dacapo
A framework for easy application of established machine learning techniques on large, multi-dimensional images.
Science Score: 67.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
✓CITATION.cff file
Found CITATION.cff file -
✓codemeta.json file
Found codemeta.json file -
✓.zenodo.json file
Found .zenodo.json file -
✓DOI references
Found 2 DOI reference(s) in README -
✓Academic publication links
Links to: pubmed.ncbi, ncbi.nlm.nih.gov -
○Academic email domains
-
○Institutional organization owner
-
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (13.4%) to scientific vocabulary
Keywords
Repository
A framework for easy application of established machine learning techniques on large, multi-dimensional images.
Basic Info
- Host: GitHub
- Owner: janelia-cellmap
- License: bsd-3-clause
- Language: Python
- Default Branch: main
- Homepage: https://janelia-cellmap.github.io/dacapo/
- Size: 54.7 MB
Statistics
- Stars: 60
- Watchers: 4
- Forks: 10
- Open Issues: 28
- Releases: 5
Topics
Metadata Files
README.md

DaCapo 

A framework for easy application of established machine learning techniques on large, multi-dimensional images.
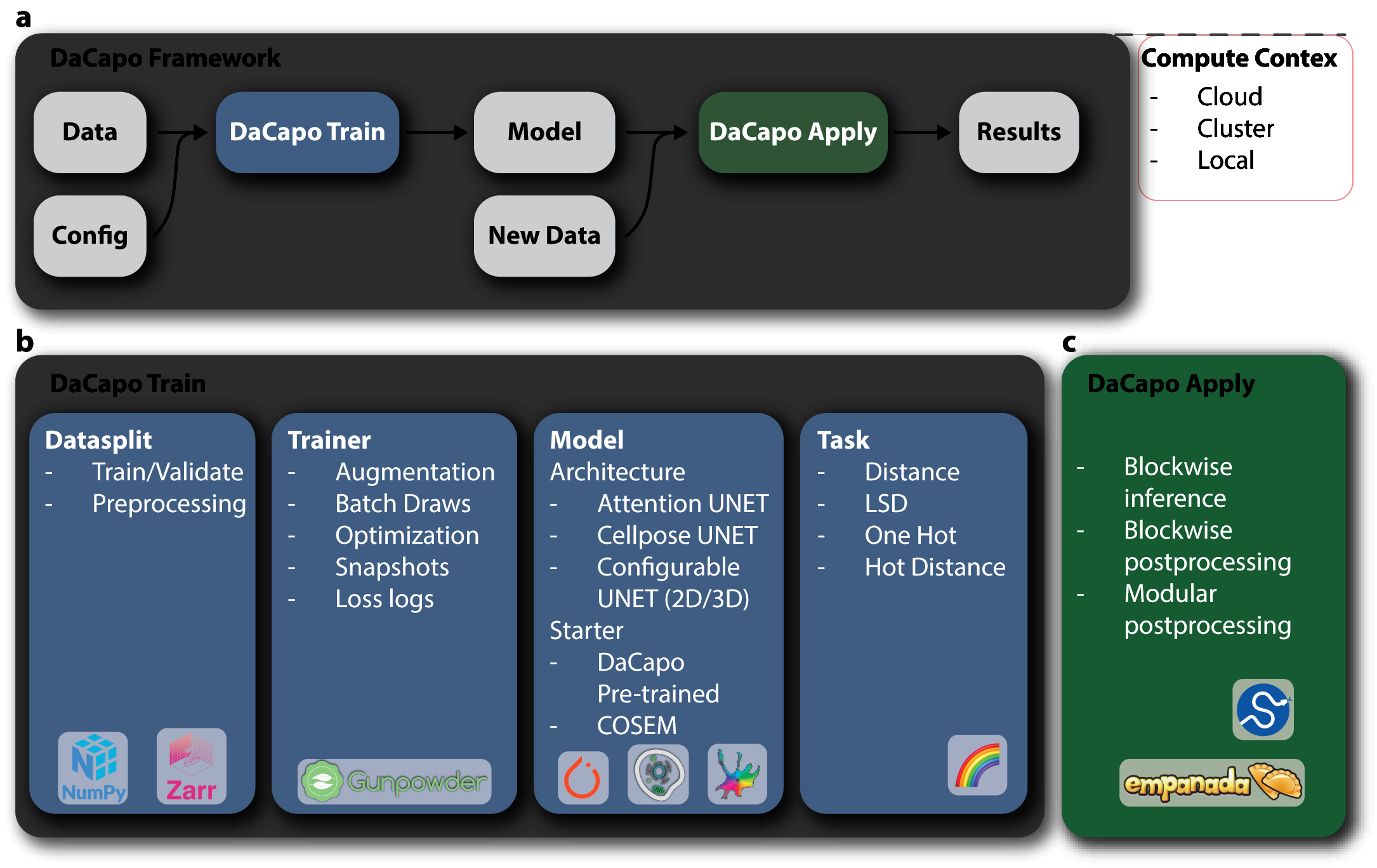
dacapo allows you to configure machine learning jobs as combinations of
DataSplits,
Architectures,
Tasks,
Trainers,
on arbitrarily large volumes of
multi-dimensional images. dacapo is not tied to a particular learning
framework, but currently only supports torch with
plans to support tensorflow.
Installation and Setup
Currently, python>=3.10 is supported. We recommend creating a new conda environment for dacapo with python 3.10.
conda create -n dacapo python=3.10
conda activate dacapo
Then install DaCapo using pip with the following command:
pip install dacapo-ml
This will install the minimum required dependencies.
You may additionally utilize a MongoDB server for storing outputs. To install and run MongoDB locally, refer to the MongoDB documentation here.
The use of MongoDB, as well as specifying the compute context (on cluster or not) should be specified in the dacapo.yaml in the main directory.
Functionality Overview
Tasks we support and approaches for those tasks: - Instance Segmentation - Affinities - Local Shape Descriptors - Semantic segmentation - Signed distances - One-hot encoding of different types of objects
Example Tutorial
A minimal example tutorial can be found in the examples directory and opened in colab here:
Helpful Resources & Tools
- Chunked data, zarr, and n5
- OME-Zarr: a cloud-optimized bioimaging file format with international community support (doi: 10.1101/2023.02.17.528834)
- Videos about N5 and Fiji can be found in this playlist. For other questions, join the discussion on the Image.sc forum.
- Read about chunked storage plugins in Fiji in this blog: N5 plugins for Fiji
- Script for converting tiff to zarr can be found here
- Segmentations
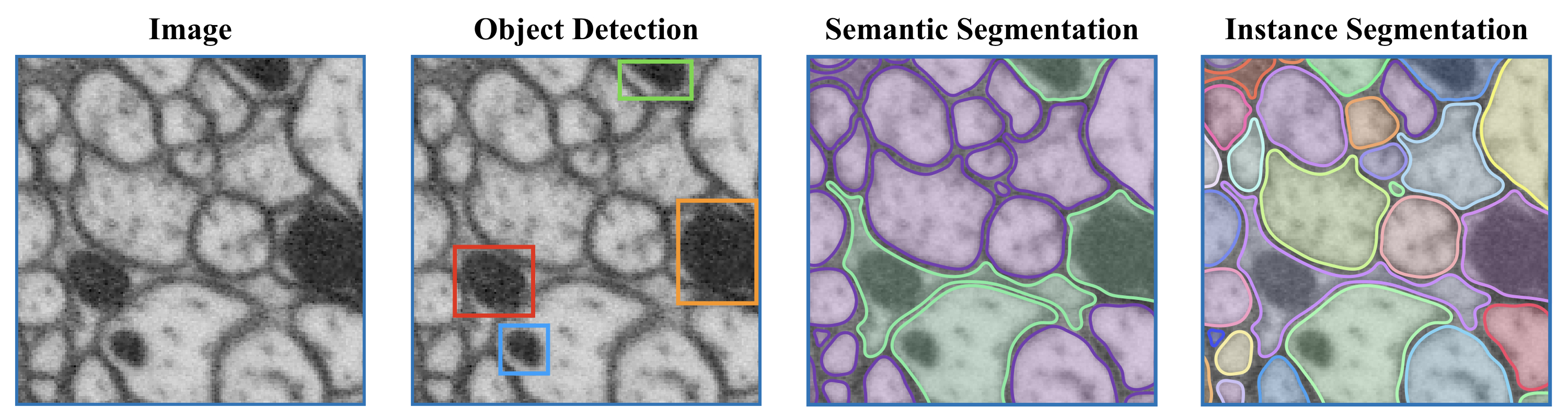
- A description of local shape descriptors used for affinities task. Read the blog here. Example image from the blog showing the difference between segmentations:

- CellMap Models
- GitHub Repo of published models
- For example, the COSEM trained pytorch networks are located here.
- OpenOrganelle.org
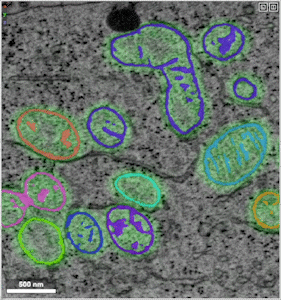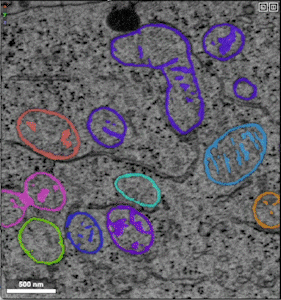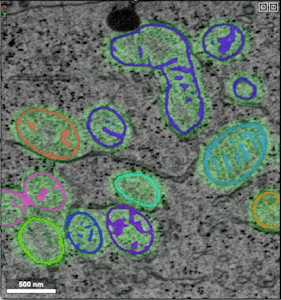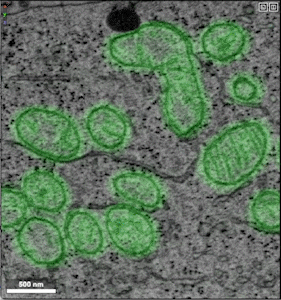
- Example of unprocessed distance predictions
- Example of refined segmentations that have undergone post-processing (e.g., thresholding, masking, smoothing)
- Example of groundtruth data
- Visualization
Citing this repo
If you use our code, please cite us and spread the news!
@article{Patton_DaCapo_a_modular_2024,
author = {Patton, William and Rhoades, Jeff L. and Zouinkhi, Marwan and Ackerman, David G. and Malin-Mayor, Caroline and Adjavon, Diane and Heinrich, Larissa and Bennett, Davis and Zubov, Yurii and Project Team, CellMap and Weigel, Aubrey V. and Funke, Jan},
doi = {10.48550/arXiv.2408.02834},
journal = {arXiv-cs.CV},
title = {{DaCapo: a modular deep learning framework for scalable 3D image segmentation}},
year = {2024}
}
Owner
- Name: CellMap Project Team
- Login: janelia-cellmap
- Kind: organization
- Location: United States of America
- Website: https://www.janelia.org/project-team/cellmap
- Repositories: 24
- Profile: https://github.com/janelia-cellmap
Using computer vision and machine learning techniques to scalably detect subcellular structures, such as organelles, in datasets generated with next-gen vEM
Citation (CITATION.cff)
cff-version: 1.2.0
message: "If you use this software, please cite it as below."
authors:
- family-names: "Patton"
given-names: "William"
orcid: "https://orcid.org/0000-0002-9652-3222"
- family-names: "Rhoades"
given-names: "Jeff L."
orcid: "https://orcid.org/0000-0001-5077-2533"
- family-names: "Zouinkhi"
given-names: "Marwan"
orcid: "https://orcid.org/0000-0002-9441-2908"
- family-names: "Funke"
given-names: "Jan"
orcid: "http://orcid.org/0000-0003-4388-7783"
title: "DaCapo"
version: 0.3.0
doi: 10.48550/arXiv.2408.02834
date-released: 2024-08-05
url: "https://github.com/janelia-cellmap/dacapo"
preferred-citation:
type: article
authors:
- family-names: "Patton"
given-names: "William"
orcid: "https://orcid.org/0000-0002-9652-3222"
- family-names: "Rhoades"
given-names: "Jeff L."
orcid: "https://orcid.org/0000-0001-5077-2533"
- family-names: "Zouinkhi"
given-names: "Marwan"
orcid: "https://orcid.org/0000-0002-9441-2908"
- family-names: "Ackerman"
given-names: "David G."
orcid: "http://orcid.org/0000-0003-0172-6594"
- family-names: "Malin-Mayor"
given-names: "Caroline"
orcid: "https://orcid.org/0000-0002-9627-6030"
- family-names: "Adjavon"
given-names: "Diane"
- family-names: "Heinrich"
given-names: "Larissa"
orcid: "http://orcid.org/0000-0003-2852-6664"
- family-names: "Bennett"
given-names: "Davis"
orcid: "http://orcid.org/0000-0001-7579-2848"
- family-names: "Zubov"
given-names: "Yurii"
orcid: "https://orcid.org/0000-0003-1988-8081"
- family-names: "Project Team"
given-names: "CellMap"
- family-names: "Weigel"
given-names: "Aubrey V."
orcid: "http://orcid.org/0000-0003-1694-4420"
- family-names: "Funke"
given-names: "Jan"
orcid: "http://orcid.org/0000-0003-4388-7783"
doi: 10.48550/arXiv.2408.02834
journal: "arXiv-cs.CV"
title: "DaCapo: a modular deep learning framework for scalable 3D image segmentation"
year: 2024
GitHub Events
Total
- Create event: 30
- Release event: 2
- Issues event: 23
- Watch event: 17
- Delete event: 20
- Member event: 1
- Issue comment event: 112
- Push event: 142
- Pull request review comment event: 1
- Pull request review event: 12
- Pull request event: 125
- Fork event: 5
Last Year
- Create event: 30
- Release event: 2
- Issues event: 23
- Watch event: 17
- Delete event: 20
- Member event: 1
- Issue comment event: 112
- Push event: 142
- Pull request review comment event: 1
- Pull request review event: 12
- Pull request event: 125
- Fork event: 5
Issues and Pull Requests
Last synced: 9 months ago
All Time
- Total issues: 28
- Total pull requests: 211
- Average time to close issues: 27 days
- Average time to close pull requests: 18 days
- Total issue authors: 10
- Total pull request authors: 13
- Average comments per issue: 3.32
- Average comments per pull request: 0.62
- Merged pull requests: 151
- Bot issues: 0
- Bot pull requests: 64
Past Year
- Issues: 14
- Pull requests: 79
- Average time to close issues: 22 days
- Average time to close pull requests: 15 days
- Issue authors: 5
- Pull request authors: 9
- Average comments per issue: 3.71
- Average comments per pull request: 1.13
- Merged pull requests: 51
- Bot issues: 0
- Bot pull requests: 29
Top Authors
Issue Authors
- mzouink (25)
- davidackerman (12)
- rhoadesScholar (10)
- cmalinmayor (5)
- jdeschamps (3)
- atc3 (2)
- neptunes5thmoon (1)
- pattonw (1)
- AJFSalomon (1)
- d-v-b (1)
- adjavon (1)
Pull Request Authors
- github-actions[bot] (151)
- mzouink (127)
- rhoadesScholar (96)
- pattonw (34)
- davidackerman (17)
- cmalinmayor (11)
- vaxenburg (6)
- atc3 (5)
- avweigel (4)
- d-v-b (4)
- adjavon (3)
- psobolewskiPhD (2)
- gbhuang (2)
- yuriyzubov (2)
- eschombu (1)
Top Labels
Issue Labels
Pull Request Labels
Dependencies
- actions/checkout master composite
- actions/setup-python v1 composite
- actions/checkout master composite
- actions/setup-python v2 composite
- ad-m/github-push-action master composite
- sphinx-notes/pages v2 composite
- actions/checkout v2 composite
- actions/setup-python v2 composite
- actions/checkout v2 composite
- actions/setup-python v2 composite
- sphinx-autodoc-typehints *
- sphinx-material *
- attrs *
- bokeh *
- cattrs *
- click *
- daisy >=1.0
- fibsem_tools *
- funlib.evaluate @ git+https://github.com/pattonw/funlib.evaluate
- funlib.geometry >=0.2
- funlib.math >=0.1
- funlib.persistence @ git+https://github.com/janelia-cellmap/funlib.persistence
- gunpowder >=1.3
- lazy-property *
- lsds @ git+https://github.com/funkelab/lsd
- mwatershed >=0.1
- neuroglancer *
- numpy *
- numpy-indexed >=0.3.7
- numpy-indexed *
- pymongo *
- pyyaml *
- simpleitk *
- toml *
- torch *
- tqdm *
- xarray *
- zarr *



