r2rtf
Easily Create Production-Ready Rich Text Format (RTF) Table and Figure
Science Score: 44.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
✓CITATION.cff file
Found CITATION.cff file -
✓codemeta.json file
Found codemeta.json file -
✓.zenodo.json file
Found .zenodo.json file -
○DOI references
-
○Academic publication links
-
○Committers with academic emails
-
○Institutional organization owner
-
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (20.2%) to scientific vocabulary
Keywords from Contributors
clinical-trials
ectd
packaging-tool
pharmaverse
Last synced: 7 months ago
·
JSON representation
·
Repository
Easily Create Production-Ready Rich Text Format (RTF) Table and Figure
Basic Info
- Host: GitHub
- Owner: Merck
- License: gpl-3.0
- Language: R
- Default Branch: master
- Homepage: https://merck.github.io/r2rtf
- Size: 11.6 MB
Statistics
- Stars: 78
- Watchers: 10
- Forks: 22
- Open Issues: 0
- Releases: 19
Created about 6 years ago
· Last pushed 7 months ago
Metadata Files
Readme
Changelog
License
Citation
README.Rmd
--- output: github_document --- # r2rtf[](https://CRAN.R-project.org/package=r2rtf) [](https://app.codecov.io/gh/Merck/r2rtf) [](https://cran.r-project.org/package=r2rtf) [](https://github.com/Merck/r2rtf/actions/workflows/R-CMD-check.yaml) [](https://cran.r-project.org/package=r2rtf) ```{r setup, include=FALSE} knitr::opts_chunk$set(echo = TRUE) ``` ```{r, echo = FALSE} pkgname <- "r2rtf" ``` ## Overview r2rtf is an R package to create production-ready tables and figures in RTF format. The package is designed with these principles: - Provide simple "verb" functions that correspond to each component of a table, to help you translate data frame(s) to a table in RTF file. - Functions are chainable with pipes (`%>%`). - Only focus on **table format**. - Data manipulation and analysis should be handled by other R packages, for example, tidyverse. - Minimize package dependency. The [R for clinical study reports and submission](https://r4csr.org/) book provides tutorials by using real world examples. ## Installation You can install the package via CRAN: ```r install.packages("r2rtf") ``` Or, install from GitHub: ```r remotes::install_github("Merck/r2rtf") ``` ## Highlighted features The R package r2rtf provides flexibility to enable features below: - Create highly customized RTF tables and figures ready for production. - Simple to use parameters and data structure. - Customized column header: split by `"|"`. - Three required parameters for the output tables (data, filename, column relative width). - Flexible and detail control of table structure. - Format control in cell, row, column and table level for: - Border Type: single, double, dash, dot, etc. - Alignment: left, right, center, decimal. - Column width. - Text appearance: **bold**, *italics*, ~~strikethrough~~, underline and any combinations. - Font size. - Text and border color (`r length(colors())` different colors named in `color()` function). - Special characters: any character in UTF-8 encoding (e.g., Greek, Symbol, Chinese, Japanese, Korean). - Append several tables into one file. - Pagination. - Built-in raw data for validation. ## Simple example ```{r, eval = FALSE} library(dplyr) library(r2rtf) head(iris) %>% rtf_body() %>% # Step 1 Add attributes rtf_encode() %>% # Step 2 Convert attributes to RTF encode write_rtf(file = "ex-tbl.rtf") # Step 3 Write to a .rtf file ``` ```{r, include=FALSE} library(r2rtf) try( { pdftools::pdf_convert("vignettes/pdf/ex-tbl.pdf", dpi = 200, filenames = "ex-tbl.png") pdftools::pdf_convert("vignettes/pdf/efficacy_example.pdf", dpi = 200, filenames = "efficacy_example.png") pdftools::pdf_convert("vignettes/pdf/ae_example.pdf", dpi = 200, filenames = "ae_example.png") files <- c("ex-tbl.png", "efficacy_example.png", "ae_example.png") file.copy(from = files, to = file.path("vignettes/fig", files), overwrite = TRUE) file.remove(files) }, silent = TRUE ) ```
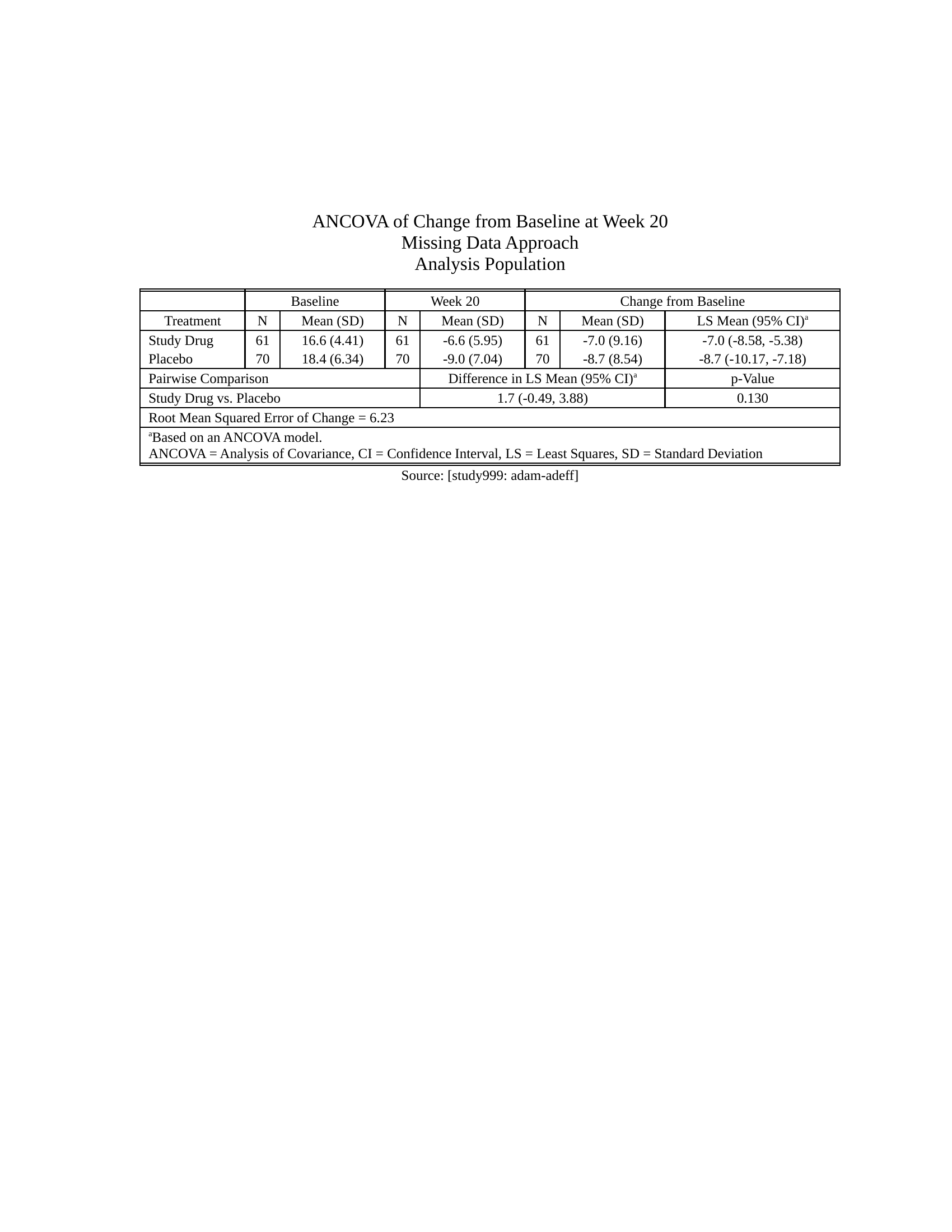
- [More Examples](https://merck.github.io/r2rtf/articles/index.html) ## Example efficacy table - [Source code](https://merck.github.io/r2rtf/articles/example-efficacy.html)Click here to see the output
 ## Example safety table - [Source code](https://merck.github.io/r2rtf/articles/example-ae-summary.html)
## Example safety table - [Source code](https://merck.github.io/r2rtf/articles/example-ae-summary.html)Click here to see the output
 ```{r, include=FALSE, eval = FALSE} # Transfer all RTF files to PDF files files <- list.files("vignettes/rtf", pattern = "*.rtf", full.names = TRUE) r2rtf:::rtf_convert_format( input = files, format = "pdf", output_dir = "vignettes/pdf" ) r2rtf:::rtf_convert_format( input = files, format = "docx", output_dir = "vignettes/docx" ) r2rtf:::rtf_convert_format( input = files, format = "html", output_dir = "vignettes/html" ) files <- list.files("vignettes/rtf", pattern = "*\\.rtf") files_pdf <- list.files("vignettes/pdf", pattern = "*\\.pdf") files_docx <- list.files("vignettes/docx", pattern = "*\\.docx") files_html <- list.files("vignettes/html", pattern = "*\\.html") # Compare RTF and PDF folder filenames. waldo::compare( tools::file_path_sans_ext(files), tools::file_path_sans_ext(files_pdf) ) waldo::compare( tools::file_path_sans_ext(files), tools::file_path_sans_ext(files_docx) ) waldo::compare( tools::file_path_sans_ext(files), tools::file_path_sans_ext(files_html) ) ``` ## Citation If you use this software, please cite it as below. > Wang, S., Ye, S., Anderson, K., & Zhang, Y. (2020). > r2rtf---an R Package to Produce Rich Text Format (RTF) Tables and Figures. > _PharmaSUG_. https://pharmasug.org/proceedings/2020/DV/PharmaSUG-2020-DV-198.pdf A BibTeX entry for LaTeX users is ```bibtex @inproceedings{wang2020r2rtf, title = {{r2rtf}---an {R} Package to Produce {Rich Text Format} ({RTF}) Tables and Figures}, author = {Wang, Siruo and Ye, Simiao and Anderson, Keaven M and Zhang, Yilong}, booktitle = {PharmaSUG}, year = {2020}, url = {https://pharmasug.org/proceedings/2020/DV/PharmaSUG-2020-DV-198.pdf} } ```
```{r, include=FALSE, eval = FALSE} # Transfer all RTF files to PDF files files <- list.files("vignettes/rtf", pattern = "*.rtf", full.names = TRUE) r2rtf:::rtf_convert_format( input = files, format = "pdf", output_dir = "vignettes/pdf" ) r2rtf:::rtf_convert_format( input = files, format = "docx", output_dir = "vignettes/docx" ) r2rtf:::rtf_convert_format( input = files, format = "html", output_dir = "vignettes/html" ) files <- list.files("vignettes/rtf", pattern = "*\\.rtf") files_pdf <- list.files("vignettes/pdf", pattern = "*\\.pdf") files_docx <- list.files("vignettes/docx", pattern = "*\\.docx") files_html <- list.files("vignettes/html", pattern = "*\\.html") # Compare RTF and PDF folder filenames. waldo::compare( tools::file_path_sans_ext(files), tools::file_path_sans_ext(files_pdf) ) waldo::compare( tools::file_path_sans_ext(files), tools::file_path_sans_ext(files_docx) ) waldo::compare( tools::file_path_sans_ext(files), tools::file_path_sans_ext(files_html) ) ``` ## Citation If you use this software, please cite it as below. > Wang, S., Ye, S., Anderson, K., & Zhang, Y. (2020). > r2rtf---an R Package to Produce Rich Text Format (RTF) Tables and Figures. > _PharmaSUG_. https://pharmasug.org/proceedings/2020/DV/PharmaSUG-2020-DV-198.pdf A BibTeX entry for LaTeX users is ```bibtex @inproceedings{wang2020r2rtf, title = {{r2rtf}---an {R} Package to Produce {Rich Text Format} ({RTF}) Tables and Figures}, author = {Wang, Siruo and Ye, Simiao and Anderson, Keaven M and Zhang, Yilong}, booktitle = {PharmaSUG}, year = {2020}, url = {https://pharmasug.org/proceedings/2020/DV/PharmaSUG-2020-DV-198.pdf} } ```Click here to see the output
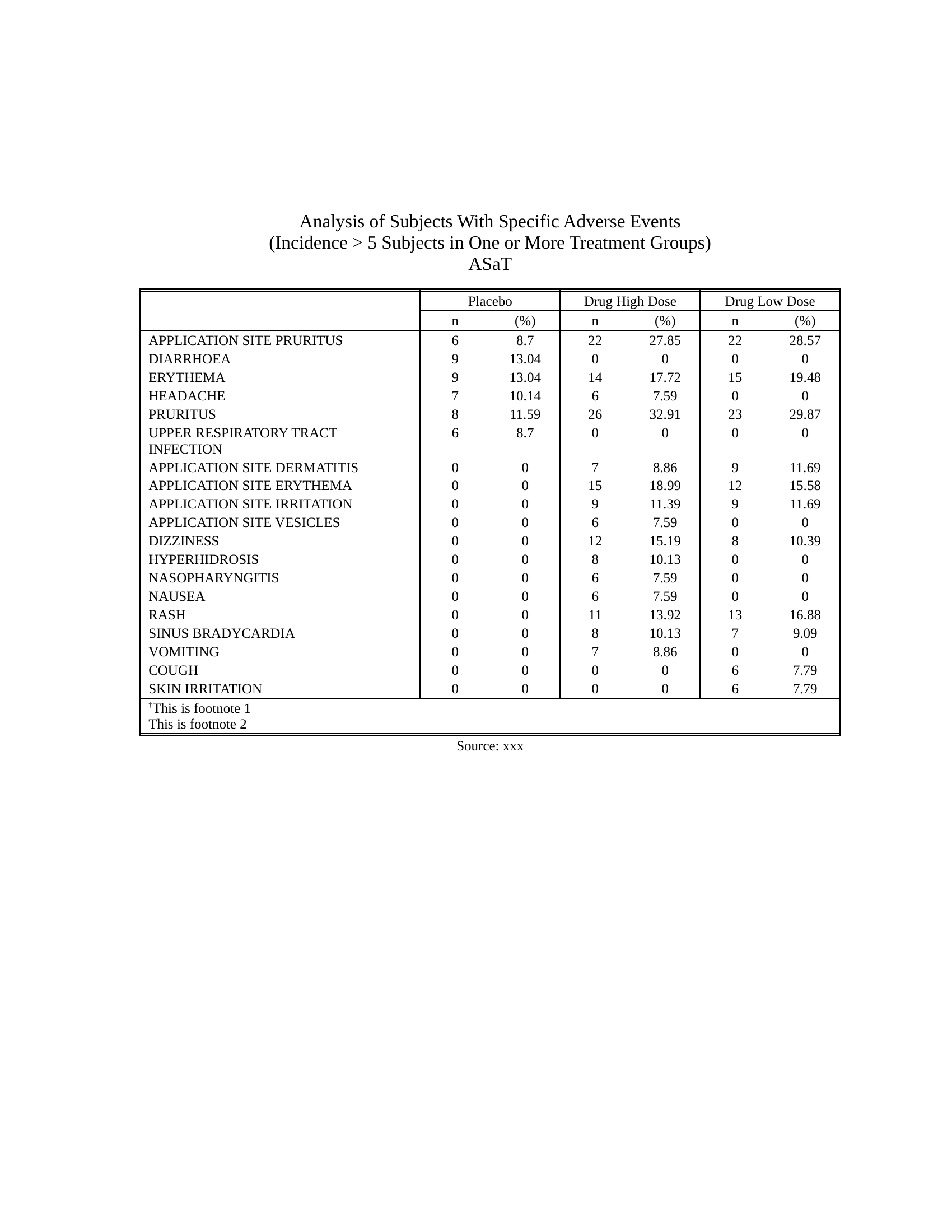
Owner
- Name: Merck Sharp & Dohme Corp. a subsidiary of Merck & Co., Inc.
- Login: Merck
- Kind: organization
- Location: Kenilworth, NJ
- Website: http://www.merck.com
- Repositories: 34
- Profile: https://github.com/Merck
Citation (CITATION.cff)
cff-version: 1.2.0
message: "If you use this software, please cite it as below."
authors:
- family-names: "Zhang"
given-names: "Yilong"
- family-names: "Wang"
given-names: "Siruo"
- family-names: "Ye"
given-names: "Simiao"
- family-names: "Kong"
given-names: "Fansen"
- family-names: "Lang"
given-names: "Brian"
title: "r2rtf: Easily Create Production-Ready Rich Text Format (RTF) Tables and Figures"
version: 1.1.2
date-released: 2024-05-28
url: "https://github.com/Merck/r2rtf"
preferred-citation:
type: conference-paper
authors:
- family-names: "Wang"
given-names: "Siruo"
- family-names: "Ye"
given-names: "Simiao"
- family-names: "Anderson"
given-names: "Keaven"
- family-names: "Zhang"
given-names: "Yilong"
title: "r2rtf---an R Package to Produce Rich Text Format (RTF) Tables and Figures"
year: 2020
collection-title: "PharmaSUG"
url: "https://pharmasug.org/proceedings/2020/DV/PharmaSUG-2020-DV-198.pdf"
GitHub Events
Total
- Create event: 17
- Release event: 3
- Issues event: 34
- Watch event: 7
- Delete event: 16
- Issue comment event: 60
- Push event: 50
- Pull request review event: 13
- Pull request event: 31
- Fork event: 3
Last Year
- Create event: 17
- Release event: 3
- Issues event: 34
- Watch event: 7
- Delete event: 16
- Issue comment event: 60
- Push event: 50
- Pull request review event: 13
- Pull request event: 31
- Fork event: 3
Committers
Last synced: 11 months ago
Top Committers
| Name | Commits | |
|---|---|---|
| Yilong Zhang | y****g@m****m | 171 |
| Nan Xiao | me@n****e | 73 |
| Yilong Zhang | e****7@g****m | 72 |
| BrianLang | b****9@g****m | 26 |
| Yilong Zhang | e****7@f****m | 19 |
| Wang, Benjamin | b****g@m****m | 7 |
| Yihui Xie | x****e@y****e | 5 |
| Fansen Kong | f****g@m****m | 3 |
| Yilong Zhang | z****o@c****m | 3 |
| Shirazi | s****m@c****m | 2 |
| Chao Cheng | 4****4@q****m | 1 |
| Howard Baek | 5****k | 1 |
| Victor | v****o@u****m | 1 |
| Ye, Simiao | s****1@m****m | 1 |
| “Yilong | “****7@f****” | 1 |
Committer Domains (Top 20 + Academic)
merck.com: 4
fb.com”: 1
usbrngspri496f0.merck.com: 1
qq.com: 1
ctchpcvarst002.merck.com: 1
ctchpcpe001.merck.com: 1
yihui.name: 1
fb.com: 1
nanx.me: 1
Issues and Pull Requests
Last synced: 7 months ago
All Time
- Total issues: 99
- Total pull requests: 123
- Average time to close issues: 2 months
- Average time to close pull requests: 3 days
- Total issue authors: 39
- Total pull request authors: 10
- Average comments per issue: 2.73
- Average comments per pull request: 0.55
- Merged pull requests: 108
- Bot issues: 0
- Bot pull requests: 0
Past Year
- Issues: 23
- Pull requests: 32
- Average time to close issues: 13 days
- Average time to close pull requests: 2 days
- Issue authors: 13
- Pull request authors: 3
- Average comments per issue: 2.52
- Average comments per pull request: 0.13
- Merged pull requests: 24
- Bot issues: 0
- Bot pull requests: 0
Top Authors
Issue Authors
- nanxstats (18)
- elong0527 (17)
- Rednose22 (6)
- kaipingyang (4)
- BrianLang (3)
- Chuck1111 (3)
- tdwils (3)
- GlicketySplit (2)
- cglaze11 (2)
- Amex-AZ (2)
- mchastkofsky (2)
- dylanjohnlukes (2)
- schenexel (2)
- cdbccccc (2)
- Danielsite (2)
Pull Request Authors
- elong0527 (57)
- nanxstats (42)
- fb-elong (13)
- wangben718 (8)
- BrianLang (4)
- yihui (2)
- kongfans53 (2)
- howardbaek (1)
- fenguoerbian (1)
- yesimiao (1)
Top Labels
Issue Labels
question (23)
bug (14)
enhancement (10)
cran (10)
documentation (3)
chore (1)
Pull Request Labels
chore (5)
documentation (2)
Packages
- Total packages: 1
-
Total downloads:
- cran 3,175 last-month
- Total docker downloads: 41,971
- Total dependent packages: 7
- Total dependent repositories: 9
- Total versions: 19
- Total maintainers: 1
cran.r-project.org: r2rtf
Easily Create Production-Ready Rich Text Format (RTF) Tables and Figures
- Homepage: https://merck.github.io/r2rtf/
- Documentation: http://cran.r-project.org/web/packages/r2rtf/r2rtf.pdf
- License: GPL-3
-
Latest release: 1.2.0
published 7 months ago
Rankings
Docker downloads count: 0.4%
Forks count: 4.1%
Stargazers count: 5.1%
Average: 7.1%
Downloads: 9.5%
Dependent repos count: 9.9%
Dependent packages count: 13.2%
Maintainers (1)
Last synced:
7 months ago
Dependencies
.github/workflows/R-CMD-check.yaml
actions
- actions/checkout v2 composite
- r-lib/actions/check-r-package v2 composite
- r-lib/actions/setup-pandoc v2 composite
- r-lib/actions/setup-r v2 composite
- r-lib/actions/setup-r-dependencies v2 composite
.github/workflows/pkgdown.yaml
actions
- JamesIves/github-pages-deploy-action 4.1.4 composite
- actions/checkout v2 composite
- r-lib/actions/setup-pandoc v2 composite
- r-lib/actions/setup-r v2 composite
- r-lib/actions/setup-r-dependencies v2 composite
.github/workflows/test-coverage.yaml
actions
- actions/checkout v2 composite
- r-lib/actions/setup-r v2 composite
- r-lib/actions/setup-r-dependencies v2 composite
DESCRIPTION
cran
- R >= 3.5.0 depends
- grDevices * imports
- tools * imports
- covr * suggests
- dplyr * suggests
- emmeans * suggests
- ggplot2 * suggests
- knitr * suggests
- magrittr * suggests
- officer * suggests
- rmarkdown * suggests
- stringi * suggests
- testthat * suggests
- tidyr * suggests
- xml2 * suggests
.github/workflows/style.yaml
actions
- actions/cache v3 composite
- actions/checkout v3 composite
- r-lib/actions/setup-r v2 composite
- r-lib/actions/setup-r-dependencies v2 composite