Science Score: 49.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
○CITATION.cff file
-
✓codemeta.json file
Found codemeta.json file -
✓.zenodo.json file
Found .zenodo.json file -
✓DOI references
Found 8 DOI reference(s) in README -
✓Academic publication links
Links to: nature.com -
○Committers with academic emails
-
○Institutional organization owner
-
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (11.5%) to scientific vocabulary
Repository
Processing Hi-C raw data within R
Basic Info
- Host: GitHub
- Owner: js2264
- License: other
- Language: R
- Default Branch: devel
- Homepage: https://jserizay.com/OHCA/docs/devel/pages/principles.html#hicool-hicstuff-within-r
- Size: 3.64 MB
Statistics
- Stars: 2
- Watchers: 1
- Forks: 0
- Open Issues: 2
- Releases: 0
Metadata Files
README.md
HiCool
Please cite:
Serizay J, Matthey-Doret C, Bignaud A, Baudry L, Koszul R (2024). “Orchestrating chromosome conformation capture analysis with Bioconductor.” Nature Communications, 15, 1-9. doi:10.1038/s41467-024-44761-x.
The HiCool R/Bioconductor package provides an end-to-end interface to
process and normalize Hi-C paired-end fastq reads into .(m)cool files.
- The heavy lifting (fastq mapping, pairs parsing and pairs filtering) is
performed by the underlying lightweight
hicstuffpython library (https://github.com/koszullab/hicstuff). - Pairs filering is done using the approach described in
Cournac et al., 2012 and implemented
in
hicstuff. Cooler(https://github.com/open2c/cooler) library is used to parse pairs into a multi-resolution, balanced.mcoolfile..(m)coolis a compact, indexed HDF5 file format specifically tailored for efficiently storing HiC-based data. The.(m)coolfile format was developed by Abdennur and Mirny and published in 2019.- Internally, all these external dependencies are automatically installed and
managed in R by a
basiliskenvironment.
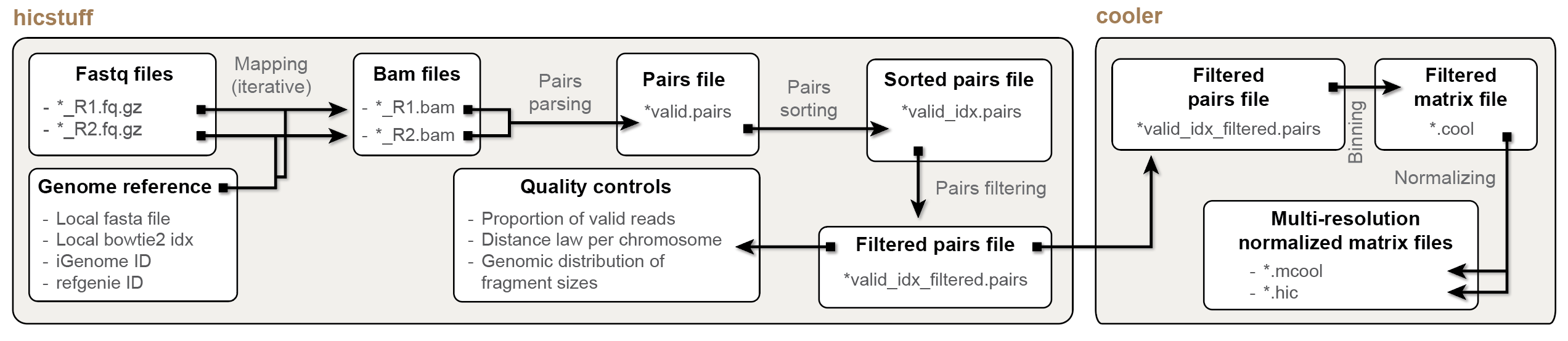
Processing .fastq paired-end files into a .mcool Hi-C contact matrix
The main processing function offered in this package is HiCool().
One simply needs to specify:
- The path to each fastq file;
- The genome reference, as a
.fastasequence, a pre-computedbowtie2index or a supported ID (hg38,mm10,dm6,R64-1-1,WBcel235,GRCz10,Galgal4); - The restriction enzyme(s) used for Hi-C.
r
library(HiCool)
x <- HiCool(
r1 = '<PATH-TO-R1.fq.gz>',
r2 = '<PATH-TO-R2.fq.gz>',
restriction = 'DpnII,HinfI',
genome = 'R64-1-1'
)
```sh
HiCool :: Recovering bowtie2 genome index from AWS iGenomes...
HiCool :: Initiating processing of fastq files [tmp folder: /tmp/RtmpARIRQo/DZ28I8]...
HiCool :: Mapping fastq files...
HiCool :: Best-suited minimum resolution automatically inferred: 1000
HiCool :: Remove unwanted chromosomes...
HiCool :: Generating multi-resolution .mcool file...
HiCool :: Balancing .mcool file...
HiCool :: Tidying up everything for you...
HiCool :: .fastq to .mcool processing done!
HiCool :: Check /home/rsg/repos/HiCool/HiCool folder to find the generated files
HiCool :: Generating HiCool report. This might take a while.
HiCool :: Report generated and available @ sample^mapped-R64-1-1^DZ28I8.html
HiCool :: All processing successfully achieved. Congrats!
```
r
x
```sh
CoolFile object
.mcool file: sample^mapped-R64-1-1^55IONQ.mcool
resolution: 1000
pairs file: sample^55IONQ.pairs
metadata(3): log args stats
```
Output files
```sh
HiCool/
|-- sample^mapped-R64-1-1^55IONQ.html
|-- logs
| |-- sample^mapped-R64-1-1^55IONQ.log
|-- matrices
| |-- sample^mapped-R64-1-1^55IONQ.mcool
|-- pairs
| |-- sample^mapped-R64-1-1^55IONQ.pairs
`-- plots
|-- sample^mapped-R64-1-1^55IONQeventdistance.pdf
|-- sample^mapped-R64-1-1^55IONQeventdistribution.pdf
```
Reporting
On top of processing fastq reads, HiCool provides convenient reports for single/multiple sample(s).
r
x <- importHiCoolFolder(output = 'HiCool/', hash = '55IONQ')
HiCReport(x)
Installation
As an R/Bioconductor package, HiCool should be very easy to install. The only
dependency is R (>= 4.2). In R, one can run:
r
if (!require("BiocManager", quietly = TRUE)) install.packages("BiocManager")
BiocManager::install("HiCool")
The first time a HiCool() function is executed, a basilisk environment
will be automatically set up. In this environment, few dependencies will be
installed:
- python (pinned 3.9.1)
- numpy (pinned 1.23.4)
- bowtie2 (pinned 2.4.5)
- samtools (pinned 1.7)
- hicstuff (pinned 3.1.5)
- cooler (pinned 0.8.11)
HiCExperiment ecosystem
HiCool is integrated within the HiCExperiment ecosystem in Bioconductor.
Read more about the HiCExperiment class and handling Hi-C data in R
here.
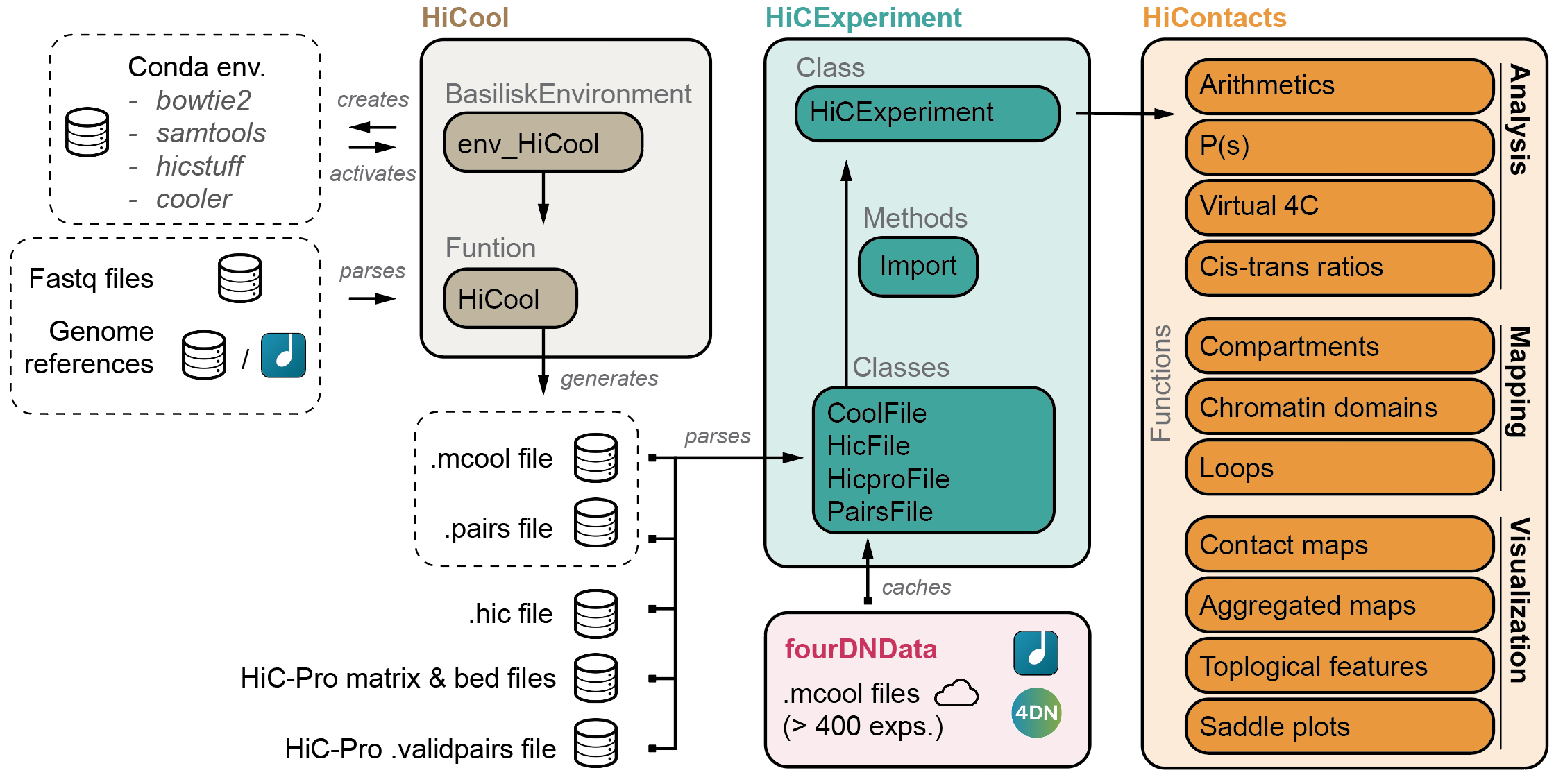
- HiCExperiment: Parsing Hi-C files in R
- HiCool: End-to-end integrated workflow to process fastq files into .cool and .pairs files
- HiContacts: Investigating Hi-C results in R
- HiContactsData: Data companion package
- fourDNData: Gateway package to 4DN-hosted Hi-C experiments
Owner
- Name: Jacques Serizay
- Login: js2264
- Kind: user
- Location: Paris, FR
- Website: js2264.github.io
- Repositories: 12
- Profile: https://github.com/js2264
GitHub Events
Total
- Issues event: 6
- Watch event: 1
- Issue comment event: 16
- Push event: 10
Last Year
- Issues event: 6
- Watch event: 1
- Issue comment event: 16
- Push event: 10
Committers
Last synced: over 2 years ago
Top Committers
| Name | Commits | |
|---|---|---|
| js2264 | j****y@g****m | 50 |
| J Wokaty | j****y | 2 |
Packages
- Total packages: 1
-
Total downloads:
- bioconductor 5,508 total
- Total dependent packages: 0
- Total dependent repositories: 0
- Total versions: 5
- Total maintainers: 1
bioconductor.org: HiCool
HiCool
- Homepage: https://github.com/js2264/HiCool
- Documentation: https://bioconductor.org/packages/release/bioc/vignettes/HiCool/inst/doc/HiCool.pdf
- License: MIT + file LICENSE
-
Latest release: 1.8.0
published 11 months ago


