paperscraper
Tools to scrape publications & their metadata from pubmed, arxiv, medrxiv, biorxiv and chemrxiv.
Science Score: 49.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
○CITATION.cff file
-
✓codemeta.json file
Found codemeta.json file -
✓.zenodo.json file
Found .zenodo.json file -
✓DOI references
Found 2 DOI reference(s) in README -
✓Academic publication links
Links to: biorxiv.org, medrxiv.org, scholar.google, ncbi.nlm.nih.gov, wiley.com -
○Committers with academic emails
-
○Institutional organization owner
-
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (14.5%) to scientific vocabulary
Keywords
Keywords from Contributors
Repository
Tools to scrape publications & their metadata from pubmed, arxiv, medrxiv, biorxiv and chemrxiv.
Basic Info
- Host: GitHub
- Owner: jannisborn
- License: mit
- Language: Python
- Default Branch: main
- Homepage: https://jannisborn.github.io/paperscraper/
- Size: 2.49 MB
Statistics
- Stars: 412
- Watchers: 11
- Forks: 47
- Open Issues: 1
- Releases: 21
Topics
Metadata Files
README.md
paperscraper
paperscraper is a python package for scraping publication metadata or full text files (PDF or XML) from
PubMed or preprint servers such as arXiv, medRxiv, bioRxiv and chemRxiv.
It provides a streamlined interface to scrape metadata, allows to retrieve citation counts
from Google Scholar, impact factors from journals and comes with simple postprocessing functions
and plotting routines for meta-analysis.
Table of Contents
Getting started
console
pip install paperscraper
This is enough to query PubMed, arXiv or Google Scholar.
Download X-rxiv Dumps
However, to scrape publication data from the preprint servers biorxiv, medrxiv and chemrxiv, the setup is different. The entire dump is downloaded and stored in the server_dumps folder in a .jsonl format (one paper per line).
py
from paperscraper.get_dumps import biorxiv, medrxiv, chemrxiv
medrxiv() # Takes ~30min and should result in ~35 MB file
biorxiv() # Takes ~1h and should result in ~350 MB file
chemrxiv() # Takes ~45min and should result in ~20 MB file
NOTE: Once the dumps are stored, please make sure to restart the python interpreter so that the changes take effect.
NOTE: If you experience API connection issues (ConnectionError), since v0.2.12 there are automatic retries which you can even control and raise from the default of 10, as in biorxiv(max_retries=20).
Since v0.2.5 paperscraper also allows to scrape {med/bio/chem}rxiv for specific dates.
py
medrxiv(start_date="2023-04-01", end_date="2023-04-08")
But watch out. The resulting .jsonl file will be labelled according to the current date and all your subsequent searches will be based on this file only. If you use this option you might want to keep an eye on the source files (paperscraper/server_dumps/*jsonl) to ensure they contain the paper metadata for all papers you're interested in.
Arxiv local dump
If you prefer local search rather than using the arxiv API:
py
from paperscraper.get_dumps import arxiv
arxiv(start_date='2024-01-01', end_date=None) # scrapes all metadata from 2024 until today.
Afterwards you can search the local arxiv dump just like the other x-rxiv dumps.
The direct endpoint is paperscraper.arxiv.get_arxiv_papers_local. You can also specify the
backend directly in the get_and_dump_arxiv_papers function:
py
from paperscraper.arxiv import get_and_dump_arxiv_papers
get_and_dump_arxiv_papers(..., backend='local')
Examples
paperscraper is build on top of the packages arxiv, pymed, and scholarly.
Publication keyword search
Consider you want to perform a publication keyword search with the query:
COVID-19 AND Artificial Intelligence AND Medical Imaging.
- Scrape papers from PubMed:
```py from paperscraper.pubmed import getanddumppubmedpapers covid19 = ['COVID-19', 'SARS-CoV-2'] ai = ['Artificial intelligence', 'Deep learning', 'Machine learning'] mi = ['Medical imaging'] query = [covid19, ai, mi]
getanddumppubmedpapers(query, outputfilepath='covid19ai_imaging.jsonl') ```
- Scrape papers from arXiv:
```py from paperscraper.arxiv import getanddumparxivpapers
getanddumparxivpapers(query, outputfilepath='covid19ai_imaging.jsonl') ```
- Scrape papers from bioRiv, medRxiv or chemRxiv:
```py from paperscraper.xrxiv.xrxiv_query import XRXivQuery
querier = XRXivQuery('serverdumps/chemrxiv2020-11-10.jsonl') querier.searchkeywords(query, outputfilepath='covid19aiimaging.jsonl') ```
You can also use dump_queries to iterate over a bunch of queries for all available databases.
```py from paperscraper import dump_queries
queries = [[covid19, ai, mi], [covid19, ai], [ai]] dump_queries(queries, '.') ```
Or use the harmonized interface of QUERY_FN_DICT to query multiple databases of your choice:
```py
from paperscraper.loaddumps import QUERYFNDICT
print(QUERYFN_DICT.keys())
QUERYFNDICT'biorxiv' QUERYFNDICT'medrxiv' ```
- Scrape papers from Google Scholar:
Thanks to scholarly, there is an endpoint for Google Scholar too. It does not understand Boolean expressions like the others, but should be used just like the Google Scholar search fields.
py
from paperscraper.scholar import get_and_dump_scholar_papers
topic = 'Machine Learning'
get_and_dump_scholar_papers(topic)
NOTE: The scholar endpoint does not require authentication but since it regularly prompts with captchas, it's difficult to apply large scale.
Full-Text Retrieval (PDFs & XMLs)
paperscraper allows you to download full text of publications using DOIs. The basic functionality works reliably for preprint servers (arXiv, bioRxiv, medRxiv, chemRxiv), but retrieving papers from PubMed dumps is more challenging due to publisher restrictions and paywalls.
Standard Usage
The main download functions work for all paper types with automatic fallbacks:
py
from paperscraper.pdf import save_pdf
paper_data = {'doi': "10.48550/arXiv.2207.03928"}
save_pdf(paper_data, filepath='gt4sd_paper.pdf')
To batch download full texts from your metadata search results:
```py from paperscraper.pdf import savepdffrom_dump
Save PDFs/XMLs in current folder and name the files by their DOI
savepdffromdump('medrxivcovidaiimaging.jsonl', pdfpath='.', keyto_save='doi') ```
Automatic Fallback Mechanisms
When the standard text retrieval fails, paperscraper automatically tries these fallbacks:
- BioC-PMC: For biomedical papers in PubMed Central (open-access repository), it retrieves open-access full-text XML from the BioC-PMC API.
- eLife Papers: For eLife journal papers, it fetches XML files from eLife's open GitHub repository.
These fallbacks are tried automatically without requiring any additional configuration.
Enhanced Retrieval with Publisher APIs
For more comprehensive access to papers from major publishers, you can provide API keys for:
- Wiley TDM API: Enables access to Wiley publications (2,000+ journals).
- Elsevier TDM API: Enables access to Elsevier publications (The Lancet, Cell, ...).
- bioRxiv TDM API Enable access to bioRxiv publications (since May 2025 bioRxiv is protected with Cloudflare)
To use publisher APIs:
Create a file with your API keys:
WILEY_TDM_API_TOKEN=your_wiley_token_here ELSEVIER_TDM_API_KEY=your_elsevier_key_here AWS_ACCESS_KEY_ID=your_aws_access_key_here AWS_SECRET_ACCESS_KEY=your_aws_secret_key_hereNOTE: The AWS keys can be created in your AWS/IAM account. When creating the key, make sure you tick theAmazonS3ReadOnlyAccesspermission policy. NOTE: If you name the file.envit will be loaded automatically (if it is in the cwd or anywhere above the tree to home).Pass the file path when calling retrieval functions:
```py from paperscraper.pdf import savepdffrom_dump
savepdffromdump( 'pubmedqueryresults.jsonl', pdfpath='./papers', keytosave='doi', apikeys='path/to/your/apikeys.txt' ) ```
For obtaining API keys: - Wiley TDM API: Visit Wiley Text and Data Mining (free for academic users with institutional subscription) - Elsevier TDM API: Visit Elsevier's Text and Data Mining (free for academic users with institutional subscription)
NOTE: While these fallback mechanisms improve retrieval success rates, they cannot guarantee access to all papers due to various access restrictions.
Citation search
You can fetch the number of citations of a paper from its title or DOI
```py from paperscraper.citations import getcitationsfromtitle, getcitationsbydoi title = 'Über formal unentscheidbare Sätze der Principia Mathematica und verwandter Systeme I.' print(getcitationsfrom_title(title))
doi = '10.1021/acs.jcim.3c00132' getcitationsby_doi(doi) ```
Journal impact factor
You can also retrieve the impact factor for all journals: ```py
from paperscraper.impact import Impactor i = Impactor() i.search("Nat Comms", threshold=85, sort_by='impact') [ {'journal': 'Nature Communications', 'factor': 17.694, 'score': 94}, {'journal': 'Natural Computing', 'factor': 1.504, 'score': 88} ]
This performs a fuzzy search with a threshold of 85. `threshold` defaults to 100 in which case an exact search is performed. You can also search by journal abbreviation, [E-ISSN](https://portal.issn.org) or [NLM ID](https://portal.issn.org).py i.search("Nat Rev Earth Environ") # [{'journal': 'Nature Reviews Earth & Environment', 'factor': 37.214, 'score': 100}] i.search("101771060") # [{'journal': 'Nature Reviews Earth & Environment', 'factor': 37.214, 'score': 100}] i.search('2662-138X') # [{'journal': 'Nature Reviews Earth & Environment', 'factor': 37.214, 'score': 100}]
Filter results by impact factor
i.search("Neural network", threshold=85, minimpact=1.5, maximpact=20)
[
{'journal': 'IEEE Transactions on Neural Networks and Learning Systems', 'factor': 14.255, 'score': 93},
{'journal': 'NEURAL NETWORKS', 'factor': 9.657, 'score': 91},
{'journal': 'WORK-A Journal of Prevention Assessment & Rehabilitation', 'factor': 1.803, 'score': 86},
{'journal': 'NETWORK-COMPUTATION IN NEURAL SYSTEMS', 'factor': 1.5, 'score': 92}
]
Show all fields
i.search("quantum information", threshold=90, return_all=True)
[
{'factor': 10.758, 'jcr': 'Q1', 'journalabbr': 'npj Quantum Inf', 'eissn': '2056-6387', 'journal': 'npj Quantum Information', 'nlmid': '101722857', 'issn': '', 'score': 92},
{'factor': 1.577, 'jcr': 'Q3', 'journalabbr': 'Nation', 'eissn': '0027-8378', 'journal': 'NATION', 'nlmid': '9877123', 'issn': '0027-8378', 'score': 91}
]
```
Plotting
When multiple query searches are performed, two types of plots can be generated automatically: Venn diagrams and bar plots.
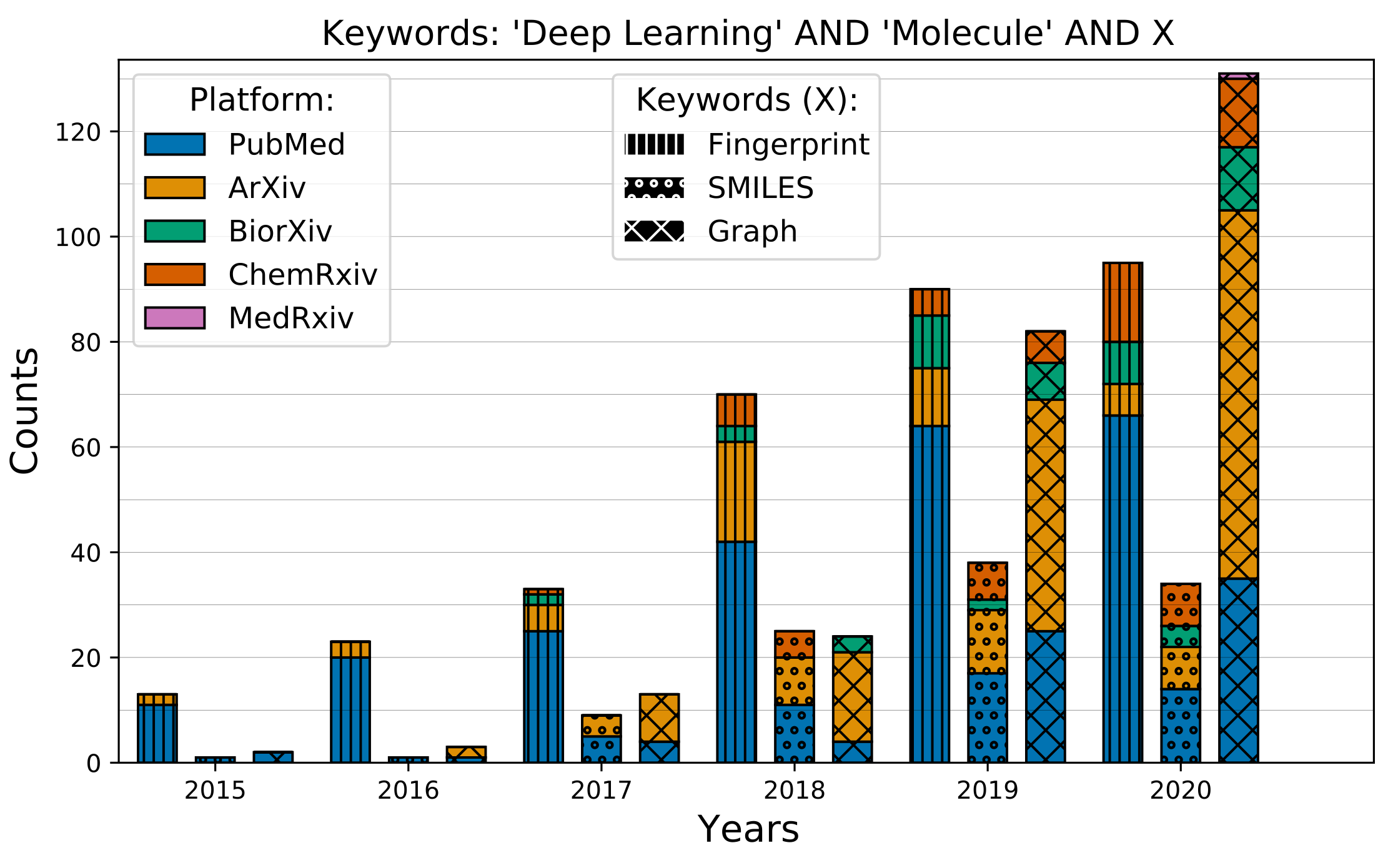
Barplots
Compare the temporal evolution of different queries across different servers.
```py from paperscraper import QUERYFNDICT from paperscraper.postprocessing import aggregatepaper from paperscraper.utils import getfilenamefromquery, load_jsonl
Define search terms and their synonyms
ml = ['Deep learning', 'Neural Network', 'Machine learning'] mol = ['molecule', 'molecular', 'drug', 'ligand', 'compound'] gnn = ['gcn', 'gnn', 'graph neural', 'graph convolutional', 'molecular graph'] smiles = ['SMILES', 'Simplified molecular'] fp = ['fingerprint', 'molecular fingerprint', 'fingerprints']
Define queries
queries = [[ml, mol, smiles], [ml, mol, fp], [ml, mol, gnn]]
root = '../keyword_dumps'
datadict = dict() for query in queries: filename = getfilenamefromquery(query) datadict[filename] = dict() for db, in QUERYFNDICT.items(): # Assuming the keyword search has been performed already data = load_jsonl(os.path.join(root, db, filename))
# Unstructured matches are aggregated into 6 bins, 1 per year
# from 2015 to 2020. Sanity check is performed by having
# `filtering=True`, removing papers that don't contain all of
# the keywords in query.
data_dict[filename][db], filtered = aggregate_paper(
data, 2015, bins_per_year=1, filtering=True,
filter_keys=query, return_filtered=True
)
Plotting is now very simple
from paperscraper.plotting import plot_comparison
datakeys = [ 'deeplearningmoleculefingerprint.jsonl', 'deeplearningmoleculesmiles.jsonl', 'deeplearningmoleculegcn.jsonl' ] plotcomparison( datadict, datakeys, titletext="'Deep Learning' AND 'Molecule' AND X", keywordtext=['Fingerprint', 'SMILES', 'Graph'], figpath='mol_representation' ) ```
Venn Diagrams
```py from paperscraper.plotting import ( plotvenntwo, plotvennthree, plotmultiplevenn )
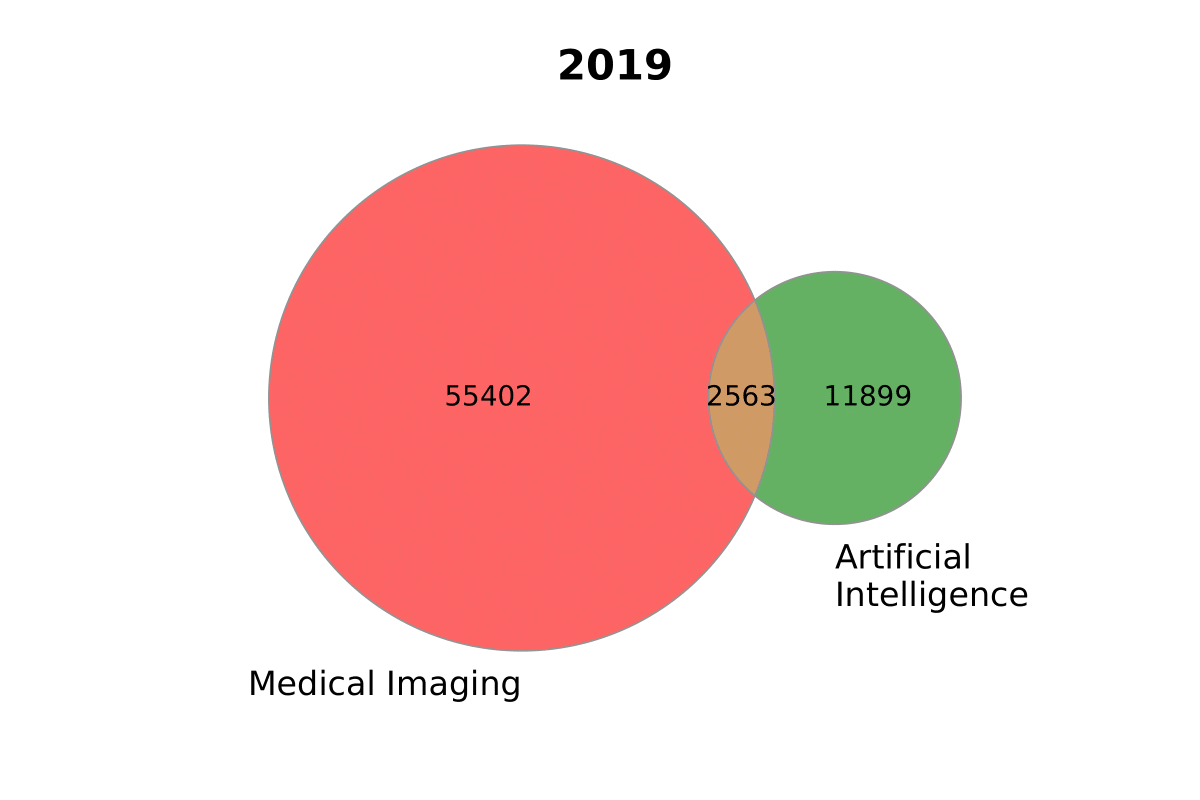
sizes2020 = (30842, 14474, 2292, 35476, 1904, 1408, 376) sizes2019 = (55402, 11899, 2563) labels2020 = ('Medical\nImaging', 'Artificial\nIntelligence', 'COVID-19') labels2019 = ['Medical Imaging', 'Artificial\nIntelligence']
plotvenntwo(sizes2019, labels2019, title='2019', figname='ai_imaging') ```
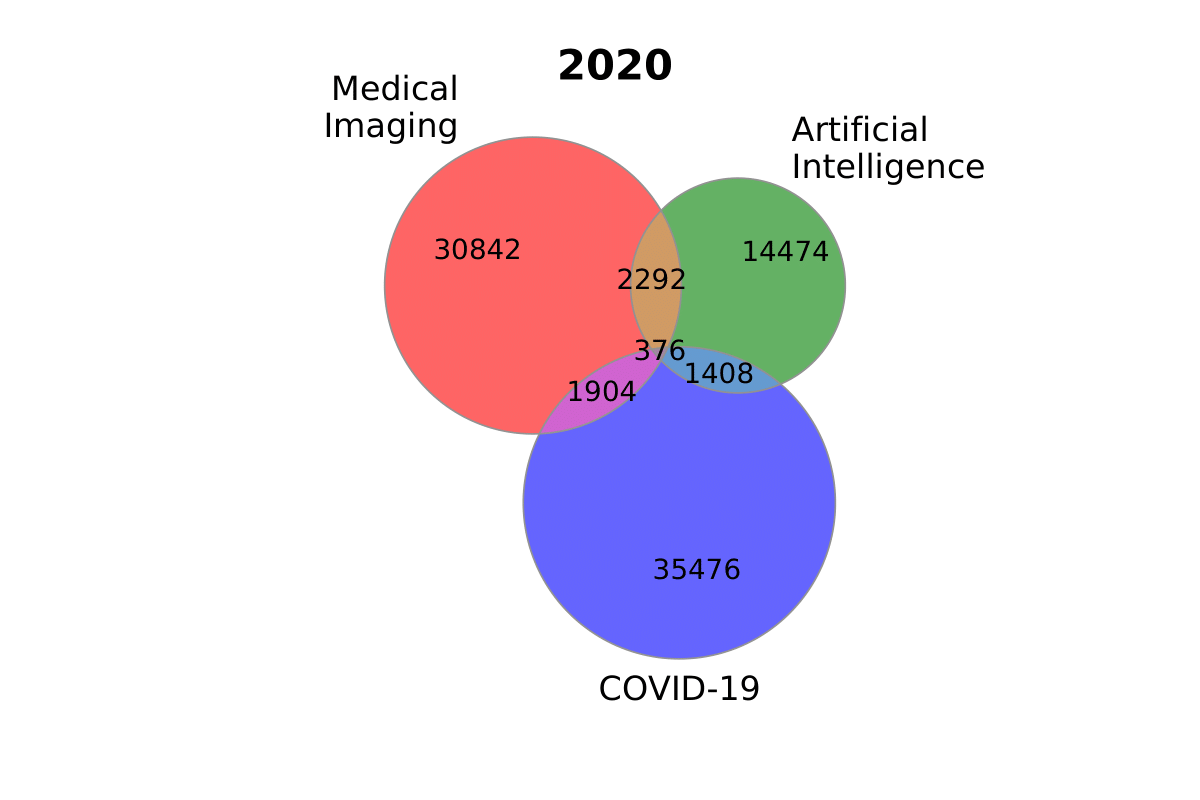
py
plot_venn_three(
sizes_2020, labels_2020, title='2020', figname='ai_imaging_covid'
)
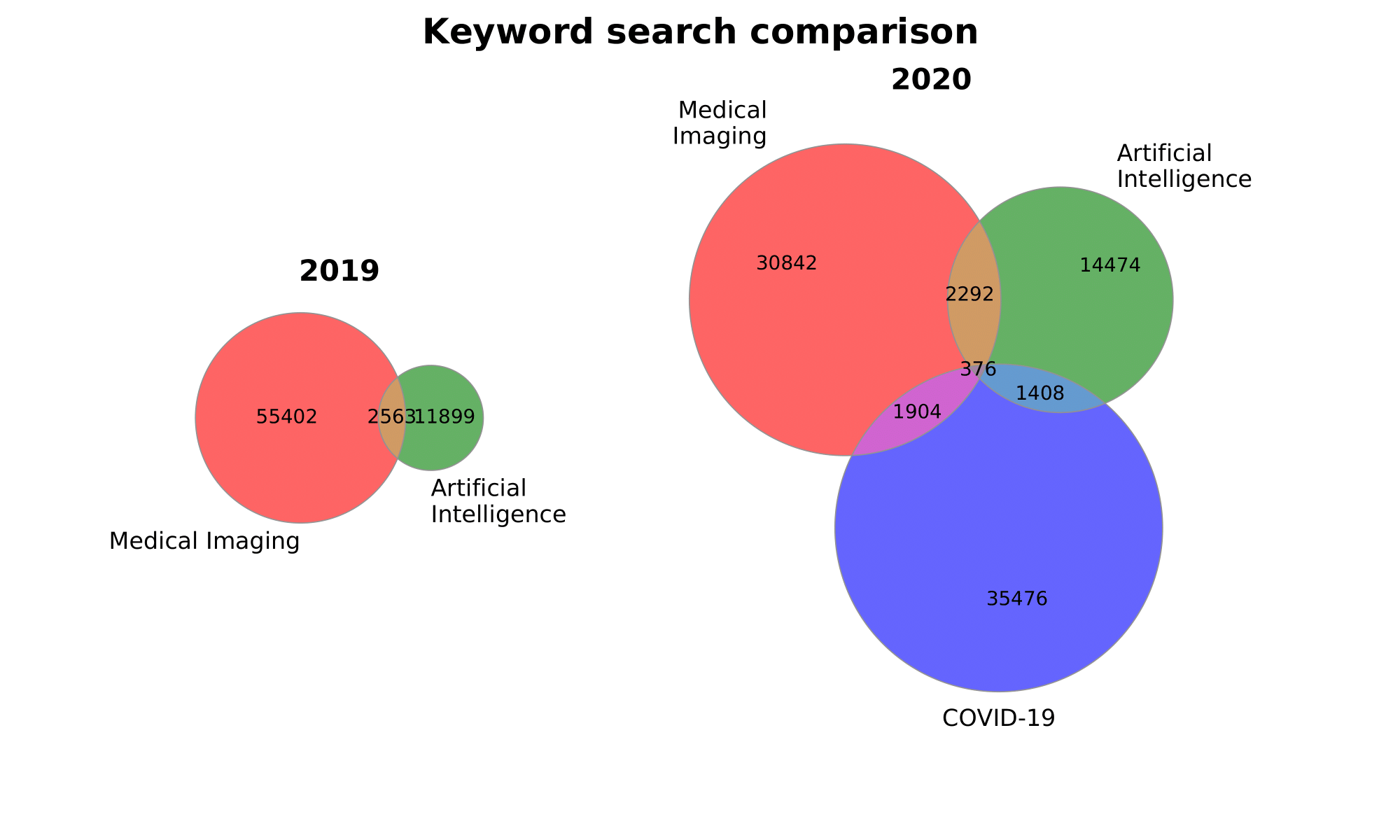
Or plot both together:
py
plot_multiple_venn(
[sizes_2019, sizes_2020], [labels_2019, labels_2020],
titles=['2019', '2020'], suptitle='Keyword search comparison',
gridspec_kw={'width_ratios': [1, 2]}, figsize=(10, 6),
figname='both'
)
Citation
If you use paperscraper, please cite a paper that motivated our development of this tool.
bibtex
@article{born2021trends,
title={Trends in Deep Learning for Property-driven Drug Design},
author={Born, Jannis and Manica, Matteo},
journal={Current Medicinal Chemistry},
volume={28},
number={38},
pages={7862--7886},
year={2021},
publisher={Bentham Science Publishers}
}
Contributions
Thanks to the following contributors:
- @mathinic: Since v0.3.0 improved PubMed full text retrieval with additional fallback mechanisms (BioC-PMC, eLife and optional Wiley/Elsevier APIs).
@memray: Since
v0.2.12there are automatic retries when downloading the {med/bio/chem}rxiv dumps.@achouhan93: Since
v0.2.5{med/bio/chem}rxiv can be scraped for specific dates!@daenuprobst: Since
v0.2.4PDF files can be scraped directly (paperscraper.pdf.save_pdf)@oppih: Since
v0.2.3chemRxiv API also provides DOI and URL if available@lukasschwab: Enabled support for
arxiv>1.4.2in paperscraperv0.1.0.@juliusbierk: Bugfixes
Owner
- Name: Jannis Born
- Login: jannisborn
- Kind: user
- Location: Zurich
- Company: @IBM Research
- Repositories: 44
- Profile: https://github.com/jannisborn
Researcher @IBM. Previous @ETH. AI 4 Science, Language Models, Quantum ML
GitHub Events
Total
- Create event: 25
- Release event: 6
- Issues event: 24
- Watch event: 157
- Delete event: 18
- Member event: 1
- Issue comment event: 59
- Push event: 96
- Pull request review comment event: 18
- Pull request review event: 14
- Pull request event: 38
- Fork event: 18
Last Year
- Create event: 25
- Release event: 6
- Issues event: 24
- Watch event: 157
- Delete event: 18
- Member event: 1
- Issue comment event: 59
- Push event: 96
- Pull request review comment event: 18
- Pull request review event: 14
- Pull request event: 38
- Fork event: 18
Committers
Last synced: 10 months ago
Top Committers
| Name | Commits | |
|---|---|---|
| Jannis Born | j****b@z****m | 104 |
| Matteo Manica | d****g@g****m | 16 |
| dependabot[bot] | 4****] | 2 |
| OPPIH.X | l****8@g****m | 2 |
| Ashish Chouhan | 3****3 | 2 |
| Yaroslav Halchenko | d****n@o****m | 1 |
| Rui Meng | m****0@g****m | 1 |
| Nicolas Mathis | 5****c | 1 |
| Lukas Schwab | l****b@g****m | 1 |
| Julius Bier Kirkegaard | j****k@g****m | 1 |
Committer Domains (Top 20 + Academic)
Issues and Pull Requests
Last synced: 7 months ago
All Time
- Total issues: 32
- Total pull requests: 65
- Average time to close issues: about 1 month
- Average time to close pull requests: 2 days
- Total issue authors: 22
- Total pull request authors: 10
- Average comments per issue: 2.72
- Average comments per pull request: 1.25
- Merged pull requests: 62
- Bot issues: 0
- Bot pull requests: 2
Past Year
- Issues: 14
- Pull requests: 34
- Average time to close issues: 10 days
- Average time to close pull requests: 3 days
- Issue authors: 9
- Pull request authors: 4
- Average comments per issue: 2.0
- Average comments per pull request: 1.41
- Merged pull requests: 31
- Bot issues: 0
- Bot pull requests: 0
Top Authors
Issue Authors
- jannisborn (9)
- mathinic (2)
- vmkalbskopf (2)
- mohammad-gh009 (1)
- dongwonmoon (1)
- yiouyou (1)
- TestF15 (1)
- peiyaoli (1)
- MorganRO8 (1)
- richardehughes (1)
- mwarqee (1)
- sleepingcat4 (1)
- pathway (1)
- xlhd19980327 (1)
- arifin-chemist89 (1)
Pull Request Authors
- jannisborn (59)
- dependabot[bot] (3)
- MoonDavid (2)
- mathinic (2)
- oppih (2)
- yarikoptic (2)
- drugilsberg (1)
- juliusbierk (1)
- memray (1)
- achouhan93 (1)
- lukasschwab (1)
Top Labels
Issue Labels
Pull Request Labels
Packages
- Total packages: 1
-
Total downloads:
- pypi 1,388 last-month
- Total dependent packages: 0
- Total dependent repositories: 2
- Total versions: 26
- Total maintainers: 2
pypi.org: paperscraper
paperscraper: Package to scrape papers.
- Homepage: https://github.com/jannisborn/paperscraper
- Documentation: https://paperscraper.readthedocs.io/
- License: MIT
-
Latest release: 0.3.2
published 8 months ago
Rankings
Maintainers (2)
Dependencies
- arxiv >=1.4.2
- bs4 >=0.0.1
- matplotlib >=3.3.2
- matplotlib-venn >=0.11.5
- pandas >=1.0.4
- pymed ==0.8.9
- requests ==2.24.0
- scholarly ==0.5.1
- seaborn >=0.11.0
- tqdm >=4.51.0
- arxiv >=1.4.2
- bs4 *
- matplotlib *
- matplotlib_venn *
- pandas *
- pymed *
- requests *
- scholarly ==0.5.1
- seaborn *
- tqdm *
- actions/checkout v2 composite
- actions/setup-python v2 composite
- actions/setup-python v2 composite
- actions/checkout v2 composite
- actions/setup-python v2 composite
- actions/checkout v4 composite
- codespell-project/actions-codespell v2 composite
- codespell-project/codespell-problem-matcher v1 composite

