Science Score: 46.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
○CITATION.cff file
-
✓codemeta.json file
Found codemeta.json file -
○.zenodo.json file
-
✓DOI references
Found 5 DOI reference(s) in README -
✓Academic publication links
Links to: nature.com -
✓Committers with academic emails
1 of 3 committers (33.3%) from academic institutions -
○Institutional organization owner
-
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (10.5%) to scientific vocabulary
Keywords from Contributors
Repository
Analyzing Hi-C data in R with HiCExperiment objects
Basic Info
- Host: GitHub
- Owner: js2264
- License: other
- Language: R
- Default Branch: devel
- Homepage: https://js2264.github.io/OHCA/
- Size: 52.9 MB
Statistics
- Stars: 14
- Watchers: 1
- Forks: 3
- Open Issues: 4
- Releases: 0
Metadata Files
README.md
HiContacts
Please cite:
Serizay J, Matthey-Doret C, Bignaud A, Baudry L, Koszul R (2024). “Orchestrating chromosome conformation capture analysis with Bioconductor.” Nature Communications, 15, 1-9. doi:10.1038/s41467-024-44761-x.
HiContacts provides tools to investigate (m)cool matrices imported in R by HiCExperiment.
It leverages the HiCExperiment class of objects, built on pre-existing Bioconductor objects, namely InteractionSet, GInterations and ContactMatrix (Lun, Perry & Ing-Simmons, F1000Research 2016), and provides analytical and visualization tools to investigate contact maps.
Installation
HiContacts is available in Bioconductor. To install the current release, use:
r
if (!requireNamespace("BiocManager", quietly = TRUE))
install.packages("BiocManager")
BiocManager::install("HiContacts")
To install the most recent version of HiContacts, you can use:
r
install.packages("devtools")
devtools::install_github("js2264/HiContacts")
library(HiContacts)
Citation
If you are using HiContacts in your research, please cite:
Serizay J (2022). HiContacts: HiContacts: R interface to cool files. R package version 1.1.0 https://github.com/js2264/HiContacts.
How to use HiContacts
HiContacts includes a introduction vignette where its usage is
illustrated. To access the vignette, please use:
r
vignette('HiContacts')
Visualising Hi-C contact maps and features
Importing a Hi-C contact maps file with HiCExperiment
r
mcool_file <- HiContactsData::HiContactsData('yeast_wt', format = 'mcool')
range <- 'I:20000-80000' # range of interest
availableResolutions(mcool_file)
hic <- HiCExperiment::import(mcool_file, format = 'mcool', focus = range, resolution = 1000)
hic
Plotting matrices (square or horizontal)
r
plotMatrix(hic, use.scores = 'count')
plotMatrix(hic, use.scores = 'balanced', limits = c(-4, -1))
plotMatrix(hic, use.scores = 'balanced', limits = c(-4, -1), maxDistance = 100000)
Plotting matrices with topological features
r
library(rtracklayer)
mcool_file <- HiContactsData::HiContactsData('yeast_wt', format = 'mcool')
hic <- import(mcool_file, format = 'mcool', focus = 'IV')
loops <- system.file("extdata", 'S288C-loops.bedpe', package = 'HiContacts') |>
import() |>
InteractionSet::makeGInteractionsFromGRangesPairs()
borders <- system.file("extdata", 'S288C-borders.bed', package = 'HiContacts') |>
import()
p <- plotMatrix(
hic, loops = loops, borders = borders,
limits = c(-4, -1), dpi = 120
)
Plotting aggregated matrices (a.k.a. APA plots)
r
contacts <- contacts_yeast()
contacts <- zoom(contacts, resolution = 2000)
aggr_centros <- aggregate(contacts, targets = topologicalFeatures(contacts, 'centromeres'))
plotMatrix(aggr_centros, use.scores = 'detrended', limits = c(-1, 1), scale = 'linear')
Mapping topological features
Chromosome compartments
```r microCmcool <- fourDNData::fourDNData('4DNES14CNC1I', 'mcool') hic <- import(microCmcool, format = 'mcool', resolution = 10000000) genome <- BSgenome.Mmusculus.UCSC.mm10::BSgenome.Mmusculus.UCSC.mm10
- Get compartments
hic <- getCompartments( hic, resolution = 100000, genome = genome, chromosomes = c('chr17', 'chr19') )
- Export compartments as bigwig and bed files
export(IRanges::coverage(metadata(hic)$eigens, weight = 'eigen'), 'microCcompartments.bw') export( topologicalFeatures(hic, 'compartments')[topologicalFeatures(hic, 'compartments')$compartment == 'A'], 'microCA-compartments.bed' ) export( topologicalFeatures(hic, 'compartments')[topologicalFeatures(hic, 'compartments')$compartment == 'B'], 'microC_B-compartments.bed' )
- Generate saddle plot
plotSaddle(hic) ```
Diamond insulation score and chromatin domains borders
```r
- Compute insulation score
hic <- refocus(hic, 'chr19:1-30000000') |> zoom(resolution = 10000) |> getDiamondInsulation(window_size = 100000) |> getBorders()
- Export insulation as bigwig track and borders as bed file
export(IRanges::coverage(metadata(hic)$insulation, weight = 'insulation'), 'microCinsulation.bw') export(topologicalFeatures(hic, 'borders'), 'microCborders.bed') ```
In-depth analysis of HiCExperiment objects
Arithmetics
Detrend
Autocorrelate
Divide
Merge
Distance law, a.k.a. P(s)
r
hic <- import(CoolFile(
mcool_file,
pairs = HiContactsData::HiContactsData('yeast_wt', format = 'pairs.gz')
))
ps <- distanceLaw(hic)
plotPs(ps, ggplot2::aes(x = binned_distance, y = norm_p))
Virtual 4C
r
hic <- import(CoolFile(mcool_file))
v4C <- virtual4C(hic, viewpoint = GRanges('V:150000-170000'))
plot4C(v4C)
Cis-trans ratios
r
hic <- import(CoolFile(mcool_file))
cisTransRatio(hic)
Scalograms
```r
```
HiCExperiment ecosystem
HiCool is integrated within the HiCExperiment ecosystem in Bioconductor.
Read more about the HiCExperiment class and handling Hi-C data in R
here.
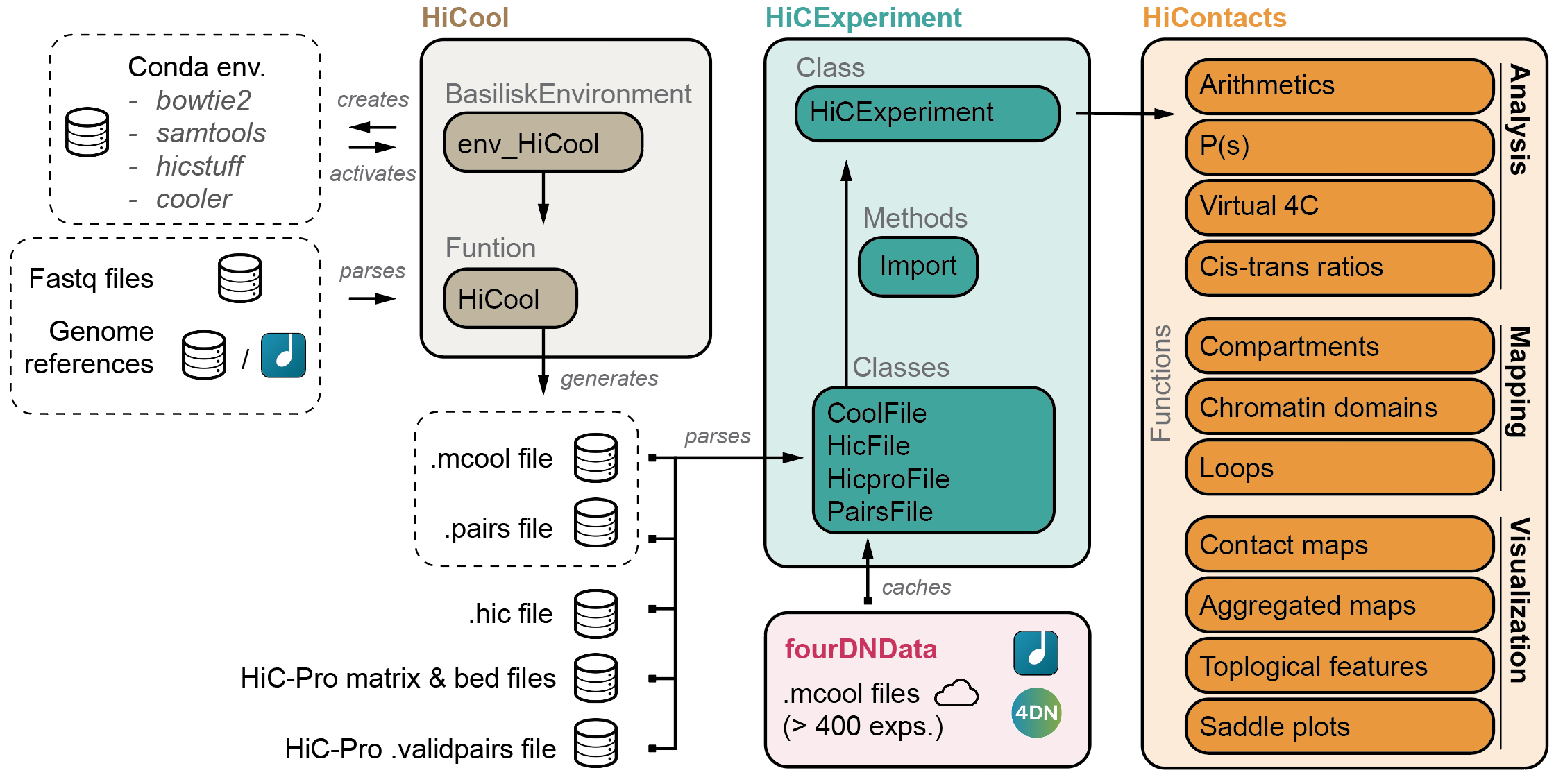
- HiCExperiment: Parsing Hi-C files in R
- HiCool: End-to-end integrated workflow to process fastq files into .cool and .pairs files
- HiContacts: Investigating Hi-C results in R
- HiContactsData: Data companion package
- fourDNData: Gateway package to 4DN-hosted Hi-C experiments
Owner
- Name: Jacques Serizay
- Login: js2264
- Kind: user
- Location: Paris, FR
- Website: js2264.github.io
- Repositories: 12
- Profile: https://github.com/js2264
GitHub Events
Total
- Issues event: 4
- Watch event: 3
- Issue comment event: 5
- Push event: 5
- Fork event: 1
Last Year
- Issues event: 4
- Watch event: 3
- Issue comment event: 5
- Push event: 5
- Fork event: 1
Committers
Last synced: over 2 years ago
Top Committers
| Name | Commits | |
|---|---|---|
| js2264 | j****y@g****m | 215 |
| J Wokaty | j****y@s****u | 2 |
| J Wokaty | j****y | 2 |
Committer Domains (Top 20 + Academic)
Issues and Pull Requests
Last synced: 8 months ago
All Time
- Total issues: 16
- Total pull requests: 0
- Average time to close issues: 2 months
- Average time to close pull requests: N/A
- Total issue authors: 14
- Total pull request authors: 0
- Average comments per issue: 2.81
- Average comments per pull request: 0
- Merged pull requests: 0
- Bot issues: 0
- Bot pull requests: 0
Past Year
- Issues: 8
- Pull requests: 0
- Average time to close issues: about 1 month
- Average time to close pull requests: N/A
- Issue authors: 7
- Pull request authors: 0
- Average comments per issue: 1.25
- Average comments per pull request: 0
- Merged pull requests: 0
- Bot issues: 0
- Bot pull requests: 0
Top Authors
Issue Authors
- xiasijian (2)
- LiNk-NY (1)
- ggmirizio (1)
- narzouni (1)
- GuidoBarzaghi (1)
- mannekim (1)
- hackkr (1)
- matteozoia4 (1)
- ratheraarif (1)
- evaham1 (1)
- PengfeiInTuebingen (1)
- rpetroll (1)
- xl5525 (1)
Pull Request Authors
Top Labels
Issue Labels
Pull Request Labels
Packages
- Total packages: 1
-
Total downloads:
- bioconductor 8,528 total
- Total dependent packages: 2
- Total dependent repositories: 0
- Total versions: 5
- Total maintainers: 1
bioconductor.org: HiContacts
Analysing cool files in R with HiContacts
- Homepage: https://github.com/js2264/HiContacts
- Documentation: https://bioconductor.org/packages/release/bioc/vignettes/HiContacts/inst/doc/HiContacts.pdf
- License: MIT + file LICENSE
-
Latest release: 1.10.0
published 12 months ago
Rankings
Maintainers (1)
Dependencies
- R >= 4.2 depends
- BiocParallel * imports
- GenomeInfoDb * imports
- GenomicInteractions * imports
- GenomicRanges * imports
- HiContactsData * imports
- IRanges * imports
- InteractionSet * imports
- Matrix * imports
- S4Vectors * imports
- corrr * imports
- data.table * imports
- dplyr * imports
- ggplot2 * imports
- ggrastr * imports
- glue * imports
- methods * imports
- plyranges * imports
- purrr * imports
- reticulate * imports
- rhdf5 * imports
- scales * imports
- stringr * imports
- tibble * imports
- tidyr * imports
- tools * imports
- zeallot * imports
- cowplot * suggests
- knitr * suggests
- testthat >= 3.0.0 suggests
- neurogenomics/rworkflows master composite


