motu-profiler
motus - a tool for marker gene-based OTU (mOTU) profiling
Science Score: 67.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
✓CITATION.cff file
Found CITATION.cff file -
✓codemeta.json file
Found codemeta.json file -
✓.zenodo.json file
Found .zenodo.json file -
✓DOI references
Found 2 DOI reference(s) in README -
○Academic publication links
-
✓Committers with academic emails
6 of 12 committers (50.0%) from academic institutions -
○Institutional organization owner
-
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (13.8%) to scientific vocabulary
Repository
motus - a tool for marker gene-based OTU (mOTU) profiling
Basic Info
- Host: GitHub
- Owner: motu-tool
- License: gpl-3.0
- Language: Python
- Default Branch: master
- Size: 1.66 MB
Statistics
- Stars: 156
- Watchers: 10
- Forks: 30
- Open Issues: 5
- Releases: 1
Metadata Files
README.md
mOTU profiler
The mOTU profiler is a computational tool that estimates relative taxonomic abundance of known and currently unknown microbial community members using metagenomic shotgun sequencing data.
Check the wiki for more information.
If you use mOTUs, please cite:
Reference genome-independent taxonomic profiling of microbiomes with mOTUs3
Hans-Joachim Ruscheweyh, Alessio Milanese, Lucas Paoli, Nicolai Karcher, Quentin Clayssen, Marisa Isabell Metzger, Jakob Wirbel, Peer Bork, Daniel R. Mende, Georg Zeller# & Shinichi Sunagawa#
Microbiome (2022)
Pre-requisites
The mOTU profiler requires: * Python 3 (or higher) * the Burrow-Wheeler Aligner v0.7.15 or higher (bwa) * SAMtools v1.5 or higher (link)
In order to use the command snv_call you need:
* metaSNV v1.0.3, available also on bioconda (we assume metaSNV.py to be in the system path)
Check installation wiki to see how to install the dependencies with conda.
Installation
mOTUs can be installed either by using pip or via conda.
Installation with conda has the advantage that it will also download and install dependencies:
```bash
Install in the base environment
conda install motus
OR, create a new environment
conda create -n motu-env motus conda activate motu-env ```
Installation with pip:
```bash
Download and install mOTUs
pip install motu-profiler
Download the mOTUs database
motus downloadDB ```
You can test that motus is intalled correctly with:
motus profile --test
Basic examples
Here is a simple example on how to obtain a taxonomic profiling from a raw read file:
bash
motus profile -s metagenomic_sample.fastq > taxonomy_profile.txt
You can separate the previous call as:
bash
motus map_tax -s metagenomic_sample.fastq -o mapped_reads.sam
motus calc_mgc -i mapped_reads.sam -o mgc_ab_table.count
motus calc_motu -i mgc_ab_table.count > taxonomy_profile.txt
rm mapped_reads.sam mgc_ab_table.count
The use of multiple threads (-t) is recommended, since bwa will finish faster. Here is an example with Paired-End reads:
bash
motus profile -f for_sample.fastq -r rev_sample.fastq -s no_pair.fastq -t 6 > taxonomy_profile.txt
You can merge taxonomy files from different samples with mOTU merge:
shell
motus profile -s metagenomic_sample_1.fastq -o taxonomy_profile_1.txt
motus profile -s metagenomic_sample_2.fastq -o taxonomy_profile_2.txt
motus merge -i taxonomy_profile_1.txt,taxonomy_profile_2.txt > all_sample_profiles.txt
You can profile samples that have been sequenced through different runs:
shell
motus profile -f sample1_run1_for.fastq,sample1_run2_for.fastq -r sample1_run1_rev.fastq,sample1_run2_rev.fastq -s sample1_run1_single.fastq > taxonomy_profile.txt
How mOTUs works
The mOTUs tool performs taxonomic profiling of metagenomics and metatrancriptomics samples, i.e. it identifies species and their relative abundance present in a sample. It is based on a set of mOTUs (~species) contained in the mOTUs database. The mOTUs database is created from reference genomes, metagenomic samples and metagenome assembled genomes (MAGs):
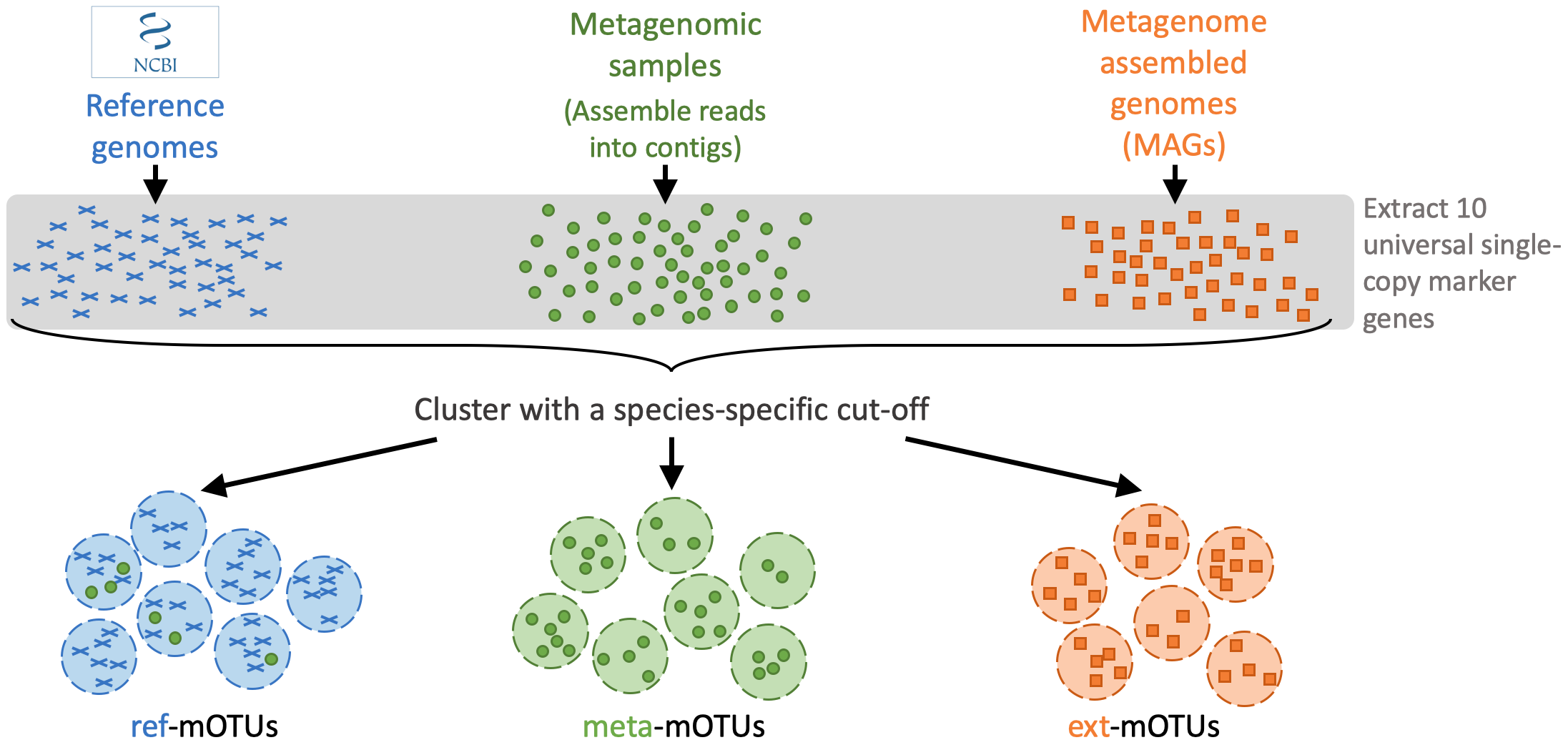
A mOTUs database is composed of three types of mOTUs: - ref-mOTUs, which represent known species, - meta-mOTUs, which represent unknown species obtained from metagenomic samples, - ext-mOTUs, which represent unknown species obtained from MAGs.
Note that meta- and ext-mOTUs will not have a species level annotation.
The mOTUs database is updated periodically, e.g the latest version (3.0.3), which doubles the number of profilable species by including ~600,000 draft genomes. Major releases are represented in the following graph (where the numbers represents the number of mOTUs for each of the three groups, with the same color-code as the previous graph):
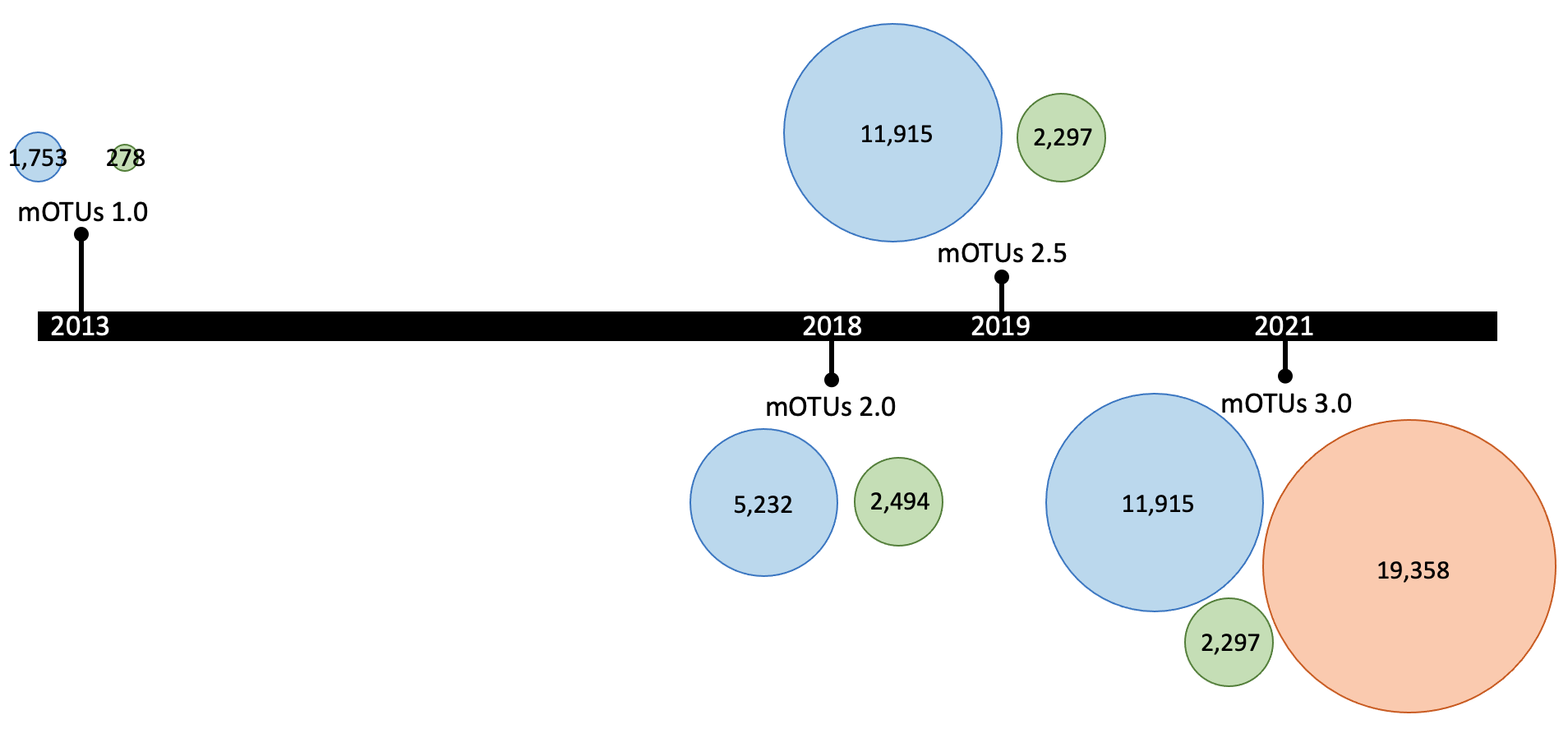
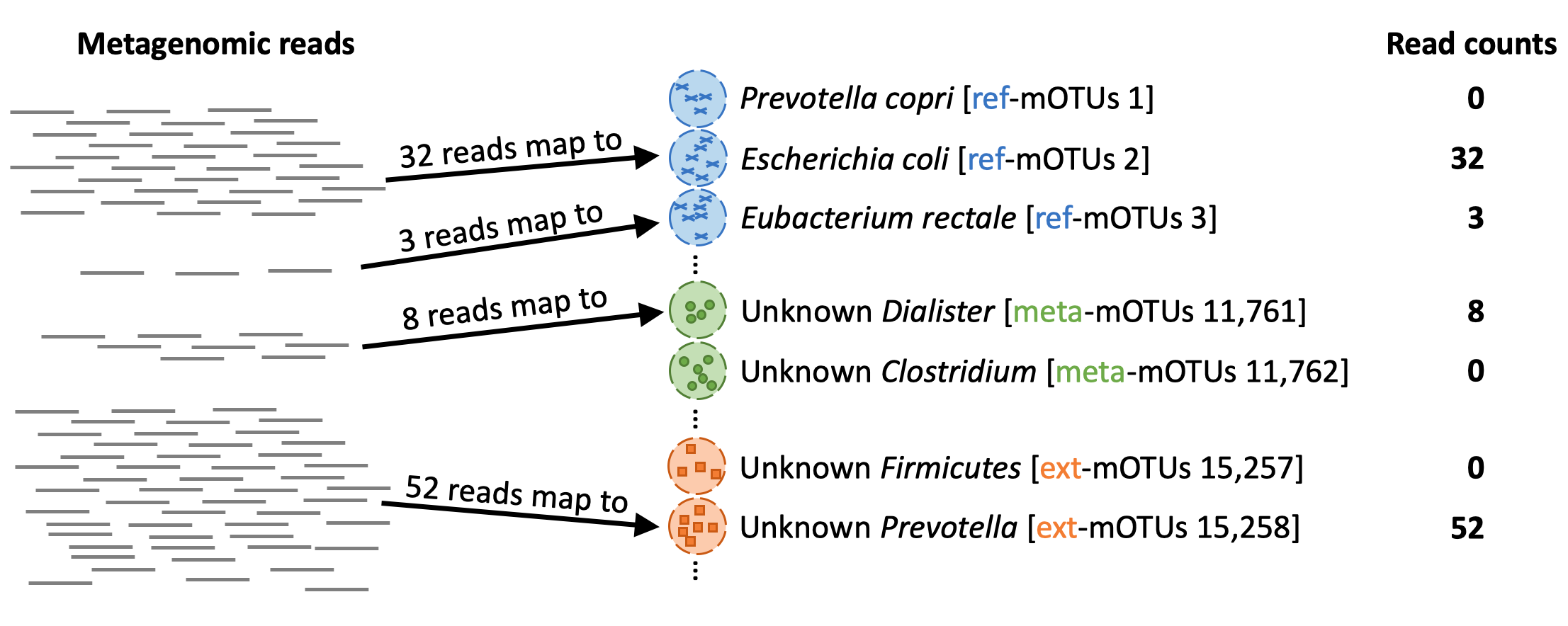
When profiling (motus profile) a metagenomic sample, the mOTUs tool maps the reads from the sample to the genes in the different mOTUs:
ChangeLog
Version 3.1.0 2023-03-28 by AlessioMilanese * Improve database clustering algorithm and update the database (change the number of ext-mOTUs from 19,358 to 20,128)
Version 3.0.3 2022-07-13 by AlessioMilanese
* Add command prep_long to allow the profiling of long reads (more information here)
Version 3.0.2 2022-01-31 by AlessioMilanese * Convert the repository to a python package and submit to PyPI
Version 3.0.1 2021-07-27 by AlessioMilanese
* Improve ref-mOTUs taxonomy according to #76
* Solve bug with -A option
Version 3.0.0 2021-06-22 by AlessioMilanese * Improve code base * Minor bug fixes
Version 2.6.1 2021-04-27 by AlessioMilanese * Minor bug fixes * Improved the taxonomy of 32 ref-mOTUs (#45)
Version 2.6.0 2021-03-08 by AlessioMilanese
* Add 19,358 new mOTUs
* Add taxonomic profiles of > 11k metagenomic and metatranscriptomic samples. The updated merge function can integrate those in to the users results.
* Minor bug fixes
* Change -1 to unassigned
Version 2.5.1 2019-08-17 by AlessioMilanese * Update the taxonomy to participate to the CAMI 2 challenge
Version 2.5.0 2019-08-09 by AlessioMilanese * Add -db option to use a database from another directory * Add -A to print all taxonomy levels together * Update the database with more than 60k new reference genomes. There are 11,915 ref-mOTUs and 2,297 meta-mOTUs.
Version 2.1.1 2019-03-04 by AlessioMilanese * Correct problem with samtools when installing with conda
Version 2.1.0 2019-03-03 by AlessioMilanese * Correct error \'\t\t\' when printing -C recall * Update database (gene coordinates)
Version 2.0.1 2018-08-23 by AlessioMilanese * Add -C to print the result in CAMI format (BioBoxes format 0.9.1) * Add -K to snv_call command to keep all the directories produced by metaSNV
Version 2.0.0 2018-06-12 by AlessioMilanese * Set relative abundances as default (instead of counts) * Add -B to print the result in BIOM format * Add test directory * Python2 is not supported anymore * Minor bug fixes
Version 2.0.0-rc1 2018-05-10 by AlessioMilanese * First release supporting all basic functionality.
Owner
- Name: motu-tool
- Login: motu-tool
- Kind: organization
- Repositories: 7
- Profile: https://github.com/motu-tool
Citation (CITATION.cff)
cff-version: 1.2.0
message: If you use this software, please cite it as below.
title: 'Microbial abundance, activity and population genomic profiling with mOTUs2'
doi: 10.1038/s41467-019-08844-4
authors:
- given-names: Alessio
family-names: Milanese
affiliation: European Molecular Biology Laboratory, Heidelberg, Germany
orcid: https://orcid.org/0000-0002-7050-2239
- given-names: Daniel R.
family-names: Mende
affiliation: Daniel K. Inouye Center for Microbial Oceanography Research and Education, University of Hawaii at Mānoa, Honolulu, United States
orcid: https://orcid.org/0000-0001-6831-4557
- given-names: Lucas
family-names: Paoli
affiliation: Department of Biology, Institute of Microbiology and Swiss Institute of Bioinformatics, ETH Zürich
& Department of Biology, École normale supérieure, Paris, France
orcid: https://orcid.org/0000-0003-0771-8309
- given-names: Guillem
family-names: Salazar
affiliation: Department of Biology, Institute of Microbiology and Swiss Institute of Bioinformatics, ETH Zürich
orcid: https://orcid.org/0000-0002-9786-1493
- given-names: Miguelangel
family-names: Cuenca
affiliation: Department of Biology, Institute of Microbiology and Swiss Institute of Bioinformatics, ETH Zürich
orcid: https://orcid.org/0000-0003-3435-9102
- given-names: Pascal
family-names: Hingamp
affiliation: Aix Marseille Univ, Université de Toulon, Marseille, France
- given-names: Renato
family-names: Alves
affiliation: European Molecular Biology Laboratory, Heidelberg, Germany
orcid: https://orcid.org/0000-0002-7212-0234
- given-names: Paul I.
family-names: Costea
affiliation: European Molecular Biology Laboratory, Heidelberg, Germany
orcid: https://orcid.org/0000-0003-1645-3947
- given-names: Luis Pedro
family-names: Coelho
affiliation: European Molecular Biology Laboratory, Heidelberg, Germany
orcid: https://orcid.org/0000-0002-9280-7885
- given-names: Thomas S. B.
family-names: Schmidt
affiliation: European Molecular Biology Laboratory, Heidelberg, Germany
orcid: https://orcid.org/0000-0001-8587-4177
- given-names: Alexandre
family-names: Almeida
affiliation: European Molecular Biology Laboratory,
European Bioinformatics Institute (EMBL-EBI), Wellcome Genome Campus, Hinxton, UK
& Wellcome Trust Sanger Institute, Wellcome Genome Campus, Hinxton, UK
orcid: https://orcid.org/0000-0001-8803-0893
- given-names: Alex L
family-names: Mitchell
affiliation: European Molecular Biology Laboratory,
European Bioinformatics Institute (EMBL-EBI), Wellcome Genome Campus, Hinxton, UK
- given-names: Robert D.
family-names: Finn
affiliation: European Molecular Biology Laboratory,
European Bioinformatics Institute (EMBL-EBI), Wellcome Genome Campus, Hinxton, UK
orcid: https://orcid.org/0000-0001-8626-2148
- given-names: Jaime
family-names: Huerta-Cepas
affiliation: European Molecular Biology Laboratory, Heidelberg, Germany
& Centro de Biotecnología y Genómica de Plantas,
Universidad Politécnica de Madrid (UPM) - Instituto Nacional de Investigación y Tecnología Agraria y Alimentaria (INIA), Madrid, Spain
orcid: https://orcid.org/0000-0003-4195-5025
- given-names: Peer
family-names: Bork
affiliation: European Molecular Biology Laboratory, Heidelberg, Germany
& Max Delbrück Centre for Molecular Medicine, Berlin, Germany
& Molecular Medicine Partnership Unit, Heidelberg, Germany
& Department of Bioinformatics, Biocenter, University of Würzburg, Würzburg, Germany
orcid: https://orcid.org/0000-0002-2627-833X
- given-names: Georg
family-names: Zeller
affiliation: European Molecular Biology Laboratory, Heidelberg, Germany
orcid: https://orcid.org/0000-0003-1429-7485
- given-names: Sunagawa
family-names: Shinichi
affiliation: Department of Biology, Institute of Microbiology and Swiss Institute of Bioinformatics, ETH Zürich
orcid: https://orcid.org/0000-0003-3065-0314
version: 3.0.3
date-released: 2022-07-13
repository-code: https://github.com/motu-tool/mOTUs
license: GNU General Public License v3.0
keywords:
- "Metagenomics"
- "Microbiome"
- "Software"
preferred-citation:
type: article
authors:
- given-names: Alessio
family-names: Milanese
affiliation: European Molecular Biology Laboratory, Heidelberg, Germany
orcid: https://orcid.org/0000-0002-7050-2239
- given-names: Daniel R.
family-names: Mende
affiliation: Daniel K. Inouye Center for Microbial Oceanography Research and Education, University of Hawaii at Mānoa, Honolulu, United States
orcid: https://orcid.org/0000-0001-6831-4557
- given-names: Lucas
family-names: Paoli
affiliation: Department of Biology, Institute of Microbiology and Swiss Institute of Bioinformatics, ETH Zürich
& Department of Biology, École normale supérieure, Paris, France
orcid: https://orcid.org/0000-0003-0771-8309
- given-names: Guillem
family-names: Salazar
affiliation: Department of Biology, Institute of Microbiology and Swiss Institute of Bioinformatics, ETH Zürich
orcid: https://orcid.org/0000-0002-9786-1493
- given-names: Miguelangel
family-names: Cuenca
affiliation: Department of Biology, Institute of Microbiology and Swiss Institute of Bioinformatics, ETH Zürich
orcid: https://orcid.org/0000-0003-3435-9102
- given-names: Pascal
family-names: Hingamp
affiliation: Aix Marseille Univ, Université de Toulon, Marseille, France
- given-names: Renato
family-names: Alves
affiliation: European Molecular Biology Laboratory, Heidelberg, Germany
orcid: https://orcid.org/0000-0002-7212-0234
- given-names: Paul I.
family-names: Costea
affiliation: European Molecular Biology Laboratory, Heidelberg, Germany
orcid: https://orcid.org/0000-0003-1645-3947
- given-names: Luis Pedro
family-names: Coelho
affiliation: European Molecular Biology Laboratory, Heidelberg, Germany
orcid: https://orcid.org/0000-0002-9280-7885
- given-names: Thomas S. B.
family-names: Schmidt
affiliation: European Molecular Biology Laboratory, Heidelberg, Germany
orcid: https://orcid.org/0000-0001-8587-4177
- given-names: Alexandre
family-names: Almeida
affiliation: European Molecular Biology Laboratory,
European Bioinformatics Institute (EMBL-EBI), Wellcome Genome Campus, Hinxton, UK
& Wellcome Trust Sanger Institute, Wellcome Genome Campus, Hinxton, UK
orcid: https://orcid.org/0000-0001-8803-0893
- given-names: Alex L
family-names: Mitchell
affiliation: European Molecular Biology Laboratory,
European Bioinformatics Institute (EMBL-EBI), Wellcome Genome Campus, Hinxton, UK
- given-names: Robert D.
family-names: Finn
affiliation: European Molecular Biology Laboratory,
European Bioinformatics Institute (EMBL-EBI), Wellcome Genome Campus, Hinxton, UK
orcid: https://orcid.org/0000-0001-8626-2148
- given-names: Jaime
family-names: Huerta-Cepas
affiliation: European Molecular Biology Laboratory, Heidelberg, Germany
& Centro de Biotecnología y Genómica de Plantas,
Universidad Politécnica de Madrid (UPM) - Instituto Nacional de Investigación y Tecnología Agraria y Alimentaria (INIA), Madrid, Spain
orcid: https://orcid.org/0000-0003-4195-5025
- given-names: Peer
family-names: Bork
affiliation: European Molecular Biology Laboratory, Heidelberg, Germany
& Max Delbrück Centre for Molecular Medicine, Berlin, Germany
& Molecular Medicine Partnership Unit, Heidelberg, Germany
& Department of Bioinformatics, Biocenter, University of Würzburg, Würzburg, Germany
orcid: https://orcid.org/0000-0002-2627-833X
- given-names: Georg
family-names: Zeller
affiliation: European Molecular Biology Laboratory, Heidelberg, Germany
orcid: https://orcid.org/0000-0003-1429-7485
- given-names: Sunagawa
family-names: Shinichi
affiliation: Department of Biology, Institute of Microbiology and Swiss Institute of Bioinformatics, ETH Zürich
orcid: https://orcid.org/0000-0003-3065-0314
doi: "10.1038/s41467-019-08844-4"
journal: "Nature Communications"
month: 3
year: 2019
title: "Microbial abundance, activity and population genomic profiling with mOTUs2"
abstract: 'Metagenomic sequencing has greatly improved our ability to profile the composition
of environmental and host-associated microbial communities. However, the dependency of most methods
on reference genomes, which are currently unavailable for a substantial fraction of microbial species,
introduces estimation biases. We present an updated and functionally extended tool
based on universal (i.e., reference-independent), phylogenetic marker gene (MG)-based
operational taxonomic units (mOTUs) enabling the profiling of >7700 microbial species.
As more than 30% of them could not previously be quantified at this taxonomic resolution,
relative abundance estimates based on mOTUs are more accurate compared to other methods.
As a new feature, we show that mOTUs, which are based on essential housekeeping genes,
are demonstrably well-suited for quantification of basal transcriptional activity of community members.
Furthermore, single nucleotide variation profiles estimated using mOTUs reflect those from whole genomes,
which allows for comparing microbial strain populations (e.g., across different human body sites).'
GitHub Events
Total
- Issues event: 26
- Watch event: 10
- Issue comment event: 28
- Pull request event: 1
- Fork event: 6
Last Year
- Issues event: 26
- Watch event: 10
- Issue comment event: 28
- Pull request event: 1
- Fork event: 6
Committers
Last synced: over 1 year ago
Top Committers
| Name | Commits | |
|---|---|---|
| Alessio Milanese | m****o@g****m | 278 |
| Renato Alves | a****c@g****m | 18 |
| Hans-Joachim Ruscheweyh | h****r@e****h | 11 |
| Lucas Paoli | l****i@g****m | 8 |
| Alessio Milanese | a****e@e****e | 4 |
| AlessioMilanese | a****m@K****h | 3 |
| SuShiAtGit | 3****t | 2 |
| AlessioMilanese | a****m@m****h | 1 |
| Hans-Joachim Ruscheweyh (ID SIS) | h****r@b****h | 1 |
| Hans-Joachim Ruscheweyh | h****r@p****h | 1 |
| Florian Plaza Oñate | f****a | 1 |
| Valentyn Bezshapkin | 6****z | 1 |
Committer Domains (Top 20 + Academic)
Issues and Pull Requests
Last synced: 6 months ago
All Time
- Total issues: 110
- Total pull requests: 14
- Average time to close issues: 7 months
- Average time to close pull requests: about 7 hours
- Total issue authors: 82
- Total pull request authors: 6
- Average comments per issue: 3.75
- Average comments per pull request: 0.43
- Merged pull requests: 11
- Bot issues: 0
- Bot pull requests: 0
Past Year
- Issues: 11
- Pull requests: 1
- Average time to close issues: 3 months
- Average time to close pull requests: N/A
- Issue authors: 11
- Pull request authors: 1
- Average comments per issue: 0.73
- Average comments per pull request: 0.0
- Merged pull requests: 0
- Bot issues: 0
- Bot pull requests: 0
Top Authors
Issue Authors
- Jigyasa3 (7)
- mikemc (6)
- AlessioMilanese (5)
- valentynbez (3)
- zckoo007 (3)
- Jibowe (2)
- Anto007 (2)
- sjaenick (2)
- fplaza (2)
- jzrapp (2)
- sturne29 (2)
- hjruscheweyh (2)
- unode (2)
- handibles (1)
- trickovicmatija (1)
Pull Request Authors
- AlessioMilanese (8)
- unode (2)
- matthpich (1)
- lijier6 (1)
- fplaza (1)
- lijierr (1)
- valentynbez (1)
Top Labels
Issue Labels
Pull Request Labels
Packages
- Total packages: 1
-
Total downloads:
- pypi 37 last-month
- Total dependent packages: 0
- Total dependent repositories: 3
- Total versions: 3
- Total maintainers: 1
pypi.org: motu-profiler
Taxonomic profiling of metagenomes from diverse environments with mOTUs3
- Homepage: https://github.com/motu-tool/mOTUs
- Documentation: https://motu-profiler.readthedocs.io/
- License: GPLv3
-
Latest release: 3.1.0
published almost 3 years ago

