pycytominer
Python package for processing image-based profiling data
Science Score: 59.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
○CITATION.cff file
-
✓codemeta.json file
Found codemeta.json file -
✓.zenodo.json file
Found .zenodo.json file -
✓DOI references
Found 5 DOI reference(s) in README -
✓Academic publication links
Links to: nature.com -
✓Committers with academic emails
4 of 22 committers (18.2%) from academic institutions -
○Institutional organization owner
-
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (17.9%) to scientific vocabulary
Keywords
Keywords from Contributors
Repository
Python package for processing image-based profiling data
Basic Info
- Host: GitHub
- Owner: cytomining
- License: bsd-3-clause
- Language: Python
- Default Branch: main
- Homepage: https://pycytominer.readthedocs.io
- Size: 690 MB
Statistics
- Stars: 124
- Watchers: 5
- Forks: 40
- Open Issues: 102
- Releases: 11
Topics
Metadata Files
README.md

Data processing for image-based profiling
Pycytominer is a suite of common functions used to process high dimensional readouts from high-throughput cell experiments. The tool is most often used for processing data through the following pipeline:
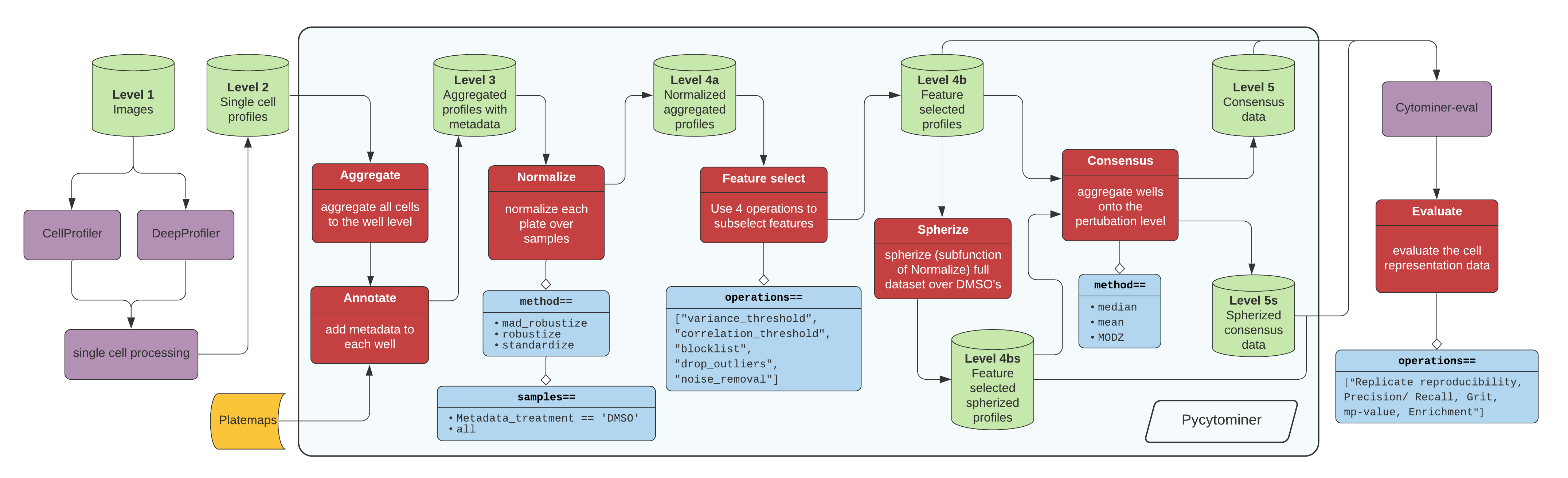
Figure 1. The standard image-based profiling experiment and the role of Pycytominer. (A) In the experimental phase, a scientist plates cells, often perturbing them with chemical or genetic agents and performs microscopy imaging. In image analysis, using CellProfiler for example, a scientist applies several data processing steps to generate image-based profiles. In addition, scientists can apply a more flexible approach by using deep learning models, such as DeepProfiler, to generate image-based profiles. (B) Pycytominer performs image-based profiling to process morphology features and make them ready for downstream analyses. (C) Pycytominer performs five fundamental functions, each implemented with a simple and intuitive API. Each function enables a user to implement various methods for executing operations.
Click here for high resolution pipeline image
Image data flow from a microscope to cell segmentation and feature extraction tools (e.g. CellProfiler or DeepProfiler) (Figure 1A). From here, additional single cell processing tools curate the single cell readouts into a form manageable for Pycytominer input. For CellProfiler, we use cytominer-database or CytoTable. For DeepProfiler, we include single cell processing tools in pycytominer.cyto_utils.
Next, Pycytominer performs reproducible image-based profiling (Figure 1B). The Pycytominer API consists of five key steps (Figure 1C). The outputs generated by Pycytominer are utilized for downstream analysis, which includes machine learning models and statistical testing to derive biological insights.
The best way to communicate with us is through GitHub Issues, where we are able to discuss and troubleshoot topics related to pycytominer.
Please see our CONTRIBUTING.md for details about communicating possible bugs, new features, or other information.
Installation
You can install Pycytominer using the following platforms.
This project follows a <major>.<minor>.<patch> semantic versioning scheme which is used for every release with small variations per platform.
pip (link):
```bash
install pycyotminer from PyPI
pip install pycytominer ```
conda (link):
```bash
install Pycytominer from conda-forge
conda install -c conda-forge pycytominer ```
Docker Hub (link):
Container images of Pycytominer are made available through Docker Hub.
These images follow a tagging scheme that extends our release sematic versioning which may be found within our CONTRIBUTING.md Docker Hub Image Releases documentation.
```bash
pull the latest Pycytominer image and run a module
docker run --platform=linux/amd64 cytomining/pycytominer:latest python -m pycytominer.
pull a commit-based version of Pycytominer (b1bb292) and run an interactive bash session within the container
docker run -it --platform=linux/amd64 cytomining/pycytominer:pycytominer-1.1.0.post16.dev0_b1bb292 bash
pull a scheduled update of pycytominer, map the present working directory to /opt within the container, and run a python script.
docker run -v $PWD:/opt --platform=linux/amd64 cytomining/pycytominer:pycytominer-1.1.0.post16.dev0b1bb292240417 python /opt/script.py ```
Frameworks
Pycytominer is primarily built on top of pandas, also using aspects of SQLAlchemy, sklearn, and pyarrow.
Pycytominer currently supports parquet and compressed text file (e.g. .csv.gz) i/o.
CellProfiler support
Currently, Pycytominer fully supports data generated by CellProfiler, adhering defaults to its specific data structure and naming conventions.
CellProfiler-generated image-based profiles typically consist of two main components:
- Metadata features: This section contains information about the experiment, such as plate ID, well position, incubation time, perturbation type, and other relevant experimental details. These feature names are prefixed with
Metadata_, indicating that the data in these columns contain metadata information. - Morphology features: These are the quantified morphological features prefixed with the default compartments (
Cells_,Cytoplasm_, andNuclei_). Pycytominer also supports non-default compartment names (e.g.,Mito_).
Note, pycytominer.cyto_utils.cells.SingleCells() contains code designed to interact with single-cell SQLite files exported from CellProfiler.
Processing capabilities for SQLite files depends on SQLite file size and your available computational resources (for ex. memory and CPU).
Handling inputs from other image analysis tools (other than CellProfiler)
We recommend pre-harmonizing data using CytoTable when working with data from image analysis tools such as CellProfiler, In Carta, or legacy data systems such as cytominer-database. CytoTable is purpose-built to help prepare data for Pycytominer and includes many presets to help you get started with your work (please also check out our CytoTable preprint).
For example, to resolve potential feature issues in the normalize() function, you must manually specify the morphological features using the features parameter.
The features parameter is also available in other key steps, such as aggregate and feature_select.
If you are using Pycytominer with these other tools, please file an issue to reach out. We'd love to hear from you so that we can learn how to best support broad and multiple use-cases.
API
Pycytominer has five major processing functions:
- Aggregate - Average single-cell profiles based on metadata information (most often "well").
- Annotate - Append metadata (most often from the platemap file) to the feature profile
- Normalize - Transform input feature data into consistent distributions
- Feature select - Exclude non-informative or redundant features
- Consensus - Average aggregated profiles by replicates to form a "consensus signature"
The API is consistent for each of these functions:
```python
Each function takes as input a pandas DataFrame or file path
and transforms the input data based on the provided options and methods
df = function( profilesorpath, features, samples, method, outputfile, additionaloptions... ) ```
Each processing function has unique arguments, see our documentation for more details.
Usage
The default way to use Pycytominer is within python scripts, and using Pycytominer is simple and fun.
The example below demonstrates how to perform normalization with a dataset generated by CellProfiler.
```python
Real world example
import pandas as pd import pycytominer
commit = "da8ae6a3bc103346095d61b4ee02f08fc85a5d98" url = f"https://media.githubusercontent.com/media/broadinstitute/lincs-cell-painting/{commit}/profiles/20160401a54948hrbatch1/SQ00014812/SQ00014812augmented.csv.gz"
df = pd.read_csv(url)
normalizeddf = pycytominer.normalize( profiles=df, method="standardize", samples="Metadatabroad_sample == 'DMSO'" ) ```
Pipeline orchestration
Pycytominer is a collection of different functions with no explicit link between steps. However, some options exist to use Pycytominer within a pipeline framework.
| Project | Format | Environment | Pycytominer usage | | :------------------------------------------------------------------------------- | :-------- | :------------------- | :---------------------- | | Profiling-recipe | yaml | agnostic | full pipeline support | | CellProfiler-on-Terra | WDL | google cloud / Terra | single-cell aggregation | | CytoSnake | snakemake | agnostic | full pipeline support |
A separate project called AuSPICES offers pipeline support up to image feature extraction.
Other functionality
Pycytominer was written with a goal of processing any high-throughput image-based profiling data. However, the initial use case was developed for processing image-based profiling experiments specifically. And, more specifically than that, image-based profiling readouts from CellProfiler measurements from Cell Painting data.
Therefore, we have included some custom tools in pycytominer/cyto_utils that provides other functionality:
CellProfiler CSV collation
If running your images on a cluster, unless you have a MySQL or similar large database set up then you will likely end up with lots of different folders from the different cluster runs (often one per well or one per site), each one containing an Image.csv, Nuclei.csv, etc.
In order to look at full plates, therefore, we first need to collate all of these CSVs into a single file (currently SQLite) per plate.
We currently do this with a library called cytominer-database.
If you want to perform this data collation inside Pycytominer using the cyto_utils function collate (and/or you want to be able to run the tests and have them all pass!), you will need cytominer-database==0.3.4; this will change your installation commands slightly:
```bash
Example for general case commit:
pip install "pycytominer[collate]"
Example for specific commit:
pip install "pycytominer[collate] @ git+https://github.com/cytomining/pycytominer@77d93a3a551a438799a97ba57d49b19de0a293ab" ```
If using pycytominer in a conda environment, in order to run collate.py, you will also want to make sure to add cytominer-database=0.3.4 to your list of dependencies.
Creating a cell locations lookup table
The CellLocation class offers a convenient way to augment a LoadData file with X,Y locations of cells in each image.
The locations information is obtained from a single cell SQLite file.
To use this functionality, you will need to modify your installation command, similar to above:
```bash
Example for general case commit:
pip install "pycytominer[cell_locations]" ```
Example using this functionality:
```bash metadatainput="s3://cellpainting-gallery/test-cpg0016-jump/source4/workspace/loaddatacsv/20210823Batch12/BR00126114/testBR00126114loaddatawithillum.parquet" singlesinglecellinput="s3://cellpainting-gallery/test-cpg0016-jump/source4/workspace/backend/20210823Batch12/BR00126114/testBR00126114.sqlite" augmentedmetadataoutput="~/Desktop/loaddatawithillumandcelllocation_subset.parquet"
python \ -m pycytominer.cytoutils.celllocationscmd \ --metadatainput ${metadatainput} \ --singlecellinput ${singlesinglecellinput} \ --augmentedmetadataoutput ${augmentedmetadataoutput} \ addcelllocation
Check the output
python -c "import pandas as pd; print(pd.readparquet('${augmentedmetadata_output}').head())"
It should look something like this (depends on the width of your terminal):
MetadataPlate MetadataWell MetadataSite ... PathNameOrigRNA ImageNumber CellCenters
0 BR00126114 A01 1 ... s3://cellpainting-gallery/cpg0016-jump/source... 1 [{'NucleiLocationCenterX': 943.512129380054...
1 BR00126114 A01 2 ... s3://cellpainting-gallery/cpg0016-jump/source... 2 [{'NucleiLocationCenterX': 29.9516027655562...
```
Generating a GCT file for morpheus
The software morpheus enables profile visualization in the form of interactive heatmaps.
Pycytominer can convert profiles into a .gct file for drag-and-drop input into morpheus.
```python
Real world example
import pandas as pd import pycytominer
commit = "da8ae6a3bc103346095d61b4ee02f08fc85a5d98" plate = "SQ00014812" url = f"https://media.githubusercontent.com/media/broadinstitute/lincs-cell-painting/{commit}/profiles/20160401a54948hrbatch1/{plate}/{plate}normalizedfeatureselect.csv.gz"
df = pd.readcsv(url) outputfile = f"{plate}.gct"
pycytominer.cytoutils.writegct( profiles=df, outputfile=outputfile ) ```
Citing Pycytominer
If you use pycytominer in your project, please cite our software.
You can see citation information in the 'cite this repository' link at the top right under about section within GitHub.
This information may also be referenced within the CITATION.cff file.
Owner
- Name: cytomining
- Login: cytomining
- Kind: organization
- Repositories: 27
- Profile: https://github.com/cytomining
GitHub Events
Total
- Create event: 47
- Release event: 3
- Issues event: 32
- Watch event: 44
- Delete event: 49
- Issue comment event: 185
- Push event: 132
- Pull request review comment event: 45
- Pull request event: 153
- Pull request review event: 101
- Fork event: 5
Last Year
- Create event: 47
- Release event: 3
- Issues event: 32
- Watch event: 44
- Delete event: 49
- Issue comment event: 185
- Push event: 132
- Pull request review comment event: 45
- Pull request event: 153
- Pull request review event: 101
- Fork event: 5
Committers
Last synced: 12 months ago
Top Committers
| Name | Commits | |
|---|---|---|
| gwaygenomics | g****y@g****m | 270 |
| Niranj | s****j@g****m | 86 |
| Dave Bunten | d****n@c****u | 63 |
| Ken Brewer | k****r | 56 |
| dependabot[bot] | 4****] | 50 |
| Beth Cimini | b****7 | 35 |
| roshankern | r****n@u****u | 23 |
| michaelbornholdt | m****t@o****m | 22 |
| Stephen Fleming | s****1@c****u | 12 |
| Ruifan Pei | r****i@g****m | 11 |
| Shantanu Singh | s****h@b****g | 9 |
| Erik Serrano | 3****a | 8 |
| John Arevalo | j****o@g****m | 5 |
| Hillary Tsang | h****g@g****m | 4 |
| Vince Rubinetti | v****i@g****m | 3 |
| Jenna Tomkinson | 1****n | 2 |
| Charlotte Bunne | b****h | 2 |
| alxndrkalinin | 1****n | 1 |
| Rebecca Senft | s****a@g****m | 1 |
| Erin Weisbart | 5****t | 1 |
| Steve Taylor | s****x@g****m | 1 |
| AdeboyeML | o****e@y****m | 1 |
Committer Domains (Top 20 + Academic)
Issues and Pull Requests
Last synced: 9 months ago
All Time
- Total issues: 156
- Total pull requests: 288
- Average time to close issues: 6 months
- Average time to close pull requests: 20 days
- Total issue authors: 22
- Total pull request authors: 19
- Average comments per issue: 1.75
- Average comments per pull request: 2.3
- Merged pull requests: 235
- Bot issues: 0
- Bot pull requests: 94
Past Year
- Issues: 30
- Pull requests: 161
- Average time to close issues: about 1 month
- Average time to close pull requests: 7 days
- Issue authors: 6
- Pull request authors: 5
- Average comments per issue: 0.47
- Average comments per pull request: 1.86
- Merged pull requests: 135
- Bot issues: 0
- Bot pull requests: 75
Top Authors
Issue Authors
- d33bs (39)
- kenibrewer (32)
- gwaybio (28)
- shntnu (11)
- jenna-tomkinson (11)
- bethac07 (9)
- axiomcura (6)
- ErinWeisbart (4)
- ethancohen123 (2)
- MikeLippincott (2)
- kvshams (1)
- arka2696 (1)
- niranjchandrasekaran (1)
- vincerubinetti (1)
- alxndrkalinin (1)
Pull Request Authors
- d33bs (97)
- dependabot[bot] (94)
- kenibrewer (34)
- gwaybio (17)
- shntnu (11)
- axiomcura (9)
- bethac07 (6)
- johnarevalo (3)
- vincerubinetti (2)
- bunnech (2)
- fefossa (2)
- staylorx (2)
- roshankern (2)
- jenna-tomkinson (2)
- alxndrkalinin (1)
Top Labels
Issue Labels
Pull Request Labels
Packages
- Total packages: 4
-
Total downloads:
- pypi 1,791 last-month
-
Total dependent packages: 1
(may contain duplicates) -
Total dependent repositories: 1
(may contain duplicates) - Total versions: 23
- Total maintainers: 3
proxy.golang.org: github.com/cytomining/pycytominer
- Documentation: https://pkg.go.dev/github.com/cytomining/pycytominer#section-documentation
- License: bsd-3-clause
-
Latest release: v1.2.4
published 10 months ago
Rankings
pypi.org: pycytominer
Python package for processing image-based profiling data
- Homepage: https://pycytominer.readthedocs.io/
- Documentation: https://pycytominer.readthedocs.io/
- License: BSD-3-Clause
-
Latest release: 1.2.4
published 10 months ago
Rankings
Maintainers (3)
conda-forge.org: pycytominer
- Homepage: https://github.com/cytomining/pycytominer
- License: BSD-3-Clause
-
Latest release: 0.2.0
published almost 4 years ago
Rankings
conda-forge.org: pycytominer.collate
- Homepage: https://github.com/cytomining/pycytominer
- License: BSD-3-Clause
-
Latest release: 0.2.0
published almost 4 years ago
Rankings
Dependencies
- 102 dependencies
- boto3 >=1.26.79
- cytominer-database 0.3.4
- fire >=0.5.0
- fsspec >=2023.1.0
- numpy >=1.16.5
- pandas >=1.2.0
- pyarrow >=8.0.0
- python >=3.8
- s3fs >=0.4.2
- scikit-learn >=0.21.2
- scipy >=1.5
- sqlalchemy >=1.3.6, <2
- readthedocs/actions/preview v1 composite
- actions/cache v3 composite
- actions/setup-python v4 composite
- ./.github/actions/setup-env * composite
- actions/checkout v4 composite
- codecov/codecov-action v3 composite
- pre-commit/action v3.0.0 composite
- ./.github/actions/setup-env * composite
- actions/checkout v4 composite
- pypa/gh-action-pypi-publish release/v1 composite
- base latest build
- production latest build
- python 3.11 build

