nelpy
Neuroelectrophysiology object model and data analysis in Python.
Science Score: 46.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
○CITATION.cff file
-
✓codemeta.json file
Found codemeta.json file -
✓.zenodo.json file
Found .zenodo.json file -
○DOI references
-
✓Academic publication links
Links to: zenodo.org -
✓Committers with academic emails
3 of 11 committers (27.3%) from academic institutions -
○Institutional organization owner
-
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (12.7%) to scientific vocabulary
Keywords
Repository
Neuroelectrophysiology object model and data analysis in Python.
Basic Info
- Host: GitHub
- Owner: nelpy
- License: mit
- Language: Jupyter Notebook
- Default Branch: master
- Homepage: https://nelpy.github.io/nelpy/
- Size: 62.5 MB
Statistics
- Stars: 50
- Watchers: 10
- Forks: 24
- Open Issues: 100
- Releases: 5
Topics
Metadata Files
README.md

Neuroelectrophysiology object model, data exploration, and analysis in Python.
Overview
Nelpy (Neuroelectrophysiology) is an open source package for analysis of neuroelectrophysiology data. Nelpy defines a number of data objects to make it easier to work with electrophysiology (ephys) data, and although it was originally designed for use with extracellular recorded data, it can be used much more broadly. Nelpy is intended to make interactive data analysis and exploration of these ephys data easy, by providing several convenience functions and common visualizations that operate directly on the nelpy objects.
More specifically, the functionality of this package includes:
- several container objects (
SpikeTrain,BinnedSpikeTrain,AnalogSignal,EpochArray, ...) with nice human-readable__repr__methods - powerful ways to interact with the data in the container objects
- hidden Markov model analysis of neural activity
- basic data exploration and visualization operating directly on the core nelpy objects
- and much more
Support
This work was supported by the National Science Foundation (CBET-1351692 and IOS-1550994) and the Human Frontiers Science Program (RGY0088). Any opinions, findings, and conclusions or recommendations expressed in this material are those of the author(s) and do not necessarily reflect the views of the National Science Foundation.
Quick examples
Let's give it a try. Create a SpikeTrainArray:
```python
import nelpy as nel # main nelpy imports
import nelpy.plotting as npl # optional plotting imports
spike_times = np.array([1, 2, 4, 5, 10])
st = nel.SpikeTrainArray(spike_times)
``` Do something:
```python
>>> print(st.n_spikes) # print out how many spikes there are in st
5
>>> print(st.supportn_spikes) # print out the underlying EpochArray on which st is defined
<EpochArray at 0x1d4812c7550: 1 epoch> of duration 9 seconds
>>> npl.raster(st) # plots the spike raster
```
As a more representative example of what nelpy can be used for, consider the estimation of place fields (spatial tuning curves) of CA1 units while an animal runs on a linear track.
Estimating the place fields can be a complicated affair, and roughly involves the following steps:
- assume we have position data and spike data available
- linearize the environment (and position data), if desired
- estimate the running velocity from the position data
- smooth the velocity estimates, since numerical differentiation is inherently noisy (and our measurements are imprecise)
- identify epochs where the animal was running, and where the animal was resting
- count the number of spikes from each unit, in each spatial bin of the environment, during run behavior
- determine how long the animal spent in each spatial bin (while running)
- estimate a firing rate within each spatial bin, by normalizing the number of observed spikes by the time spent in that spatial bin
- visualize the estimated tuning curves, and evaluate how well the tuning curves can be used to decode the animal's position
- ...
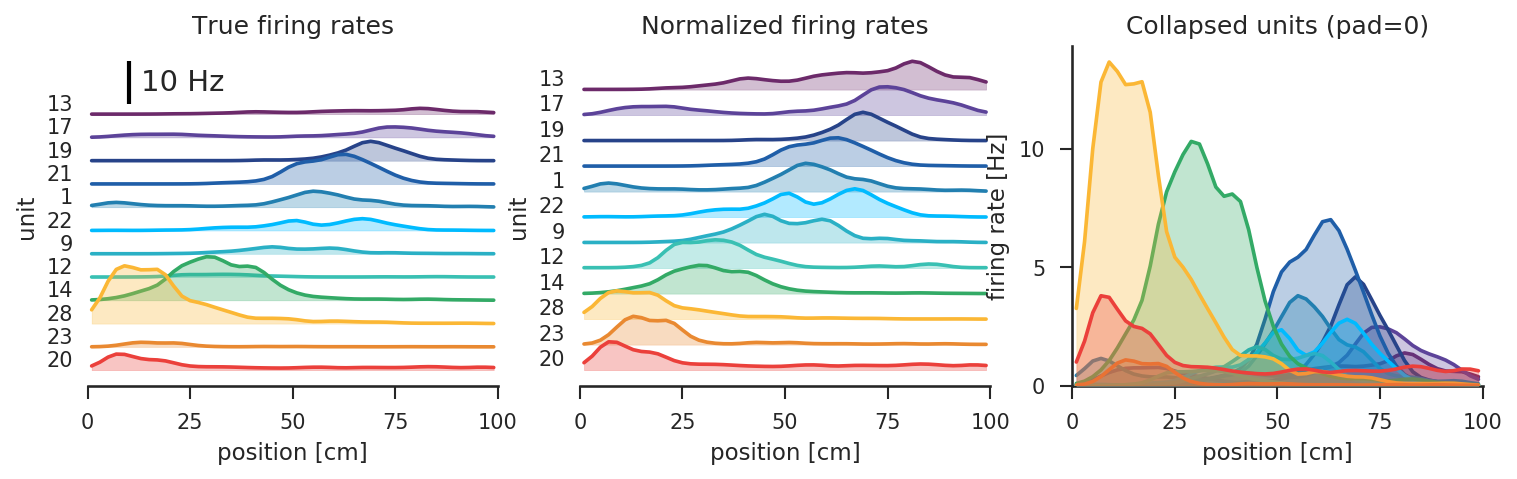
Nelpy makes it easy to do all of the above, to interact with the ephys data, and to visualize the results.
To see the full code that was used to generate the figures above, take a look at the linear track example analysis.
Getting started
The best way to get started with using nelpy is probably to take a look at
the tutorials (a work-in-progress) and
example analyses.
The tutorials are still pretty bare-bones, but will hopefully be expanded soon!
Installation
The easiest way to install nelpy is to use pip. From the terminal, run:
bash
$ pip install nelpy
Alternatively, you can install the latest version of nelpy by running the following commands:
bash
$ git clone https://github.com/nelpy/nelpy.git
$ cd nelpy
$ pip install .
If you would like to modify the code, then replace the last command with
bash
$ pip install -e .
A weak prerequisite for installing nelpy is a modified version of hmmlearn. This requirement is weak, in the sense that installation will complete successfully without it, and most of nelpy can also be used without any problems. However, as soon as any of the hidden Markov model (HMM) functions are used, you will get an error if the correct version of hmmlearn is not installed. To make things easier, there is a handy 64-bit Windows wheel in the hmmlearn directory of this repository. Installation on Linux/Unix should be almost trivial.
Related work and inspiration
Nelpy drew heavy inspiration from the python-vdmlab package (renamed to nept)
from the van der Meer lab at Dartmouth College (https://github.com/vandermeerlab),
which was created by Emily Irvine (https://github.com/emirvine). It is
also inspired by the neuralensemble.org NEO project (http://neo.readthedocs.io).
Short history: Etienne A started the nelpy project for two main reasons, namely
- he wanted / needed a
BinnedSpikeTrainobject for hidden Markov model analysis that wasn't (at the time) avaialable inneoorpython-vdmlab, and - he fundamentally wanted to add "support" attributes to all the container objects. Here "support" should be understood in the mathematical sense of "domain of definition", whereas the mathematical support technically would not include some elements for which the function maps to zero. This is critical for spike trains, for example, where it is important to differentiate "no spike at time t" from "no record at time t".
Scope of this work
The nelpy object model is expected to be quite similar to the python-vdmlab object model, which in turn has significant overlap with neuralensemble.org's neo model. However, the nelpy object model extends the former by making binned data first class citizens, and by changing the API for indexing and extracting subsets of data, as well as making "functional support" an integral part of the model. It (nelpy) is currently simpler and less comprehensive than neo, and specifically lacks in terms of physical units and complex object hierarchies and nonlinear relationships. However, nelpy again makes binned data a core object, and nelpy further aims to add additional analysis code including filtering, smoothing, position analysis, subsampling, interpolation, spike rate estimation, spike generation / synthesis, ripple detection, Bayesian decoding, and so on. In short, nelpy is more than just an object model, but the nelpy core is designed to be a flexible, readable, yet powerful object model for neuroelectrophysiology.
Where
=================== ======================================================== download https://pypi.python.org/pypi/nelpy tutorials https://github.com/nelpy/tutorials example analyses https://github.com/nelpy/example-analyses docs nelpy.github.io/nelpy/ code https://github.com/nelpy/nelpy =================== ========================================================
Cite
If you use nelpy in your research, please cite it:
@misc{nelpy17,
author = {Etienne Ackermann},
title = {Nelpy: Neuroelectrophysiology object model, data exploration, and analysis in Python},
howpublished = {\url{https://github.com/nelpy/nelpy/}},
year = {2017--2018}
}
License
Nelpy is distributed under the MIT license. See the LICENSE file for details.
GitHub Events
Total
- Issues event: 4
- Watch event: 5
- Delete event: 1
- Issue comment event: 12
- Push event: 35
- Pull request review comment event: 1
- Pull request review event: 2
- Pull request event: 25
- Fork event: 2
- Create event: 4
Last Year
- Issues event: 4
- Watch event: 5
- Delete event: 1
- Issue comment event: 12
- Push event: 35
- Pull request review comment event: 1
- Pull request review event: 2
- Pull request event: 25
- Fork event: 2
- Create event: 4
Committers
Last synced: over 2 years ago
Top Committers
| Name | Commits | |
|---|---|---|
| eackermann | e****a@r****u | 370 |
| Etienne Ackermann | E****n | 177 |
| eackermann | e****n@g****m | 112 |
| Joshua Chu | j****e@g****m | 69 |
| Etienne | e****3@r****u | 55 |
| shayok | s****a@g****m | 37 |
| Etienne Ackermann | e****n | 34 |
| Caleb Kemere | c****e@g****m | 2 |
| Sibo | u****o@g****m | 1 |
| iomidiran | m****n@g****m | 1 |
| Rice University Neuroengineering Lab | c****e@r****u | 1 |
Committer Domains (Top 20 + Academic)
Issues and Pull Requests
Last synced: 7 months ago
All Time
- Total issues: 102
- Total pull requests: 23
- Average time to close issues: 9 months
- Average time to close pull requests: 3 months
- Total issue authors: 7
- Total pull request authors: 3
- Average comments per issue: 2.38
- Average comments per pull request: 0.65
- Merged pull requests: 16
- Bot issues: 0
- Bot pull requests: 0
Past Year
- Issues: 2
- Pull requests: 20
- Average time to close issues: N/A
- Average time to close pull requests: about 20 hours
- Issue authors: 1
- Pull request authors: 1
- Average comments per issue: 0.0
- Average comments per pull request: 0.35
- Merged pull requests: 15
- Bot issues: 0
- Bot pull requests: 0
Top Authors
Issue Authors
- eackermann (75)
- jchutrue (18)
- ckemere (4)
- kushaangupta (2)
- avrodriguez (1)
- drsax93 (1)
- shayokdutta (1)
Pull Request Authors
- ryanharvey1 (20)
- kushaangupta (2)
- iomidiran (1)
Top Labels
Issue Labels
Pull Request Labels
Packages
- Total packages: 1
-
Total downloads:
- pypi 23 last-month
- Total dependent packages: 0
- Total dependent repositories: 1
- Total versions: 14
- Total maintainers: 1
pypi.org: nelpy
Neuroelectrophysiology object model and data analysis in Python.
- Homepage: https://github.com/nelpy/nelpy/
- Documentation: https://nelpy.readthedocs.io/
- License: MIT License
-
Latest release: 0.2.1
published about 7 years ago
Rankings
Maintainers (1)
Dependencies
- actions/cache v3 composite
- actions/checkout v3 composite
- actions/setup-python v4 composite
- actions/checkout v3 composite
- actions/setup-python v4 composite
- dill *
- matplotlib >=1.5.0
- numba >=0.56.0
- numpy >=1.16.0
- scikit-learn *
- scipy >=0.18.0