rabies
fMRI preprocessing pipeline and analysis tools adapted for rodent images. Visit the full documentation at https://rabies.readthedocs.io/
Science Score: 67.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
✓CITATION.cff file
Found CITATION.cff file -
✓codemeta.json file
Found codemeta.json file -
✓.zenodo.json file
Found .zenodo.json file -
✓DOI references
Found 2 DOI reference(s) in README -
○Academic publication links
-
✓Committers with academic emails
2 of 7 committers (28.6%) from academic institutions -
○Institutional organization owner
-
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (16.0%) to scientific vocabulary
Repository
fMRI preprocessing pipeline and analysis tools adapted for rodent images. Visit the full documentation at https://rabies.readthedocs.io/
Basic Info
Statistics
- Stars: 44
- Watchers: 3
- Forks: 20
- Open Issues: 67
- Releases: 8
Metadata Files
README.md
RABIES: Rodent Automated Bold Improvement of EPI Sequences.
RABIES is an open source image processing pipeline for rodent fMRI. It conducts state-of-the-art preprocessing and confound correction, and supplies standard resting-state functional connectivity analyses. Visit our documentation at https://rabies.readthedocs.io/en/latest/.
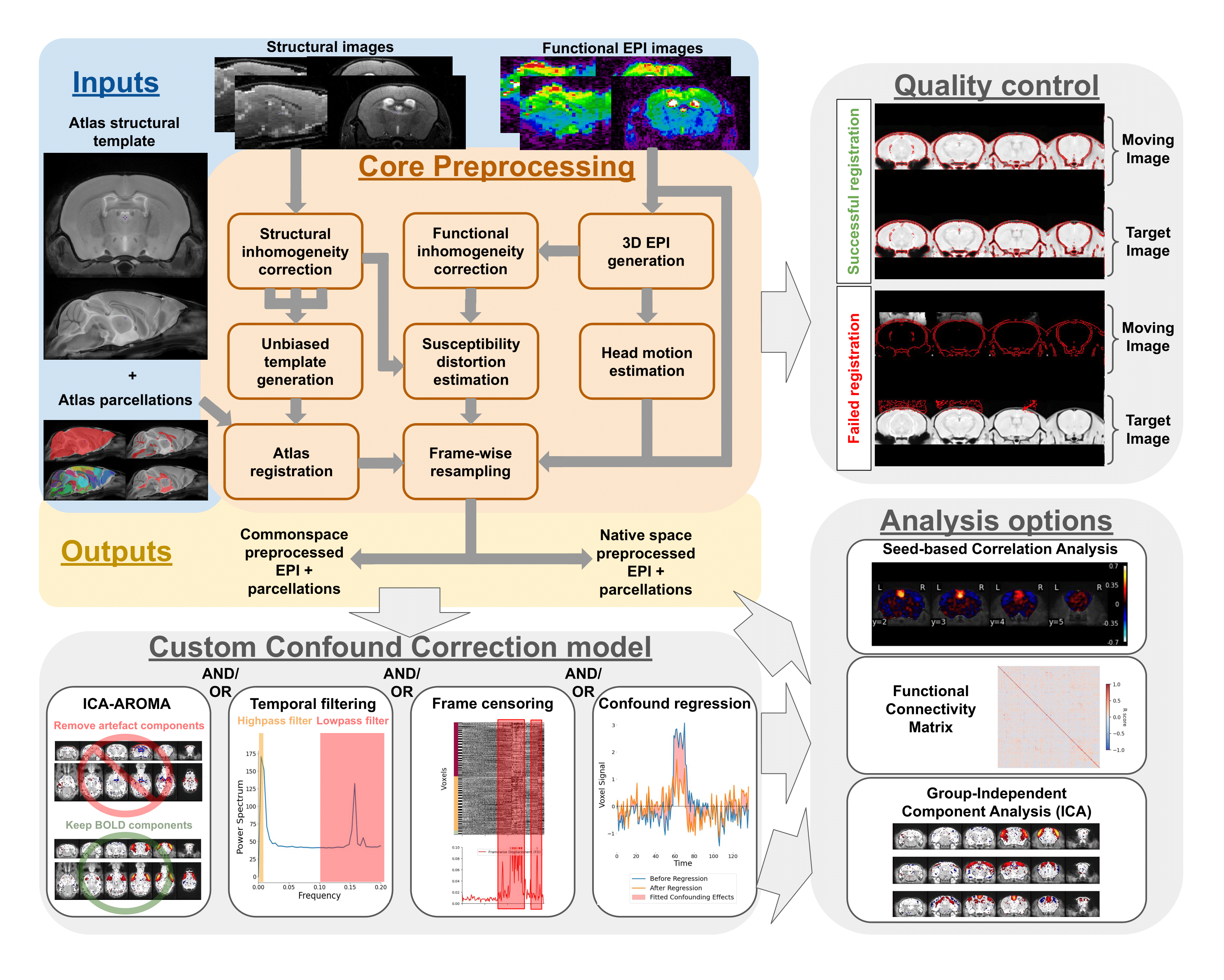
What you can do with RABIES
The primary purpose of RABIES is to provide rodent fMRI research with a standard, flexible, and reliable image processing platform. The package is complemented with informative data diagnostic features for confound characterization and encourages best practices for quality control and reproducibility. The RABIES software is structured into three main processing stages: preprocessing, confound correction and analysis.
Preprocessing
The preprocessing workflow regroups essential fMRI preprocessing steps prior to analysis. It includes a robust registration workflow with automatically-adapting parameters allowing to succesfully process diverse acquisition types (i.e. rodent species, scan field strength, coil type, ...), and can conduct the following preprocessing steps: - head motion correction - susceptibility distortion correction - resampling to native or common space - brain parcellation - slice timing correction (optional) - despiking (optional) - visual assessment of registration for quality control
Confound correction
Following preprocessing, a range of strategies to correct fMRI confounds (e.g. motion) can then be conducted within RABIES: - linear detrending - confound regression (with several options for nuisance regressors) - frequency filtering (highpass, lowpass, bandpass) - frame censoring (or scrubbing) - ICA-AROMA - spatial smoothing
Analysis
Simple resting-state connectivity analyses are made available after preprocessing and confound correction. RABIES also provides a 'data diagnosis' workflow, which generates several indices of data quality and potential confounds, and conversaly, aims to improve the correction of confounds and transparency with regards to data quality: - seed-based functional connectivity - whole-brain connectivity matrix - group-ICA - dual regression - data diagnosis
Notes on software design
Nipype workflows: The image processing pipelines are structured using the Nipype library, which allows to build dynamic workflows in the form of a computational graph. Each node in the graph consists of a processing step, and the required inputs/outputs define the links between nodes. In addition to supporting code organization, Nipype workflows also handle several plugin architectures for parallel execution as well as memory management. The computational time to run the entire RABIES pipeline will vary substantially depending on data size, but for most uses, it will range from a few hours to a day when using proper computational resources and parallel execution.
Reproducible and transparent research: RABIES aims to follow best practices for reproducible and transparent research, including the following: - open source code https://github.com/CoBrALab/RABIES - standardized input data format with BIDS - easily shared, automatically-generated visual outputs for quality control - containerized distribution of the software hosted on Docker Hub which can be downloaded via Docker and Singularity platforms
Citation
Citing RABIES: Please cite the official publication Desrosiers-Grégoire, et al. Nat Commun 15, 6708 (2024). when referencing the software.
Boilerplate: a boilerplate summarizing the preprocessing and confound correction operations is automatically generated in the output folder. You can use the boilerplate to help describe your methods in a paper.
License
The RABIES license allows for uses in academic and educational environments only. Commercial use requires a commercial license from CoBrALab contact@cobralab.ca, http://cobralab.ca
Acknowledgements
This software was developped by the CoBrALab, located at the Cerebral Imaging Center of the Douglas Mental Health University Institute, Montreal, Canada, in affiliation with McGill University, Montreal, Canada. This work was supported by funding from Healthy Brains, Healthy Lives (HBHL), the Fonds de recherche du Québec - Santé (FRQS) and - Nature et technologies (FRQNT), and the Natural Sciences and Engineering Research Council (NSERC) of Canada. fMRIPrep was an important inspirational source for this project, in particular with regards to best practices for software reproducibility and code design using Nipype. We also thank the organizers of BrainHack School Montreal, which guided the initial steps of this project in 2018.
Ask for help
If you need support in using the software or experience issues that are not documented, we'll provide support on the Github discussion.
Contributing to RABIES
Read our dedicated documentation
Owner
- Name: Computational Brain Anatomy Laboratory
- Login: CoBrALab
- Kind: organization
- Email: contact@cobralab.ca
- Location: Montreal, QC
- Website: http://cobralab.ca
- Repositories: 36
- Profile: https://github.com/CoBrALab
Computational Brain Anatomy Laboratory, located in the CIC at the Douglas Institute, McGill University
Citation (CITATION.cff)
cff-version: 1.2.0
message: "If you use this software, please cite it as below."
authors:
- family-names: "Desrosiers-Gregoire"
given-names: "Gabriel"
- family-names: "Devenyi"
given-names: "Gabriel A."
- family-names: "Grandjean"
given-names: "Joanes"
- family-names: "Chakravarty"
given-names: "M. Mallar"
title: "RABIES: Rodent Automated Bold Improvement of EPI Sequences."
version: 0.5.2
date-released: 2025-06-30
url: "https://github.com/CoBrALab/RABIES"
preferred-citation:
type: article
authors:
- family-names: "Desrosiers-Gregoire"
given-names: "Gabriel"
- family-names: "Devenyi"
given-names: "Gabriel A."
- family-names: "Grandjean"
given-names: "Joanes"
- family-names: "Chakravarty"
given-names: "M. Mallar"
doi: "10.1038/s41467-024-50826-8"
journal: "Nat. Commun."
month: 8
start: 1 # First page number
end: 15 # Last page number
title: "A standardized image processing and data quality platform for rodent fMRI"
volume: 15
year: 2024
GitHub Events
Total
- Create event: 10
- Release event: 2
- Issues event: 32
- Watch event: 8
- Delete event: 5
- Member event: 1
- Issue comment event: 84
- Push event: 36
- Pull request review comment event: 28
- Pull request review event: 23
- Pull request event: 18
- Fork event: 5
Last Year
- Create event: 10
- Release event: 2
- Issues event: 32
- Watch event: 8
- Delete event: 5
- Member event: 1
- Issue comment event: 84
- Push event: 36
- Pull request review comment event: 28
- Pull request review event: 23
- Pull request event: 18
- Fork event: 5
Committers
Last synced: almost 3 years ago
All Time
- Total Commits: 652
- Total Committers: 7
- Avg Commits per committer: 93.143
- Development Distribution Score (DDS): 0.087
Top Committers
| Name | Commits | |
|---|---|---|
| Gab-D-G | g****e@m****a | 595 |
| Gab-D-G | 3****G@u****m | 33 |
| Gabriel A. Devenyi | g****i@g****m | 17 |
| davidgruskin | 3****n@u****m | 3 |
| Jed Lamari-Saysset | j****t@m****a | 2 |
| grandjeanlab | jo@s****h | 1 |
| Cedric Barre | c****r@g****m | 1 |
Committer Domains (Top 20 + Academic)
Issues and Pull Requests
Last synced: 6 months ago
All Time
- Total issues: 113
- Total pull requests: 98
- Average time to close issues: 5 months
- Average time to close pull requests: 3 days
- Total issue authors: 31
- Total pull request authors: 9
- Average comments per issue: 2.41
- Average comments per pull request: 0.65
- Merged pull requests: 80
- Bot issues: 0
- Bot pull requests: 7
Past Year
- Issues: 23
- Pull requests: 20
- Average time to close issues: 5 days
- Average time to close pull requests: 1 day
- Issue authors: 13
- Pull request authors: 6
- Average comments per issue: 1.78
- Average comments per pull request: 2.1
- Merged pull requests: 10
- Bot issues: 0
- Bot pull requests: 1
Top Authors
Issue Authors
- Gab-D-G (33)
- gdevenyi (18)
- grandjeanlab (10)
- KarisColyerPatel (5)
- geowk (4)
- sxf-ux (4)
- mapepe18 (4)
- dortuzarm (3)
- schmidt12335 (3)
- david-roper (2)
- nmskhan (2)
- benhibou (2)
- Mushtaq82 (2)
- lvqianmri (1)
- mhanafyuwo (1)
Pull Request Authors
- Gab-D-G (75)
- dependabot[bot] (8)
- CMonnin (7)
- milaurose (3)
- davidgruskin (3)
- gdevenyi (3)
- david-roper (2)
- coderabbitai[bot] (1)
- cedricBarre (1)
Top Labels
Issue Labels
Pull Request Labels
Packages
- Total packages: 1
-
Total downloads:
- pypi 178 last-month
- Total docker downloads: 28
- Total dependent packages: 0
- Total dependent repositories: 1
- Total versions: 14
- Total maintainers: 3
pypi.org: rabies
RABIES: Rodent Automated Bold Improvement of EPI Sequences.
- Homepage: https://github.com/CoBrALab/RABIES
- Documentation: https://rabies.readthedocs.io/
- License: LICENSE
-
Latest release: 0.5.2
published 8 months ago
Rankings
Dependencies
- base latest build
- ubuntu 18.04 build
- mcin/docker-fsl latest build
- jinja2 ==3.1.1
- myst-parser ==0.17.2
- sphinx ==4.5.0
- sphinx-rtd-dark-mode *
- sphinx-rtd-theme *
- sphinxcontrib-bibtex *
- future *
- matplotlib ==2.2
- nibabel *
- numpy ==1.14
- pandas ==0.23
- seaborn ==0.9.0
- actions/checkout v3 composite
- docker/build-push-action 0565240e2d4ab88bba5387d719585280857ece09 composite
- docker/login-action 343f7c4344506bcbf9b4de18042ae17996df046d composite
- docker/metadata-action 96383f45573cb7f253c731d3b3ab81c87ef81934 composite
- docker/setup-buildx-action f95db51fddba0c2d1ec667646a06c2ce06100226 composite