusion model that further improves performance by iterative refinement of predicted sequences.
Links:
- Publication in Nature Machine Intelligence: InstaNovo enables diffusion-powered de novo peptide sequencing in large-scale proteomics experiments
- InstaNovo blog: https://instanovo.ai/
- Documentation: https://instadeepai.github.io/InstaNovo/
Developed by:
Usage
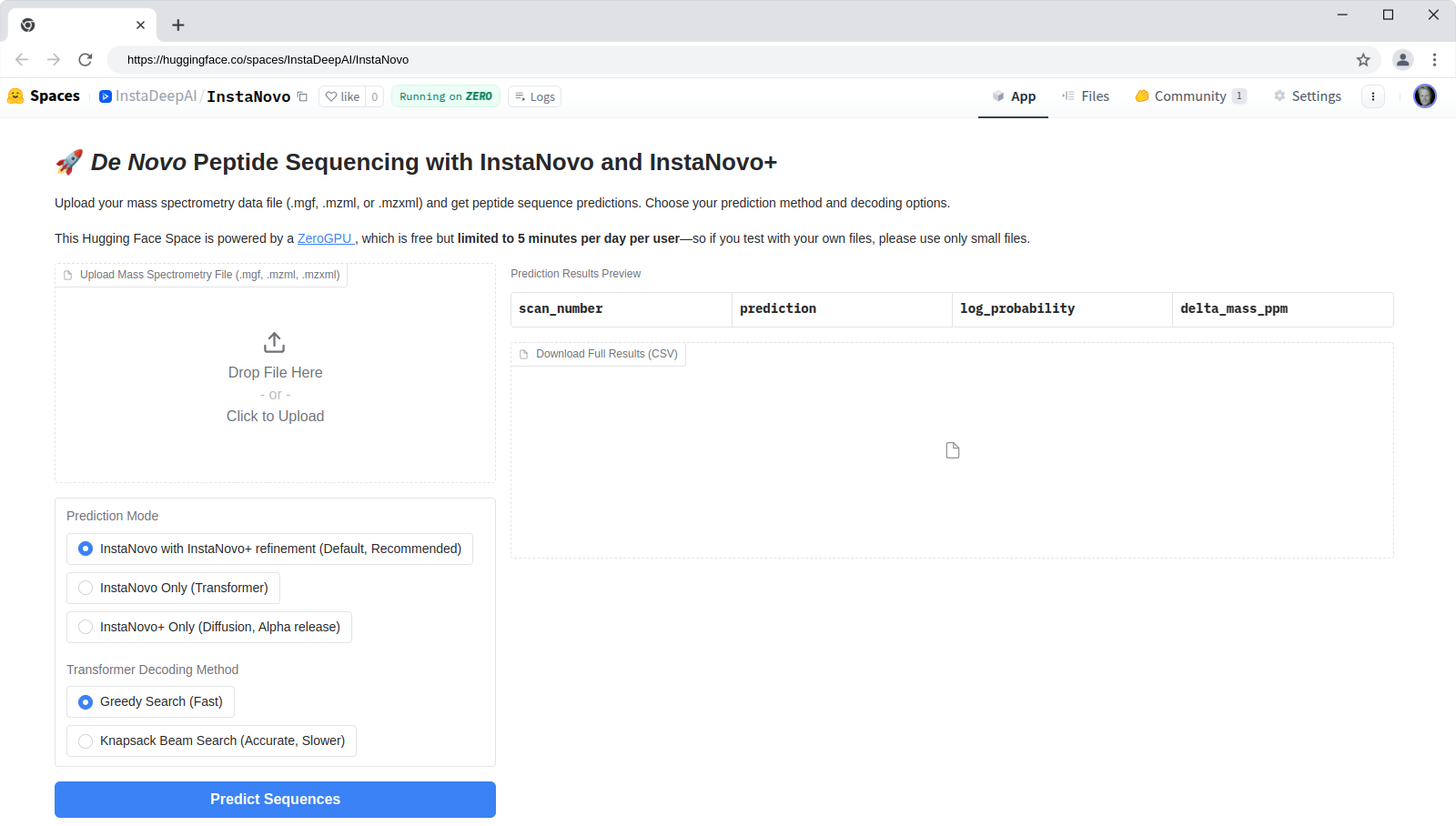
HuggingFace Space
InstaNovo is available as a HuggingFace Space at
hf.co/spaces/InstaDeepAI/InstaNovo for quick
testing and evaluation. You can upload your own spectra files in .mgf, .mzml, or .mzxml format
and run de novo predictions. The results will be displayed in a table format, and you can download
the predictions as a CSV file. The HuggingFace Space is powered by the InstaNovo model and the
InstaNovo+ model for iterative refinement.
Installation
To use InstaNovo Python package with command line interface, we need to install the module via
pip:
bash
pip install instanovo
If you have access to an NVIDIA GPU, you can install InstaNovo with the GPU version of PyTorch (recommended):
bash
pip install "instanovo[cu124]"
If you are on macOS, you can install the CPU-only version of PyTorch:
bash
pip install "instanovo[cpu]"
Command line usage
InstaNovo provides a comprehensive command line interface (CLI) for both prediction and training tasks.
To get help and see the available commands:
instanovo --help
To see the version of InstaNovo, InstaNovo+ and some of the dependencies:
instanovo version
Predicting
To get help about the prediction command line options:
instanovo predict --help
Running predictions with both InstaNovo and InstaNovo+
The default is to run predictions first with the transformer-based InstaNovo model, and then further improve the performance by iterative refinement of these predicted sequences by the diffusion-based InstaNov+ model.
instanovo predict --data-path ./sample_data/spectra.mgf --output-path predictions.csv
Which results in the following output:
scan_number,precursor_mz,precursor_charge,experiment_name,spectrum_id,diffusion_predictions_tokenised,diffusion_predictions,diffusion_log_probabilities,transformer_predictions,transformer_predictions_tokenised,transformer_log_probabilities,transformer_token_log_probabilities
0,451.25348,2,spectra,spectra:0,"['A', 'L', 'P', 'Y', 'T', 'P', 'K', 'K']",ALPYTPKK,-0.03160184621810913,LAHYNKK,"L, A, H, Y, N, K, K",-424.5889587402344,"[-0.5959059000015259, -0.0059959776699543, -0.01749008148908615, -0.03598890081048012, -0.48958998918533325, -1.5242897272109985, -0.656516432762146]"
To evaluate InstaNovo performance on an annotated dataset:
bash
instanovo predict --evaluation --data-path ./sample_data/spectra.mgf --output-path predictions.csv
Which results in the following output:
scan_number,precursor_mz,precursor_charge,experiment_name,spectrum_id,diffusion_predictions_tokenised,diffusion_predictions,diffusion_log_probabilities,targets,transformer_predictions,transformer_predictions_tokenised,transformer_log_probabilities,transformer_token_log_probabilities
0,451.25348,2,spectra,spectra:0,"['L', 'A', 'H', 'Y', 'N', 'K', 'K']",LAHYNKK,-0.06637095659971237,IAHYNKR,LAHYNKK,"L, A, H, Y, N, K, K",-424.5889587402344,"[-0.5959059000015259, -0.0059959776699543, -0.01749008148908615, -0.03598890081048012, -0.48958998918533325, -1.5242897272109985, -0.656516432762146]"
Note that the --evaluation flag includes the targets column in the output, which contains the
ground truth peptide sequence. Metrics will be calculated and displayed in the console.
Command line arguments and overriding config values
The configuration file for inference may be found under
instanovo/configs/inference/ folder. By default, the
default.yaml file is used.
InstaNovo uses command line arguments for commonly used parameters:
--data-path- Path to the dataset to be evaluated. Allows.mgf,.mzml,.mzxml,.ipcor a directory. Glob notation is supported: eg.:./experiment/*.mgf--output-path- Path to output csv file.--instanovo-model- Model to use for InstaNovo. Either a model ID (currently supported:instanovo-v1.1.0) or a path to an Instanovo checkpoint file (.ckpt format).--instanovo-plus-model- Model to use for InstaNovo+. Either a model ID (currently supported:instanovoplus-v1.1.0) or a path to an Instanovo+ checkpoint file (.ckpt format).--denovo- Whether to do de novo predictions. If you want to evaluate the model on annotated data, use the flag--evaluationflag.--with-refinement- Whether to use InstaNovo+ for iterative refinement of InstaNovo predictions. Default isTrue. If you don't want to use refinement,use the flag--no-refinement.
To override the configuration values in the config files, you can use command line arguments. For example, by default beam search with one beam is used. If you want to use beam search with 5 beams, you can use the following command:
bash
instanovo predict --data-path ./sample_data/spectra.mgf --output-path predictions.csv num_beams=5
Note the lack of prefix -- before num_beams in the command line argument because you are
overriding the value of key defined in the config file.
Output description
When output_path is specified, a CSV file will be generated containing predictions for all the
input spectra. The model will attempt to generate a peptide for every MS2 spectrum regardless of
confidence. We recommend filtering the output using the log_probabilities and deltamassppm
columns.
| Column | Description | Data Type | Notes |
| ----------------------- | -------------------------------------------------------------- | ------------ | ------------------------------------------------------------------------------------------------------------- |
| scannumber | Scan number of the MS/MS spectrum | Integer | Unique identifier from the input file |
| precursormz | Precursor m/z (mass-to-charge ratio) | Float | The observed m/z of the precursor ion |
| precursorcharge | Precursor charge state | Integer | Charge state of the precursor ion |
| experimentname | Experiment name derived from input filename | String | Based on the input file name (mgf, mzml, or mzxml) |
| spectrumid | Unique spectrum identifier | String | Combination of experiment name and scan number (e.g., yeast:17738) |
| targets | Target peptide sequence | String | Ground truth peptide sequence (if available) |
| predictions | Predicted peptide sequences | String | Model-predicted peptide sequence |
| predictionstokenised | Predicted peptide sequence tokenized by amino acids | List[String] | Each amino acid token separated by commas |
| logprobabilities | Log probability of the entire predicted sequence | Float | Natural logarithm of the sequence confidence, can be converted to probability with np.exp(logprobabilities). |
| tokenlogprobabilities | Log probability of each token in the predicted sequence | List[Float] | Natural logarithm of the sequence confidence per amino acid |
| deltamassppm | Mass difference between precursor and predicted peptide in ppm | Float | Mass deviation in parts per million |
Models
InstaNovo 1.1.0 includes new models instanovo-v1.1.0.ckpt, and instanovoplus-v1.1.0.ckpt trained
on a larger dataset with more PTMs.
Note: The InstaNovo Extended 1.0.0 training data mis-represented Cysteine as unmodified for the majority of the training data. Please update to the latest version of the model.
Training Datasets
- ProteomeTools Part
I (PXD004732),
II (PXD010595), and
III (PXD021013) \
(referred to as the all-confidence ProteomeTools
AC-PTdataset in our paper) - Additional PRIDE dataset with more modifications: \ (PXD000666, PXD000867, PXD001839, PXD003155, PXD004364, PXD004612, PXD005230, PXD006692, PXD011360, PXD011536, PXD013543, PXD015928, PXD016793, PXD017671, PXD019431, PXD019852, PXD026910, PXD027772)
- Massive-KB v1
- Additional phosphorylation dataset \ (not yet publicly released)
Natively Supported Modifications
| Amino Acid | Single Letter | Modification | Mass Delta (Da) | Unimod ID | | --------------------------- | ------------- | ----------------------- | --------------- | --------------------------------------------------------------------------- | | Methionine | M | Oxidation | +15.9949 | [UNIMOD:35] | | Cysteine | C | Carboxyamidomethylation | +57.0215 | [UNIMOD:4] | | Asparagine, Glutamine | N, Q | Deamidation | +0.9840 | [UNIMOD:7] | | Serine, Threonine, Tyrosine | S, T, Y | Phosphorylation | +79.9663 | [UNIMOD:21] | | N-terminal | - | Ammonia Loss | -17.0265 | [UNIMOD:385] | | N-terminal | - | Carbamylation | +43.0058 | [UNIMOD:5] | | N-terminal | - | Acetylation | +42.0106 | [UNIMOD:1] |
See residue configuration under instanovo/configs/residues/extended.yaml
Training
Data to train on may be provided in any format supported by the SpectrumDataHandler. See section on data conversion for preferred formatting.
Training InstaNovo
To train the auto-regressive transformer model InstaNovo using the config file instanovo/configs/instanovo.yaml, you can use the following command:
bash
instanovo transformer train --help
To update the InstaNovo model config, modify the config file under instanovo/configs/model/instanovo_base.yaml
Training InstaNovo+
To train the diffusion model InstaNovo+ using the config file instanovo/configs/instanovoplus.yaml, you can use the following command:
bash
instanovo diffusion train --help
To update the InstaNovo+ model config, modify the config file under instanovo/configs/model/instanovoplus_base.yaml
Advanced prediction options
Run predictions with only InstaNovo
If you want to run predictions with only InstaNovo, you can use the following command:
bash
instanovo transformer predict --help
Run predictions with only InstaNovo+
If you want to run predictions with only InstaNovo+, you can use the following command:
bash
instanovo diffusion predict --help
Run predictions with InstaNovo and InstaNovo+ in separate steps
You can first run predictions with InstaNovo
bash
instanovo transformer predict --data-path ./sample_data/spectra.mgf --output-path instanovo_predictions.csv
and then use the predictions as input for InstaNovo+:
bash
instanovo diffusion predict --data-path ./sample_data/spectra.mgf --output-path instanovo_plus_predictions.csv instanovo_predictions_path=instanovo_predictions.csv
Performance
We have benchmarked our latest models InstaNovo v1.1 and InstaNovo+ v1.1 against our previous models. For all results below, InstaNovo decoding was performed with knapsack beam search decoding. InstaNovo+ then refined these results. We present peptide accuracy as the metric of comparison. Peptide accuracy is a measure of precision at full coverage (no filtering).
Nine-species dataset
| Dataset | InstaNovo v0.1 | InstaNovo+ v0.1 | InstaNovo v1.1 | InstaNovo+ v1.1 | | -------- | -------------- | --------------- | -------------- | --------------- | | Bacillus | 0.624 | 0.674 | 0.652 | 0.684 | | Mouse | 0.466 | 0.490 | 0.524 | 0.542 | | Yeast | 0.559 | 0.624 | 0.618 | 0.645 |
InstaNovo and InstaNovo+ v0.1 were fine-tuned on the eight species dataset, excluding the test species, whereas InstaNovo and InstaNovo+ v1.1 were evaluated zero-shot on these datasets.
Biological validation datasets
| Dataset | InstaNovo v0.1 | InstaNovo+ v0.1 | InstaNovo v1.1 | InstaNovo+ v1.1 | | ------------------------------- | -------------- | --------------- | -------------- | --------------- | | HeLa degradome | 0.695 | 0.719 | 0.813 | 0.821 | | HeLa single-shot | 0.503 | 0.517 | 0.642 | 0.647 | | Herceptin | 0.494 | 0.562 | 0.710 | 0.720 | | Immunopeptidomics | 0.581 | 0.697 | 0.707 | 0.748 | | Candidatus "Scalindua brodae" | 0.724 | 0.736 | 0.748 | 0.762 | | Snake venoms | 0.196 | 0.198 | 0.221 | 0.238 | | Nanobodies | 0.447 | 0.464 | 0.492 | 0.508 | | Wound fluids | 0.225 | 0.229 | 0.354 | 0.364 |
Additional features
Spectrum Data Class
InstaNovo introduces a Spectrum Data Class: SpectrumDataFrame.
This class acts as an interface between many common formats used for storing mass spectrometry,
including .mgf, .mzml, .mzxml, and .csv. This class also supports reading directly from
HuggingFace, Pandas, and Polars.
When using InstaNovo, these formats are natively supported and automatically converted to the
internal SpectrumDataFrame supported by InstaNovo for training and inference. Any data path may be
specified using glob notation. For example you
could use the following command to get de novo predictions from all the files in the folder
./experiment:
bash
instanovo predict --data_path=./experiment/*.mgf
Alternatively, a list of files may be specified in the inference config.
The SpectrumDataFrame also allows for loading of much larger datasets in a lazy way. To do this, the
data is loaded and stored as .parquet files in a
temporary directory. Alternatively, the data may be saved permanently natively as .parquet for
optimal loading.
Example usage:
Converting mgf files to the native format:
```python from instanovo.utils import SpectrumDataFrame
Convert mgf files native parquet:
sdf = SpectrumDataFrame.load("/path/to/data.mgf", lazy=False, isannotated=True) sdf.save("path/to/parquet/folder", partition="train", chunksize=1e6) ```
Loading the native format in shuffle mode:
```python
Load a native parquet dataset:
sdf = SpectrumDataFrame.load("path/to/parquet/folder", partition="train", shuffle=True, lazy=True, is_annotated=True) ```
Using the loaded SpectrumDataFrame in a PyTorch DataLoader:
```python from instanovo.transformer.dataset import SpectrumDataset from torch.utils.data import DataLoader
ds = SpectrumDataset(sdf)
Note: Shuffle and workers is handled by the SpectrumDataFrame
dl = DataLoader( ds, collatefn=SpectrumDataset.collatebatch, shuffle=False, num_workers=0, ) ```
Some more examples using the SpectrumDataFrame:
```python sdf = SpectrumDataFrame.load("/path/to/experiment/*.mzml", lazy=True)
Remove rows with a charge value > 3:
sdf.filterrows(lambda row: row["precursorcharge"]<=2)
Sample a subset of the data:
sdf.sample_subset(fraction=0.5, seed=42)
Convert to pandas
df = sdf.to_pandas() # Returns a pd.DataFrame
Convert to polars LazyFrame
lazydf = sdf.topolars(return_lazy=True) # Returns a pl.LazyFrame
Save as an .mgf file
sdf.write_mgf("path/to/output.mgf") ```
SpectrumDataFrame Features:
- The SpectrumDataFrame supports lazy loading with asynchronous prefetching, mitigating wait times between files.
- Filtering and sampling may be performed non-destructively through on file loading
- A two-fold shuffling strategy is introduced to optimise sampling during training (shuffling files and shuffling within files).
Using your own datasets
To use your own datasets, you simply need to tabulate your data in either Pandas or Polars with the following schema:
The dataset is tabular, where each row corresponds to a labelled MS2 spectra.
sequence (string)\ The target peptide sequence including post-translational modificationsmodified_sequence (string) [legacy]\ The target peptide sequence including post-translational modificationsprecursor_mz (float64)\ The mass-to-charge of the precursor (from MS1)charge (int64)\ The charge of the precursor (from MS1)mz_array (list[float64])\ The mass-to-charge values of the MS2 spectrumintensity_array (list[float32])\ The intensity values of the MS2 spectrum
For example, the DataFrame for the nine species benchmark dataset (introduced in Tran et al. 2017) looks as follows:
| | sequence | precursormz | precursorcharge | mzarray | intensityarray | | --: | :----------------------------- | -----------: | ---------------: | :----------------------------------- | :---------------------------------- | | 0 | GRVEGMEAR | 335.502 | 3 | [102.05527 104.052956 113.07079 ...] | [ 767.38837 2324.8787 598.8512 ...] | | 1 | IGEYK | 305.165 | 2 | [107.07023 110.071236 111.11693 ...] | [ 1055.4957 2251.3171 35508.96 ...] | | 2 | GVSREEIQR | 358.528 | 3 | [103.039444 109.59844 112.08704 ...] | [801.19995 460.65268 808.3431 ...] | | 3 | SSYHADEQVNEASK | 522.234 | 3 | [101.07095 102.0552 110.07163 ...] | [ 989.45154 2332.653 1170.6191 ...] | | 4 | DTFNTSSTSN[UNIMOD:7]STSSSSSNSK | 676.282 | 3 | [119.82458 120.08073 120.2038 ...] | [ 487.86942 4806.1377 516.8846 ...] |
For de novo prediction, the sequence column is not required.
We also provide a conversion script for converting to native SpectrumDataFrame (sdf) format:
bash
instanovo convert --help
Development
uv setup
This project is set up to use uv to manage Python and dependencies. First, be sure you have uv installed on your system.
On Linux and macOS:
bash
curl -LsSf https://astral.sh/uv/install.sh | sh
On Windows:
powershell
powershell -c "irm https://astral.sh/uv/install.ps1 | iex"
Note: InstaNovo is built for Python >=3.10, <3.13 and tested on Linux.
Fork and clone the repository
Then fork this repo (having your own fork will make it easier to contribute) and clone it.
bash
git clone https://github.com/YOUR-USERNAME/InstaNovo.git
cd InstaNovo
And install the dependencies. If you do have access to an NVIDIA GPU, you can install the GPU version of PyTorch (recommended):
bash
uv sync --extra cu124
uv run pre-commit install
If you don't have access to a GPU, you can install the CPU-only version of PyTorch:
bash
uv sync --extra cpu
uv run pre-commit install
Both approaches above also install the development dependencies. If you also want to install the documentation dependencies, you can do so with:
bash
uv sync --extra cu124 --group docs
Activate the virtual environment:
bash
source .venv/bin/activate
To upgrade all packages to the latest versions, you can run:
bash
uv lock --upgrade
uv sync --extra cu124
Basic development workflows
Testing
InstaNovo uses pytest for testing. To run the tests, you can use the following command:
bash
uv run instanovo/scripts/get_zenodo_record.py # Download the test data
python -m pytest --cov-report=html --cov --random-order --verbose .
To see the coverage report, run:
bash
python -m coverage report -m
To view the coverage report in a browser, run:
bash
python -m http.server --directory ./coverage
and navigate to http://0.0.0.0:8000/ in your browser.
Linting
InstaNovo uses pre-commit hooks to ensure code quality. To run the linters, you can use the following command:
bash
pre-commit run --all-files
Building the documentation
To build the documentation locally, you can use the following commands:
bash
uv sync --extra cu124 --group docs
git config --global --add safe.directory "$(dirname "$(pwd)")"
rm -rf docs/reference
python ./docs/gen_ref_nav.py
mkdocs build --verbose --site-dir docs_public
mkdocs serve
Generating a requirements.txt file
If you have a pip or conda based workflow and want to generate a requirements.txt file, you
can use the following command:
bash
uv export --format requirements-txt > requirements.txt
Setting Python interpreter in VSCode
To set the Python interpreter in VSCode, open the Command Palette (Ctrl+Shift+P), search for
Python: Select Interpreter, and select ./.venv/bin/python.
License
Code is licensed under the Apache License, Version 2.0 (see LICENSE)
The model checkpoints are licensed under Creative Commons Non-Commercial (CC BY-NC-SA 4.0)
BibTeX entry and citation info
If you use InstaNovo in your research, please cite the following paper:
bibtex
@article{eloff_kalogeropoulos_2025_instanovo,
title = {InstaNovo enables diffusion-powered de novo peptide sequencing in large-scale
proteomics experiments},
author = {Eloff, Kevin and Kalogeropoulos, Konstantinos and Mabona, Amandla and Morell,
Oliver and Catzel, Rachel and Rivera-de-Torre, Esperanza and Berg Jespersen,
Jakob and Williams, Wesley and van Beljouw, Sam P. B. and Skwark, Marcin J.
and Laustsen, Andreas Hougaard and Brouns, Stan J. J. and Ljungars,
Anne and Schoof, Erwin M. and Van Goey, Jeroen and auf dem Keller, Ulrich and
Beguir, Karim and Lopez Carranza, Nicolas and Jenkins, Timothy P.},
year = 2025,
month = {Mar},
day = 31,
journal = {Nature Machine Intelligence},
doi = {10.1038/s42256-025-01019-5},
issn = {2522-5839},
url = {https://doi.org/10.1038/s42256-025-01019-5}
}
Acknowledgements
Big thanks to Pathmanaban Ramasamy, Tine Claeys, and Lennart Martens of the CompOmics research group for providing us with additional phosphorylation training data.
Owner
- Name: InstaDeep Ltd
- Login: instadeepai
- Kind: organization
- Email: hello@instadeep.com
- Location: London, UK
- Website: https://instadeep.com
- Twitter: instadeepai
- Repositories: 14
- Profile: https://github.com/instadeepai
We productise innovation
Citation (CITATION.cff)
cff-version: 1.2.0
message: "If you use this software, please cite it as below."
authors:
- family-names: "Eloff"
given-names: "Kevin"
orcid: "https://orcid.org/0000-0003-1355-8743"
- family-names: "Mabona"
given-names: "Amandla"
orcid: "https://orcid.org/0009-0009-7514-677X"
- family-names: "Catzel"
given-names: "Rachel"
orcid: "https://orcid.org/0000-0002-2983-9833"
- family-names: "Van Goey"
given-names: "Jeroen"
orcid: "https://orcid.org/0000-0003-4480-5567"
title: "InstaNovo"
version: 1.1.1
doi: 10.5281/zenodo.15101326
date-released: 2025-03-28
url: "https://github.com/instadeepai/instanovo"
preferred-citation:
type: article
authors:
- family-names: "Eloff"
given-names: "Kevin"
orcid: "https://orcid.org/0000-0003-1355-8743"
- family-names: "Kalogeropoulos"
given-names: "Konstantinos"
orcid: "https://orcid.org/0000-0003-3907-9281"
- family-names: "Mabona"
given-names: "Amandla"
orcid: "https://orcid.org/0009-0009-7514-677X"
- family-names: "Morell"
given-names: "Oliver"
orcid: "https://orcid.org/0009-0000-8702-1792"
- family-names: "Catzel"
given-names: "Rachel"
orcid: "https://orcid.org/0000-0002-2983-9833"
- family-names: "Rivera-de-Torre"
given-names: "Esperanza"
orcid: "https://orcid.org/0000-0002-0272-6150"
- family-names: "Berg Jespersen"
given-names: "Jakob"
orcid: "https://orcid.org/0000-0001-6634-6256"
- family-names: "Williams"
given-names: "Wesley"
orcid: "https://orcid.org/0000-0002-8712-7498"
- family-names: "van Beljouw"
given-names: "Sam"
orcid: "https://orcid.org/0000-0003-3892-5028"
- family-names: "Skwark"
given-names: "Marcin"
orcid: "https://orcid.org/0000-0002-2022-6766"
- family-names: "Hougaard Laustsen"
given-names: "Andreas"
orcid: "https://orcid.org/0000-0001-6918-5574"
- family-names: "Brouns"
given-names: "Stan J. J."
orcid: "https://orcid.org/0000-0002-9573-1724"
- family-names: "Ljungars"
given-names: "Anne"
orcid: "https://orcid.org/0000-0002-2158-0601"
- family-names: "Schoof"
given-names: "Erwin M."
orcid: "https://orcid.org/0000-0002-3117-7832"
- family-names: "Van Goey"
given-names: "Jeroen"
orcid: "https://orcid.org/0000-0003-4480-5567"
- family-names: "auf dem Keller"
given-names: "Ulrich"
orcid: "https://orcid.org/0000-0002-3431-7415"
- family-names: "Beguir"
given-names: "Karim"
orcid: "https://orcid.org/0000-0002-2789-6527"
- family-names: "Lopez Carranza"
given-names: "Nicolas"
orcid: "https://orcid.org/0000-0003-2235-5753"
- family-names: "Jenkins"
given-names: "Timothy P."
orcid: "https://orcid.org/0000-0003-2979-5663"
doi: 10.1038/s42256-025-01019-5
journal: "Nature Machine Intelligence"
title:
"InstaNovo enables diffusion-powered de novo peptide sequencing in large-scale proteomics
experiments"
url: "https://www.nature.com/articles/s42256-025-01019-5"
GitHub Events
Total
- Create event: 32
- Release event: 5
- Issues event: 23
- Watch event: 44
- Delete event: 8
- Member event: 2
- Issue comment event: 47
- Push event: 63
- Pull request review comment event: 3
- Pull request review event: 19
- Pull request event: 57
- Fork event: 15
Last Year
- Create event: 32
- Release event: 5
- Issues event: 23
- Watch event: 44
- Delete event: 8
- Member event: 2
- Issue comment event: 47
- Push event: 63
- Pull request review comment event: 3
- Pull request review event: 19
- Pull request event: 57
- Fork event: 15
Committers
Last synced: 10 months ago
Top Committers
| Name | Commits | |
|---|---|---|
| Jeroen Van Goey | j****y@i****m | 43 |
| Kevin | k****f@i****m | 36 |
| Amandla Mabona | a****a@i****m | 18 |
| Rachel Catzel | r****l@i****m | 10 |
| dependabot[bot] | 4****] | 3 |
| Jesper Lauridsen | 7****u | 1 |
Committer Domains (Top 20 + Academic)
Issues and Pull Requests
Last synced: 6 months ago
All Time
- Total issues: 30
- Total pull requests: 106
- Average time to close issues: 4 months
- Average time to close pull requests: 9 days
- Total issue authors: 20
- Total pull request authors: 6
- Average comments per issue: 1.8
- Average comments per pull request: 0.52
- Merged pull requests: 72
- Bot issues: 0
- Bot pull requests: 26
Past Year
- Issues: 15
- Pull requests: 72
- Average time to close issues: 22 days
- Average time to close pull requests: 9 days
- Issue authors: 13
- Pull request authors: 5
- Average comments per issue: 1.0
- Average comments per pull request: 0.51
- Merged pull requests: 45
- Bot issues: 0
- Bot pull requests: 18
Top Authors
Issue Authors
- irleader (4)
- JannikSchneider12 (2)
- denisbeslic (2)
- lostculture (2)
- 07liesin (2)
- KevinEloff (2)
- cguetot (2)
- biocc (1)
- dumbgoos (1)
- Happyers (1)
- bsphinney (1)
- ayavinash (1)
- jack-jiang01 (1)
- cliang-huanglab (1)
- AJ-2022-al (1)
Pull Request Authors
- BioGeek (57)
- dependabot[bot] (42)
- KevinEloff (13)
- rcatzel (12)
- amabinsta (3)
- jesperdlau (2)
Top Labels
Issue Labels
Pull Request Labels
Packages
- Total packages: 1
-
Total downloads:
- pypi 166 last-month
- Total dependent packages: 0
- Total dependent repositories: 0
- Total versions: 12
- Total maintainers: 1
pypi.org: instanovo
InstaNovo enables diffusion-powered de novo peptide sequencing in large scale proteomics experiments
- Homepage: https://github.com/instadeepai/InstaNovo
- Documentation: https://instanovo.readthedocs.io/
- License: apache-2.0
-
Latest release: 1.1.4
published 9 months ago
Rankings
Maintainers (1)
Dependencies
- actions/checkout v3 composite
- actions/setup-python v4 composite
- pypa/gh-action-pypi-publish 27b31702a0e7fc50959f5ad993c78deac1bdfc29 composite
- aiobotocore ==2.4.1
- boto3 ==1.24.59
- botocore ==1.27.59
- click ==8.1.7
- cloudpathlib ==0.10.0
- datasets ==2.14.5
- deepspeed ==0.7.2
- depthcharge-ms ==0.1.0
- fastprogress ==1.0.3
- jiwer ==2.5.1
- matplotlib ==3.6.0
- numpy ==1.23.3
- omegaconf ==2.2.3
- pandas ==1.5.0
- polars ==0.17.8
- protobuf ==3.19.6
- pyarrow ==11.0.0
- python-dotenv ==0.21.0
- pytorch_lightning ==1.8.6
- scikit-learn ==1.1.2
- seaborn ==0.12.0
- spectrum_utils ==0.4.1
- tensorboard ==2.10.1
- tensorboardX ==2.5.1
- torch ==1.13.1
- torchaudio ==0.13.1
- torchvision ==0.14.1
- tqdm ==4.64.1