ParaDigMa: a Python toolbox for extracting Parkinson's disease digital biomarkers from daily life wrist sensor data
ParaDigMa: a Python toolbox for extracting Parkinson's disease digital biomarkers from daily life wrist sensor data - Published in JOSS (2026)
Science Score: 92.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
✓CITATION.cff file
Found CITATION.cff file -
○codemeta.json file
-
○.zenodo.json file
-
✓DOI references
Found 23 DOI reference(s) in README and JOSS metadata -
✓Academic publication links
Links to: medrxiv.org, zenodo.org -
○Academic email domains
-
○Institutional organization owner
-
✓JOSS paper metadata
Published in Journal of Open Source Software
Keywords
Repository
Digital Biomarkers for Parkinson's Disease Toolbox.
Basic Info
- Host: GitHub
- Owner: biomarkersParkinson
- License: apache-2.0
- Language: Python
- Default Branch: main
- Homepage: https://biomarkersparkinson.github.io/paradigma/
- Size: 207 MB
Statistics
- Stars: 10
- Watchers: 2
- Forks: 1
- Open Issues: 20
- Releases: 17
Topics
Metadata Files
README.md

| Badges | |
|:----:|----|
| Packages and Releases | 


|
| Build Status |

|
| License |

Overview
ParaDigMa (Parkinson's disease Digital Markers) is a Python toolbox for extracting validated digital biomarkers from wrist sensor data in Parkinson's disease. It processes accelerometer, gyroscope, and PPG signals collected during passive monitoring in daily life.
Key Features: - Arm swing during gait analysis - Tremor analysis - Pulse rate analysis - Scientifically validated in peer-reviewed publications - Modular, extensible architecture for custom analyses
Quick Start
Installation
For regular use:
bash
pip install paradigma
Requires Python 3.11+.
For development or running tutorials:
Example data requires git-lfs. See the installation guide for setup instructions.
Basic Usage
```python from paradigma.orchestrator import run_paradigma
Example 1: Single DataFrame with default output directory
results = runparadigma( dfs=df, pipelines=['gait', 'tremor'], watchside='left', # Required for gait pipeline save_intermediate=['quantification', 'aggregation'] # Saves to ./output by default )
Example 2: Multiple DataFrames as list (assigned to 'df1', 'df2', etc.)
results = runparadigma( dfs=[df1, df2, df3], pipelines=['gait', 'tremor'], outputdir="./results", # Custom output directory watchside='left', saveintermediate=['quantification', 'aggregation'] )
Example 3: Dictionary of DataFrames (custom segment/file names)
results = runparadigma( dfs={'morningsession': df1, 'eveningsession': df2}, pipelines=['gait', 'tremor'], watchside='right', save_intermediate=[] # No files saved - results only in memory )
Example 4: Load from data directory
results = runparadigma( datapath='./mydata', pipelines=['gait', 'tremor'], watchside='left', filepattern='*.parquet', saveintermediate=['quantification', 'aggregation'] )
Access results (nested by pipeline)
For gait, results are nested by filtered/unfiltered
gaitfiltered = results['quantifications']['gait']['filtered'] gaitunfiltered = results['quantifications']['gait']['unfiltered'] tremormeasures = results['quantifications']['tremor'] gaitaggregates = results['aggregations']['gait'] # Contains 'filtered' and 'unfiltered' keys tremor_aggregates = results['aggregations']['tremor']
Check for errors
if results['errors']: print(f"Warning: {len(results['errors'])} error(s) occurred") ```
See our tutorials for complete examples.
Pipelines
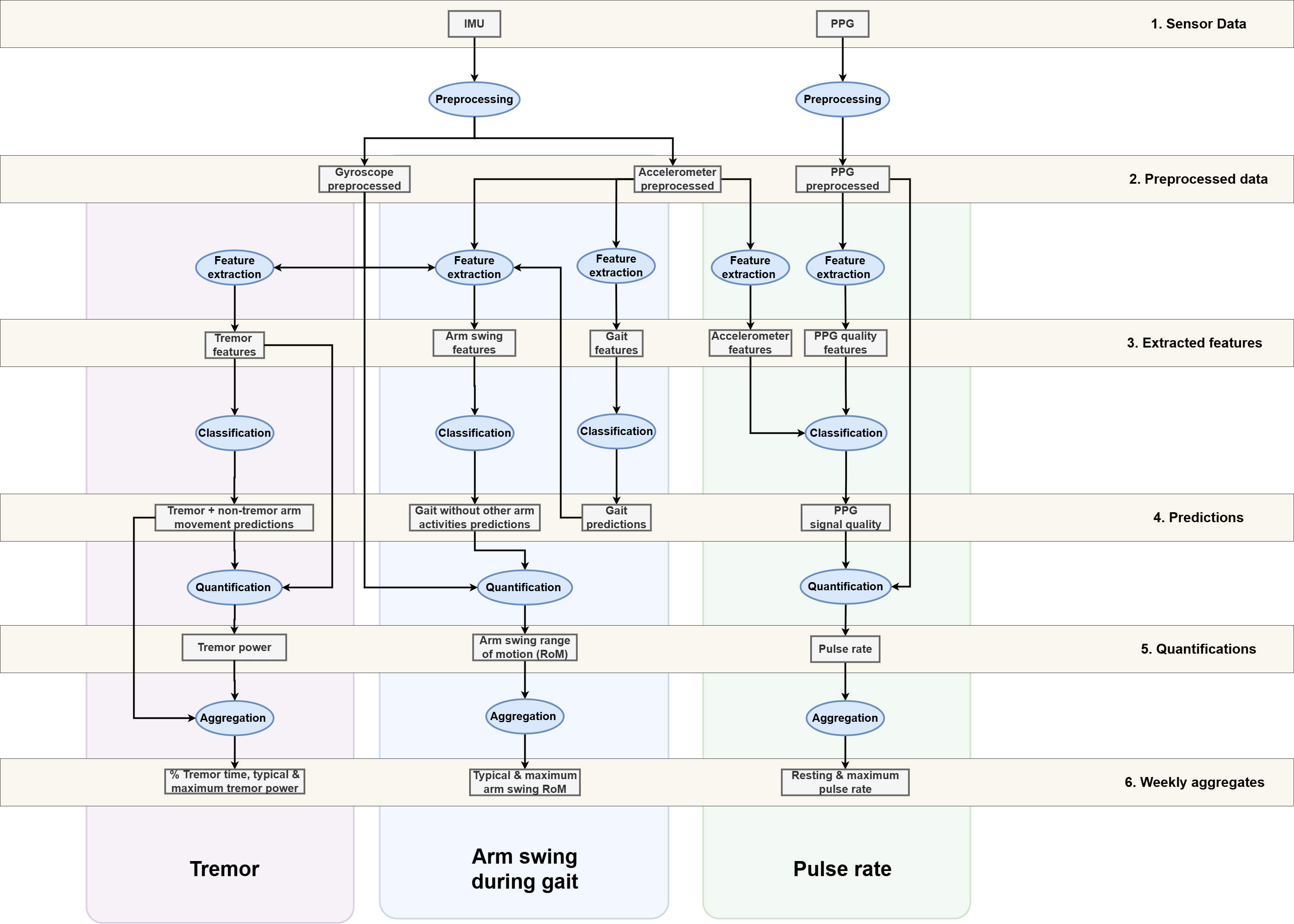
Validated Processing Pipelines
| Pipeline | Input sensors | Output week-level aggregation | Publications | Tutorial | | ---- | ---- | ------- | ---- | ---- | | Arm swing during gait | Accelerometer + Gyroscope | Typical, maximum & variability of arm swing range of motion | Post 2025, Post 2026* | Guide | | Tremor | Gyroscope | % tremor time, typical & maximum tremor power | Timmermans 2025a, Timmermans 2025b* | Guide | | Pulse rate | PPG (+ Accelerometer) | Resting & maximum pulse rate | Veldkamp 2025* | Guide |
* Indicates pre-print
Pipeline Architecture
ParaDigMa can best be understood by categorizing the sequential processes: | Process | Description | | ---- | ---- | | Preprocessing | Preparing raw sensor signals for further processing | | Feature extraction | Extracting features based on windowed sensor signals | | Classification | Detecting segments of interest using validated classifiers (e.g., gait segments) | | Quantification | Extracting specific measures from the detected segments (e.g., arm swing measures) | | Aggregation | Aggregating the measures over a specific time period (e.g., week-level aggregates)
Usage
Documentation
- Tutorials - Step-by-step usage examples
- Installation Guide - Setup and troubleshooting
- Sensor Data Requirements - Data specifications and compliance
- Supported Devices - Validated hardware
- Input Formats Guide - Input format options and data loading
- Configuration Guide - Pipeline configuration
- API Reference - Complete API documentation
Sensor Requirements & Supported Devices
ParaDigMa is designed for wrist sensor data collected during passive monitoring in persons with Parkinson's disease. While designed to work with any compliant device, it has been empirically validated on:
- Verily Study Watch (gait, tremor, pulse rate)
- Axivity AX6 (gait, tremor)
- Gait-up Physilog 4 (gait, tremor)
- Empatica EmbracePlus (data loading)
Please check before running the pipelines whether your sensor data complies with the requirements for the sensor configuration and context of use. See the sensor requirements guide for data specifications and the supported devices guide for device-specific setup instructions.
Data Formats
ParaDigMa supports the following data formats:
- In-memory (recommended): Pandas DataFrames (see examples above)
- Data loading file extensions: TSDF, Parquet, CSV, Pickle and several device-specific formats (AVRO (Empatica), CWA (Axivity))
Troubleshooting
For installation issues, see the installation guide troubleshooting section.
For other issues, check our issue tracker or contact paradigma@radboudumc.nl.
Scientific Validation
The following publications contain details and validation of the pipelines:
Arm swing during gait * Post, E. et al. (2025) - Quantifying arm swing in Parkinson's disease: a method account for arm activities during free-living gait * Post, E. et al. (2026) - (Pre-print) Longitudinal progression of digital arm swing measures during free-living gait in early Parkinson's disease
Tremor * Timmermans, N.A. et al. (2025) - A generalizable and open-source algorithm for real-life monitoring of tremor in Parkinson's disease * Timmermans, N.A. et al. (2025) - (Pre-print) Progression of daily-life tremor measures in early Parkinson disease: a longitudinal continuous monitoring study
Contributing
We welcome contributions! Please see:
Citation
If you use ParaDigMa in your research, please cite:
bibtex
@software{paradigma2024,
author = {Post, Erik and Veldkamp, Kars and Timmermans, Nienke and
Soriano, Diogo Coutinho and Kasalica, Vedran and
Kok, Peter and Evers, Luc},
title = {ParaDigMa: Parkinson's disease Digital Markers},
year = {2024},
doi = {10.5281/zenodo.13838392},
url = {https://github.com/biomarkersParkinson/paradigma}
}
License
Licensed under the Apache License 2.0. See LICENSE for details.
Acknowledgements
Core Team: Erik Post, Kars Veldkamp, Nienke Timmermans, Diogo Coutinho Soriano, Vedran Kasalica, Peter Kok, Twan van Laarhoven, Luc Evers
Advisors: Max Little, Jordan Raykov, Hayriye Cagnan, Bas Bloem
Funding: the initial release was funded by the Michael J Fox Foundation (grant #020425) and the Dutch Research Council (grants #ASDI.2020.060, #2023.010)
Contact
- Email: paradigma@radboudumc.nl
- Issue Tracker
Owner
- Name: Digital Biomarkers for Parkinson's disease
- Login: biomarkersParkinson
- Kind: organization
- Location: Netherlands
- Repositories: 1
- Profile: https://github.com/biomarkersParkinson
JOSS Publication
ParaDigMa: a Python toolbox for extracting Parkinson's disease digital biomarkers from daily life wrist sensor data
Authors

Center of Expertise for Parkinson and Movement Disorders, Department of Neurology, Donders Institute for Brain, Cognition and Behaviour, Radboud University Medical Center, Nijmegen, the Netherlands, Institute for Computing and Information Sciences, Radboud University, Nijmegen, the Netherlands

Center of Expertise for Parkinson and Movement Disorders, Department of Neurology, Donders Institute for Brain, Cognition and Behaviour, Radboud University Medical Center, Nijmegen, the Netherlands

Center of Expertise for Parkinson and Movement Disorders, Department of Neurology, Donders Institute for Brain, Cognition and Behaviour, Radboud University Medical Center, Nijmegen, the Netherlands

Center for Engineering, Modeling and Applied Social Sciences (CECS), Federal University of ABC (UFABC), Brazil

Center of Expertise for Parkinson and Movement Disorders, Department of Neurology, Donders Institute for Brain, Cognition and Behaviour, Radboud University Medical Center, Nijmegen, the Netherlands

School of Mathematical Sciences, University of Nottingham, Nottingham, United Kingdom, Horizon Digital Economy Research Institute, University of Nottingham, Nottingham, United Kingdom

Institute for Computing and Information Sciences, Radboud University, Nijmegen, the Netherlands

Center of Expertise for Parkinson and Movement Disorders, Department of Neurology, Donders Institute for Brain, Cognition and Behaviour, Radboud University Medical Center, Nijmegen, the Netherlands, Institute for Computing and Information Sciences, Radboud University, Nijmegen, the Netherlands
Citation (CITATION.cff)
cff-version: 1.3.1
title: >-
ParaDigMa: A toolbox for deriving Parkinson's disease Digital Markers from real-life wrist sensor data
message: >-
If you use this software, please cite it using the
metadata from this file.
type: software
version: 1.1.0
date-released: '2026-02-04'
authors:
- given-names: Erik
family-names: Post
email: erik.post@radboudumc.nl
affiliation: >-
Radboud University, Nijmegen, Netherlands; Radboud
University Medical Center, Nijmegen, Netherlands
orcid: 'https://orcid.org/0000-0002-1937-9976'
- given-names: Kars
family-names: Veldkamp
email: Kars.Veldkamp@radboudumc.nl
affiliation: >-
Radboud University Medical Center, Nijmegen,
Netherlands
orcid: 'https://orcid.org/0009-0007-3498-3155'
- given-names: Nienke
family-names: Timmermans
email: nienke.timmermans@radboudumc.nl
affiliation: >-
Radboud University Medical Center, Nijmegen,
Netherlands
orcid: 'https://orcid.org/0009-0001-3814-3440'
- given-names: Diogo
family-names: Soriano
email: diogo.soriano@ufabc.edu.br
affiliation: >-
Center for Engineering, Modeling and Applied Social
Sciences (CECS), Federal University of ABC (UFABC),
Brazil
- given-names: Vedran
family-names: Kasalica
email: v.kasalica@esciencecenter.nl
affiliation: Netherlands eScience Center
orcid: 'https://orcid.org/0000-0002-0097-1056'
- given-names: Peter
family-names: Kok
email: p.kok@esciencecenter.nl
affiliation: Netherlands eScience Center
orcid: 'https://orcid.org/0000-0002-6630-7326'
- given-names: Hayriye
family-names: Cagnan
email: h.cagnan@imperial.ac.uk
affiliation: Imperial College London
orcid: 'https://orcid.org/0000-0002-1641-115X'
- given-names: Bastiaan R.
family-names: Bloem
email: bas.bloem@radboudumc.nl
affiliation: >-
Radboud University Medical Center, Nijmegen,
Netherlands
- given-names: Max A.
family-names: Little
email: max.a.little@gmail.com
affiliation: University of Birmingham
- given-names: Jordan P.
family-names: Raykov
email: yordan.raykov@gmail.com
affiliation: University of Nottingham
orcid: 'https://orcid.org/0000-0003-0753-717X'
- given-names: Twan
family-names: van Laarhoven
email: tvanlaarhoven@cs.ru.nl
affiliation: Radboud University, Nijmegen, Netherlands
orcid: 'https://orcid.org/0000-0001-7597-0579'
- given-names: Luc J.W.
family-names: Evers
email: luc.evers@radboudumc.nl
affiliation: >-
Radboud University, Nijmegen, Netherlands; Radboud
University Medical Center, Nijmegen, Netherlands
orcid: 'https://orcid.org/0000-0002-8241-5087'
repository-code: 'https://github.com/biomarkersParkinson/paradigma'
url: 'https://biomarkersparkinson.github.io/paradigma/'
abstract: >-
The Parkinson's disease Digital Markers (ParaDigMa) toolbox is a Python software
package designed for processing and analyzing real-life wrist sensor data to
extract digital measures of motor and non-motor signs of Parkinson's disease (PD).
The toolbox processes accelerometer, gyroscope and photoplethysmography signals
collected during passive monitoring in daily life. It contains three scientifically
validated data processing pipelines: (1) arm swing during gait, (2) tremor, and
(3) pulse rate analysis. An orchestrator function enables end-to-end processing from
raw data loading to aggregated measures, with automatic data preparation including
flexible column mapping and sensor orientation adjustment. The toolbox also provides
general functionalities for signal processing and feature extraction, such as
filtering, peak detection, and spectral analysis.
ParaDigMa is accompanied by comprehensive documentation including tutorials,
installation guides, and API reference. The modular architecture enables researchers
to easily extend the toolbox with custom algorithms and functionalities while
maintaining compatibility with standardized data formats.
keywords:
- parkinson's disease
- digital biomarkers
- gait analysis
- ppg signal
- imu data
license: Apache-2.0
Issues and Pull Requests
Last synced: 3 months ago
All Time
- Total issues: 46
- Total pull requests: 135
- Average time to close issues: 4 months
- Average time to close pull requests: 2 days
- Total issue authors: 5
- Total pull request authors: 6
- Average comments per issue: 0.04
- Average comments per pull request: 0.5
- Merged pull requests: 89
- Bot issues: 0
- Bot pull requests: 0
Past Year
- Issues: 28
- Pull requests: 35
- Average time to close issues: about 2 months
- Average time to close pull requests: 4 days
- Issue authors: 2
- Pull request authors: 5
- Average comments per issue: 0.0
- Average comments per pull request: 0.14
- Merged pull requests: 22
- Bot issues: 0
- Bot pull requests: 0
Top Authors
Issue Authors
- Erikpostt (32)
- vedran-kasalica (10)
- movieminer (2)
- twanvl (1)
- KarsVeldkamp (1)
Pull Request Authors
- Erikpostt (73)
- nienketimmermans (19)
- KarsVeldkamp (18)
- kretep (16)
- vedran-kasalica (8)
- lucevers1991 (1)
Top Labels
Issue Labels
Pull Request Labels
Packages
- Total packages: 1
-
Total downloads:
- pypi 323 last-month
- Total dependent packages: 0
- Total dependent repositories: 0
- Total versions: 20
- Total maintainers: 5
pypi.org: paradigma
ParaDigMa - A toolbox for deriving Parkinson's disease Digital Markers from real-life wrist sensor data
- Documentation: https://paradigma.readthedocs.io/
- License: Apache-2.0
-
Latest release: 1.1.0
published 4 months ago