GitHub

https://github.com/bammri/3d_dynamic_paper_2024
Code and data for the 3D Dynamic paper 2024


https://github.com/bamresearch/probeye
A general framework for setting up parameter estimation problems.

https://github.com/bandframework/rose
The Reduced-Order Scattering Emulator (rose) is a user-friendly software for building efficient surrogate models for nuclear scattering

https://github.com/barahona-research-group/multiscalemobilitypatterns
Code for the paper "Multiscale mobility patterns and the restriction of human movement" by Juni Schindler, Jonathan M Clarke and Mauricio Barahona: https://arxiv.org/abs/2201.06323

https://github.com/barathme/hen_multiscale_lowk
Explainable Multiscale Modeling of High-Entropy Nitride Superlattices for Low-Thermal-Conductivity Coatings

https://github.com/barricklab/ltee-ecoli
Genomics resources for the Long-Term Evolution Experiment with E. coli

https://github.com/bartongroup/sack1g-analysis
This is a GitHub Repository containing the data and code used for an structural analysis of FAM83G and FAM83G-CK1A 3D models

https://github.com/batmen-lab/diamond
Error-controlled interaction discovery in machine learning models


https://github.com/battmodels/diffthermo_ocv_paper
diffthermo, a python package for thermodynamically consistent OCV model construction

https://github.com/battmoteam/pybamm
Fast and flexible physics-based battery models in Python

https://github.com/bbglab/oncodrive3d
A computational approach for identifying cancer driver genes by detecting three-dimensional clusters of somatic missense mutations in protein structures.

https://github.com/bbopt/rungekutta
The RUNGE-KUTTA blackbox optimization problem

https://github.com/beatlab-mcmaster/schlichting_msc_thesis
Code and data to reproduce my master's thesis.


https://github.com/bede/deacon
FASTX search and [host] depletion using minimizers

https://github.com/beefathima-git/cloud-computing-for-real-time-surveillance-applications-
This project utilizes fog and cloud computing to enable low-latency, real-time surveillance by processing video and sensor data closer to the source. It improves responsiveness, reduces bandwidth usage, and enhances system scalability for smart surveillance applications.

https://github.com/beehive-lab/tornadovm
TornadoVM: A practical and efficient heterogeneous programming framework for managed languages



https://github.com/benjamindecker/quantum-cellular-automaton
A classical simulation of quantum cellular automata

https://github.com/benjaminhlina/nichetools
Complementary functions to extract and work with objects created by {nicheROVER} and {SIBER}

https://github.com/benjaminhlina/lake-ontario-ecoregions-ms
Data and code for paper centred on trophic differences among ecoregions in Lake Ontario

https://github.com/benweare/em_scripts
Electron microscopy scripts not associated with any specific publications.

https://github.com/berenslab/disentangling-retinal-images
This repository contains the code for the paper "Disentangling representations of retinal images with generative models".


https://github.com/bhklab/signaturesets
Compendium of published molecular signatures


https://github.com/bihealth/fracture-healing-and-aging-scseq
single-cell bone healing project

https://github.com/billbrod/spatial-frequency-model
Model from the spatial frequency preferences paper

https://github.com/billbrod/foveated-metamers
Create metamers using models of the ventral stream and run experiments to validate them

https://github.com/bin-cao/tcgpr
[NPJ Com Mat 2023 | Small 2024] Machine Learning Algorithm : outlier identifying, feature selection






https://github.com/biodt/general-soilgrids-soil-data
Building block for obtaining selected soil data at given location from SoilGrids and derived data sources (Soilgrids REST API, HiHydroSoil map).

https://github.com/biodt/uc-grassland-model
grassland simulation model source code

https://github.com/bioinfomachinelearning/gate
Graph transformer for estimating protein model accuracy


https://github.com/biop/qupath-extension-cellpose
an extension that wraps a Cellpose environment such that WSI can be analyzed using Cellpose through QuPath.


https://github.com/bioscan-ml/bioscan-1m
A Step Towards Worldwide Biodiversity Assessment: The BIOSCAN-1M Insect Dataset

https://github.com/biswesh456/bargaining
This repo contains code for the paper Incorporating Autonomous Bargaining Capabilities into E-Commerce Systems which was accepted in ACM IVA 2020.

https://github.com/bitcs-information-retrieval-2021-2022/assignment-description
Description of Final Assignments of Information Retrieval Course 2021-2022


https://github.com/bluebrain/topological_sampling
The analysis pipeline for the topological sampling project


https://github.com/bluebrain/neurodamus
A BBP Simulation Control application for NEURON

https://github.com/bluebrain/psp-validation
Perform spontaneous minis validations

https://github.com/bluebrain/currentscape
Currentscape is a Python tool enabling scientists to easily plot the currents in electrical neuron models. The code is based on the paper Alonso and Marder, 2019.


https://github.com/bluebrain/astrovascpy
Vasculature blood flow computation and impact of astrocytic endfeet on vessels

https://github.com/bluebrain/atlas-enhancement
Tools to generate flatmaps of brain atlases & code to add barrel and barrel column annotations to Allen Mouse CCFv3

https://github.com/bluebrain/vessmorphovis
A lightweight, interactive, extensible and cross-platform framework for building, visualizing and analyzing vasculature (or blood vessels) morphologies.

https://github.com/bluegreen-labs/swift_cc894
Analysis of the 19 month flight of juvenile swift CC894

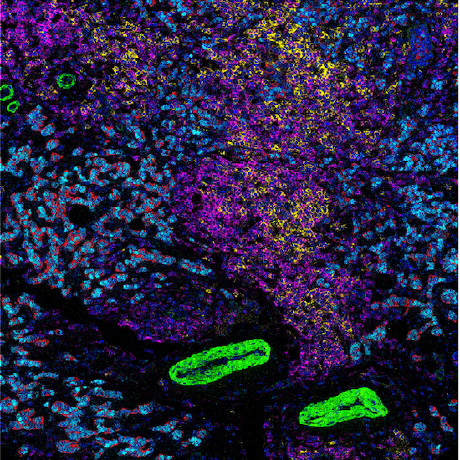
https://github.com/bodenmillergroup/scpathology_publication
The Single-Cell Pathology Landscape of Breast Cancer

https://github.com/boniolp/tsadtaxonomy
This application helps users explore and understand the vast array of existing methods, ranging from traditional statistical approaches to modern machine learning algorithms. It visualizes a structured, process-centric taxonomy of anomaly detection techniques, enabling a deeper insight into the research landscape.

https://github.com/borchlab/ibex
Using BCR and expression for sequence embedding



https://github.com/brain-development-and-disorders-lab/mars
Metadatify is an open-source web-based tool to create, manage, and search scientific metadata.

https://github.com/braingeneers/sims-web
Attempt to run SIMS inference in the browser

https://github.com/bramvanroy/mantis
Segmentation interface for the TPR-DB to manually tokenize and sentence segment

https://github.com/bridgedb/wikidata2bridgedb
Creation of BridgeDb mapping databases from Wikidata


https://github.com/britishgeologicalsurvey/ash-model-plotting
Wrapper around Iris Python library for easy plotting of NAME, FALL3D and HYSPLIT model results.

https://github.com/broadinstitute/one-shot-atlas
Siamese Neural Network for anatomical registration

https://github.com/broadinstitute/str-truth-set
Genome-wide tandem repeat (TR) truth set based on the Synthetic Diploid Benchmark [Li et al. 2018]


https://github.com/broadinstitute/sma-finder
A tool for diagnosing SMA using exome, genome or targeted sequencing data


https://github.com/broadinstitute/g2papi
Python Client Library for the G2P Portal API

https://github.com/brucewlee/conference_cheatsheet
publication-related stuff, largely for myself

https://github.com/bryanparthum/usrw_waterquality
Code and data for "Overlooked Benefits of Nutrient Reductions in the Mississippi River Basin"



https://github.com/bueler/mcd-extended
extending the multilevel constraint decomposition method of Tai (2003)

https://github.com/bugratufan/make-silicon
A streamlined toolkit for initializing FPGA and ASIC directories and scripts.




https://github.com/bytedance/pxdesignbench
A Unified Evaluation Suite for Protein Design

https://github.com/bytedance/hammer
An efficient toolkit for training deep models.

https://github.com/byuflowlab/implicitad.jl
Automates steady and unsteady adjoints (general solvers and ODEs respectively). Forward and reverse mode algorithmic differentiation around implicit functions (not propagating AD through), as well as custom rules to allow for mixed-mode AD or calling external (non-AD compatible) functions within an AD chain.



https://github.com/cadet/rdm-example-rectangular-pulse
CADET case study from paper: Analytical solutions and moment analysis of general rate model for linear liquid chromatography” (Shamsul Qamar et al.)

https://github.com/cadet/rdm-example-multi-state-steric-mass-action
CADET case study from paper: Multi-state steric mass action model and case study on complex high loading behavior of mAb on ion exchange tentacle resin (Diedrich et al.)

https://github.com/cafferychen777/awesome-scrna-annotation
A curated list of tools and methods for scRNA-seq cell type annotation

https://github.com/calliope-project/module_road_transport
A module to compute road transport timeseries and yearly values.

https://github.com/caltechlibrary/2018-03-29-r-workshop
Repo for files for 2018-03-29 Data Analysis, Visualization, and Reproducibility with R Notebooks workshop.

https://github.com/cambiotraining/stats-mixed-effects-models

https://github.com/camilogarciabotero/biosimplex.jl
Representing BioSequences as Simplex numerical matrix

https://github.com/captaincodercool/captaincodercool
Welcome to CAPTAINCODERCOOL’s GitHub! Dive into my repository of diverse projects ranging from AI-driven tools to full-stack web applications. Discover code, insights, and innovations crafted to push the boundaries of technology and software development.

https://github.com/carmonalab/genenmf
Methods to discover gene programs on single-cell data

https://github.com/carmonalab/stacas
R package for semi-supervised single-cell data integration