https://github.com/csoneson/alevinqc
Create QC and summary reports for Alevin output
Science Score: 49.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
○CITATION.cff file
-
✓codemeta.json file
Found codemeta.json file -
✓.zenodo.json file
Found .zenodo.json file -
✓DOI references
Found 1 DOI reference(s) in README -
○Academic publication links
-
✓Committers with academic emails
2 of 5 committers (40.0%) from academic institutions -
○Institutional organization owner
-
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (9.1%) to scientific vocabulary
Keywords from Contributors
Repository
Create QC and summary reports for Alevin output
Basic Info
- Host: GitHub
- Owner: csoneson
- License: other
- Language: R
- Default Branch: devel
- Homepage: https://csoneson.github.io/alevinQC/
- Size: 14.4 MB
Statistics
- Stars: 31
- Watchers: 4
- Forks: 6
- Open Issues: 2
- Releases: 0
Metadata Files
README.md
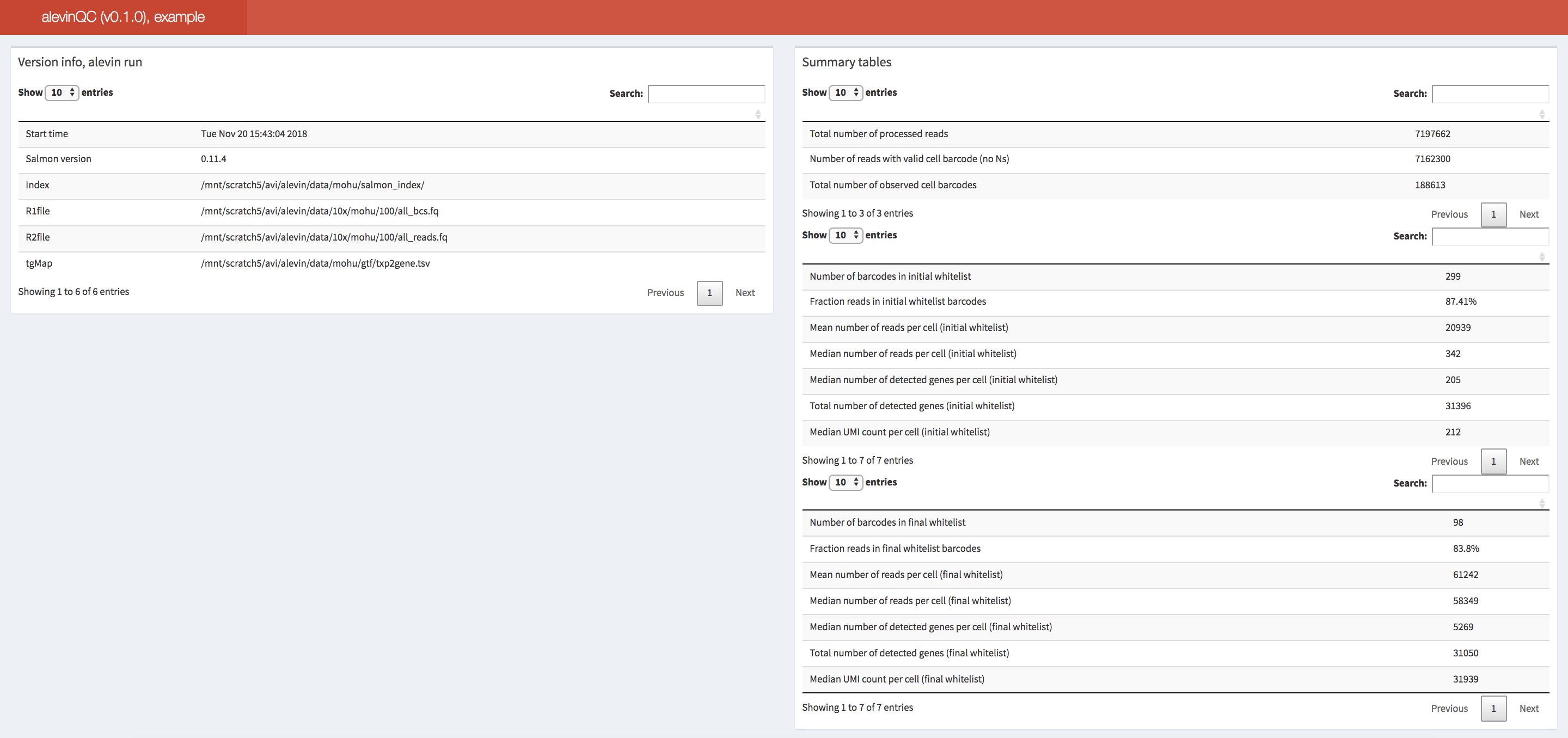
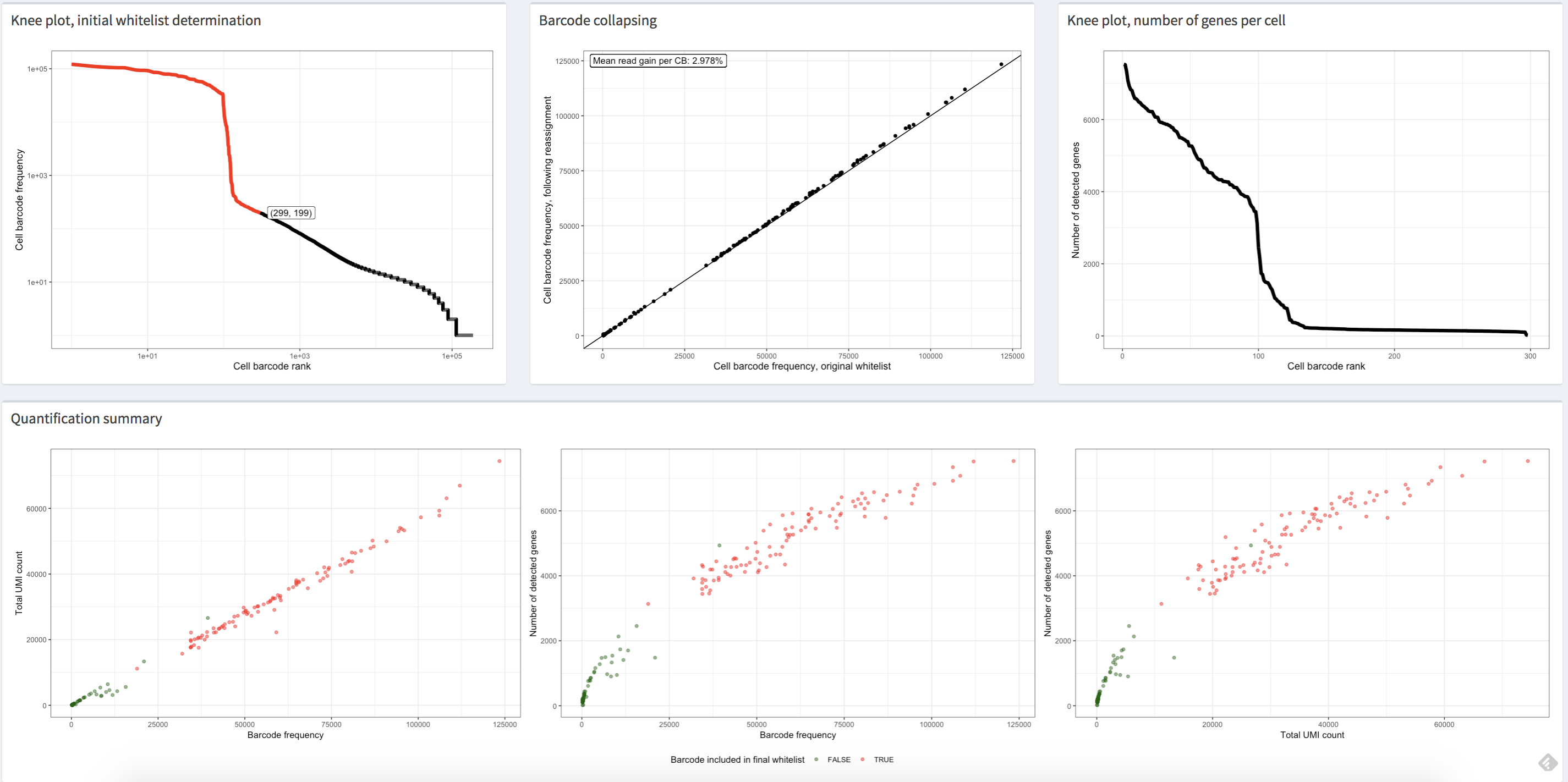
alevinQC
The alevinQC R package provides functionality for generating QC reports
summarizing the output of alevin
(Srivastava et al., Genome Biology 20:65 (2019)). The reports can
be generated in html or pdf format, or as R/Shiny applications.
Installation:
alevinQC is available from
Bioconductor, and can be
installed using the BiocManager CRAN package:
if (!requireNamespace("BiocManager", quietly = TRUE))
install.packages("BiocManager")
BiocManager::install("alevinQC")
Note that alevinQC v 1.1 or newer is required to process output from Salmon version 0.14.0 or newer.
Example usage:
alevinQCReport(baseDir = system.file("extdata/alevin_example_v0.14",
package = "alevinQC"),
sampleId = "testSample",
outputFile = "alevinReport.html",
outputFormat = "html_document",
outputDir = tempdir(), forceOverwrite = TRUE)
For more information, we refer to the package vignette.
Owner
- Name: Charlotte Soneson
- Login: csoneson
- Kind: user
- Website: http://csoneson.github.io/
- Twitter: CSoneson
- Repositories: 110
- Profile: https://github.com/csoneson
GitHub Events
Total
- Issues event: 1
- Watch event: 2
- Issue comment event: 1
- Push event: 8
- Create event: 2
Last Year
- Issues event: 1
- Watch event: 2
- Issue comment event: 1
- Push event: 8
- Create event: 2
Committers
Last synced: 10 months ago
Top Committers
| Name | Commits | |
|---|---|---|
| Charlotte Soneson | c****n@g****m | 240 |
| Nitesh Turaga | n****a@g****m | 14 |
| J Wokaty | j****y@s****u | 10 |
| DongzeHe | d****a@g****m | 5 |
| A Wokaty | a****y@s****u | 2 |
Committer Domains (Top 20 + Academic)
Issues and Pull Requests
Last synced: 7 months ago
All Time
- Total issues: 21
- Total pull requests: 12
- Average time to close issues: 23 days
- Average time to close pull requests: about 7 hours
- Total issue authors: 20
- Total pull request authors: 3
- Average comments per issue: 3.95
- Average comments per pull request: 0.75
- Merged pull requests: 11
- Bot issues: 0
- Bot pull requests: 0
Past Year
- Issues: 1
- Pull requests: 0
- Average time to close issues: N/A
- Average time to close pull requests: N/A
- Issue authors: 1
- Pull request authors: 0
- Average comments per issue: 1.0
- Average comments per pull request: 0
- Merged pull requests: 0
- Bot issues: 0
- Bot pull requests: 0
Top Authors
Issue Authors
- crazyhottommy (2)
- LisaHagenau (1)
- PavanKhosla (1)
- wmacnair (1)
- apeltzer (1)
- PietroD (1)
- jaclyn-taroni (1)
- Ryan-Zhu (1)
- csoneson (1)
- jdrnevich (1)
- hiraksarkar (1)
- vswarup (1)
- gringer (1)
- yueli8 (1)
- changostraw (1)
Pull Request Authors
- csoneson (10)
- cansavvy (1)
- DongzeHE (1)
Top Labels
Issue Labels
Pull Request Labels
Packages
- Total packages: 1
-
Total downloads:
- bioconductor 15,507 total
- Total dependent packages: 0
- Total dependent repositories: 0
- Total versions: 6
- Total maintainers: 1
bioconductor.org: alevinQC
Generate QC Reports For Alevin Output
- Homepage: https://github.com/csoneson/alevinQC
- Documentation: https://bioconductor.org/packages/release/bioc/vignettes/alevinQC/inst/doc/alevinQC.pdf
- License: MIT + file LICENSE
-
Latest release: 1.24.0
published 11 months ago
Rankings
Maintainers (1)
Dependencies
- R >= 4.0 depends
- DT * imports
- GGally * imports
- Rcpp * imports
- cowplot * imports
- dplyr * imports
- ggplot2 * imports
- methods * imports
- rjson * imports
- rlang * imports
- rmarkdown >= 2.5 imports
- shiny * imports
- shinydashboard * imports
- stats * imports
- tools * imports
- tximport >= 1.17.4 imports
- utils * imports
- BiocManager * suggests
- BiocStyle * suggests
- knitr * suggests
- testthat >= 3.0.0 suggests
- actions/cache v1 composite
- actions/checkout v2 composite
- actions/upload-artifact master composite
- grimbough/bioc-actions/build-install-check v1 composite
- grimbough/bioc-actions/run-BiocCheck v1 composite
- grimbough/bioc-actions/setup-bioc v1 composite
- r-lib/actions/setup-pandoc v2 composite