cnvpytor
a python extension of CNVnator -- a tool for CNV analysis from depth-of-coverage by mapped reads
Science Score: 49.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
○CITATION.cff file
-
✓codemeta.json file
Found codemeta.json file -
✓.zenodo.json file
Found .zenodo.json file -
✓DOI references
Found 4 DOI reference(s) in README -
○Academic publication links
-
✓Committers with academic emails
3 of 8 committers (37.5%) from academic institutions -
○Institutional organization owner
-
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (11.1%) to scientific vocabulary
Repository
a python extension of CNVnator -- a tool for CNV analysis from depth-of-coverage by mapped reads
Basic Info
- Host: GitHub
- Owner: abyzovlab
- License: mit
- Language: Python
- Default Branch: master
- Size: 135 MB
Statistics
- Stars: 202
- Watchers: 7
- Forks: 30
- Open Issues: 59
- Releases: 4
Metadata Files
README.md

3D printable CNVpytor logo (stl file)
CNVpytor - a python extension of CNVnator
CNVpytor is a Python package and command line tool for CNV/CNA analysis from depth-of-coverage by mapped reads developed in Abyzov Lab, Mayo Clinic.
Follow CNVpytor Twitter account.
New in version 1.3.1
What's new: - Reduced Pytor file size by compressing the BAF likelihood matrix - Option to avoid storing the full BAF likelihood matrix (-nolh), drastically reducing the final Pytor file size to less than 50 MB - If the full BAF likelihood matrix is not stored in Pytor file, during -call step likelihood will be calculated during run time - Introduced plotting parameter "lhlite," used when the full BAF likelihood matrix is not present in the Pytor file - Implemented log scale for Manhattan plot (#126) - Added plot RD difference/ratio between two samples (#151) - Updated the code for VCF output - Included an error log for missing annotation links in reference genome settings - Added matplotlibuse parameter to set the Matplotlib backend
Citing CNVpytor
CNVpytor: a tool for copy number variation detection and analysis from read depth and allele imbalance in whole-genome sequencing
Milovan Suvakov, Arijit Panda, Colin Diesh, Ian Holmes, Alexej Abyzov, GigaScience, Volume 10, Issue 11, November 2021, giab074
https://doi.org/10.1093/gigascience/giab074
Learn how to use CNVpytor in 10 minutes
- Getting started with command line interface
- Jupyter notebook: How to use CNVpytor from Python
- Google Colab: With CEPH trio example dataset
- Video Tutorial: 3-minute YT demo
pytor file support in igv.js and igv-webapp
- Demo Video
- Links
- https://github.com/igvteam/igv.js/wiki/cnvpytor
- https://igv.org/app/
- Test Link: https://arpanda.github.io/igv.js/examples/cnvpytor/
Gallery
| | |
|---|---|
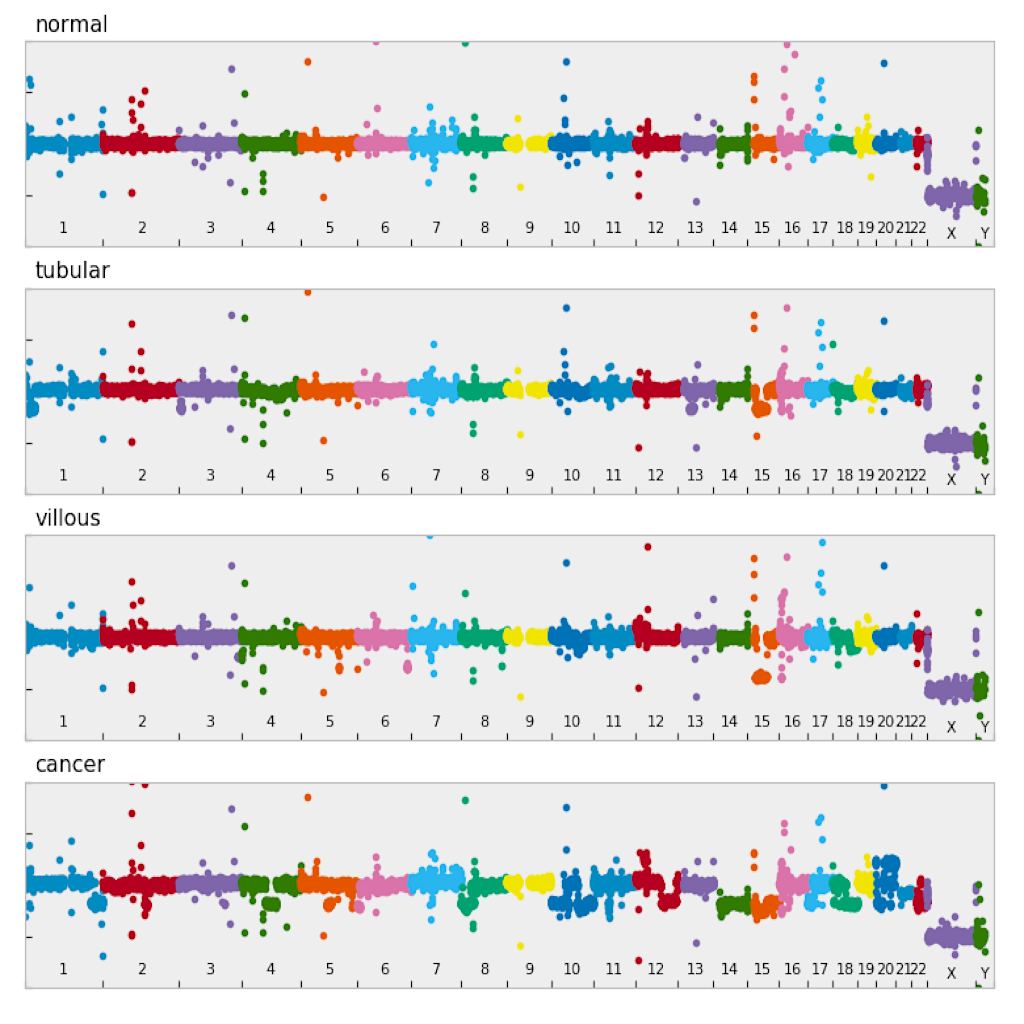
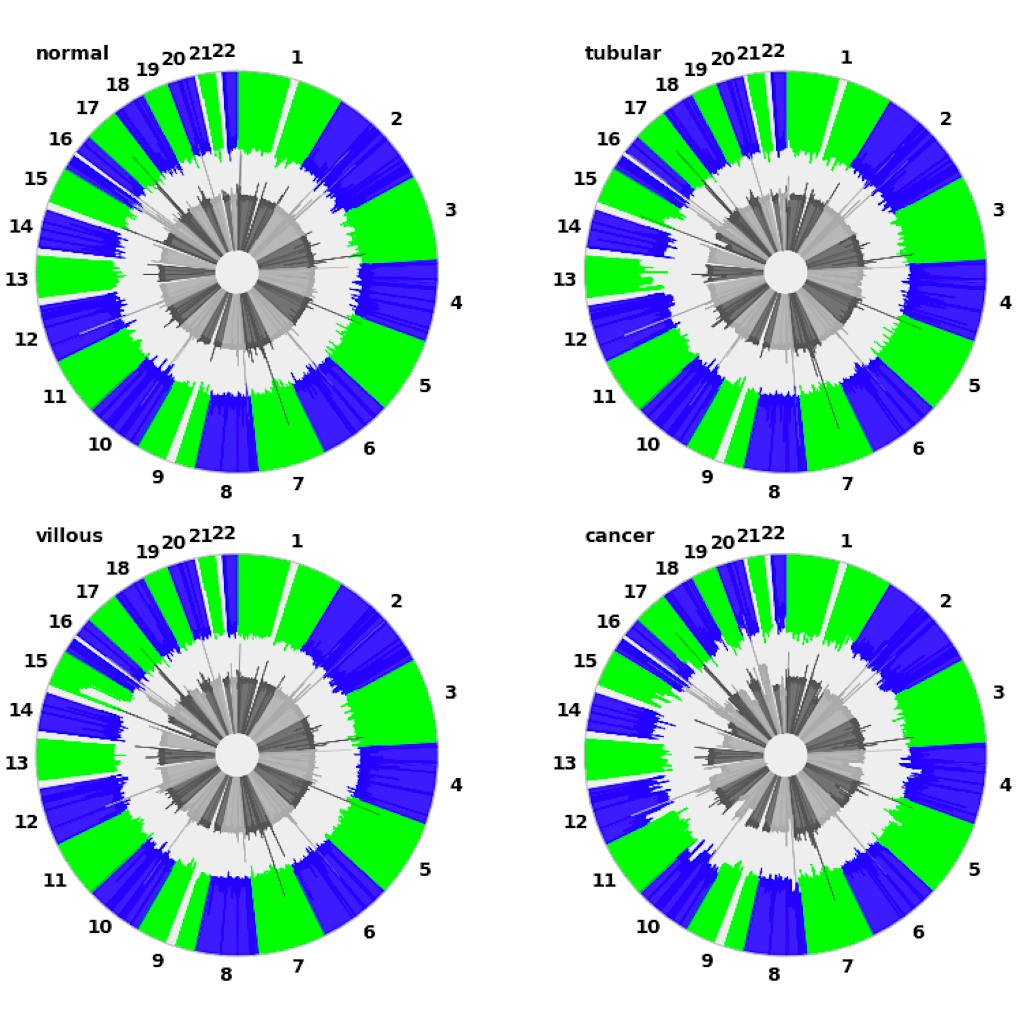
| Manhattan plot (see example)| Circular plot (see example)|
| |
| |
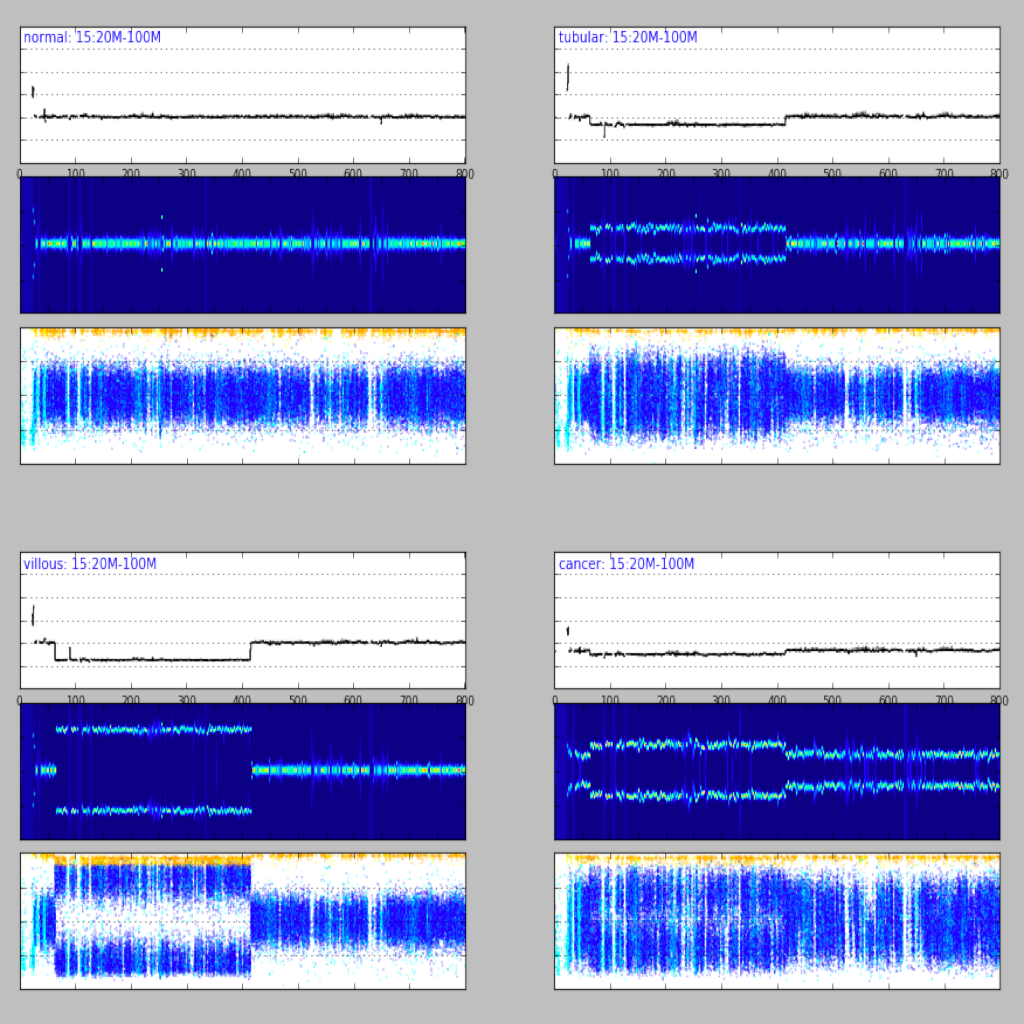
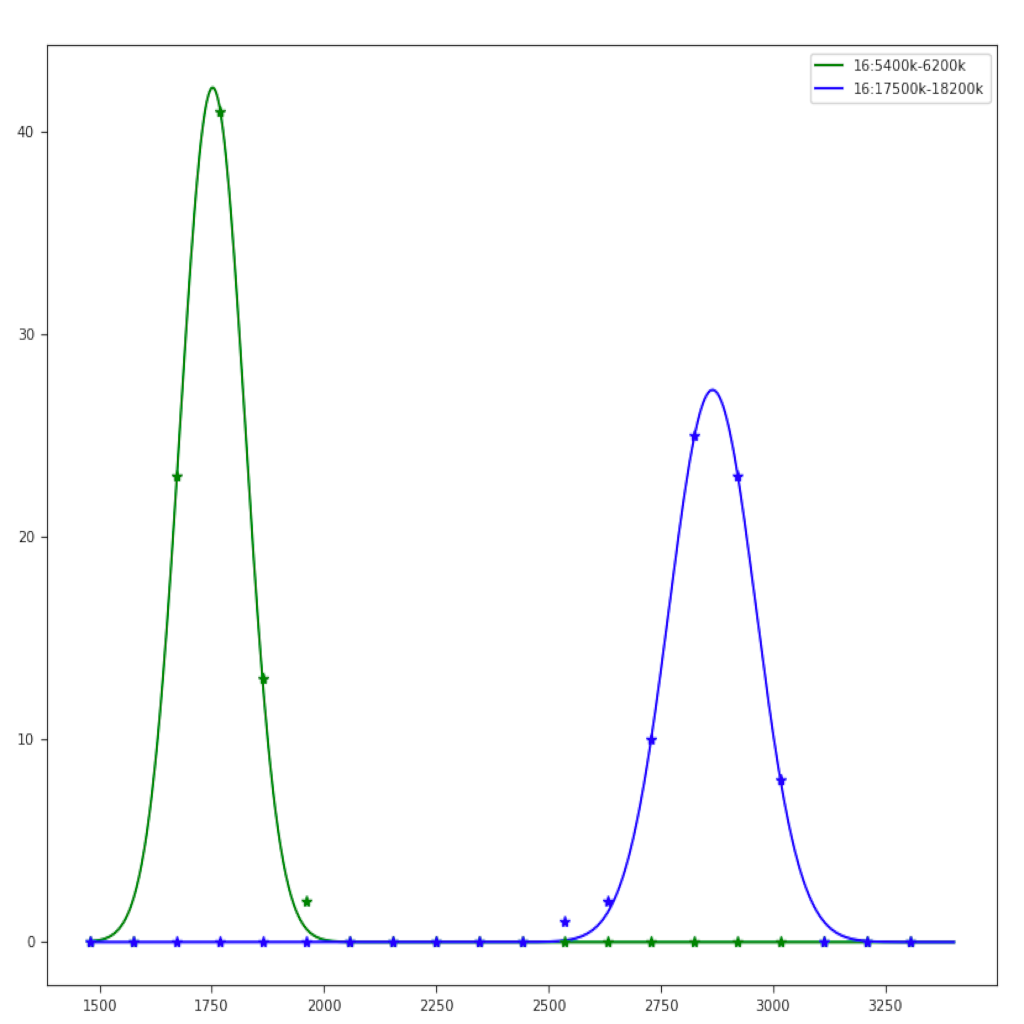
| Region plot (see example)| Compare regions (see example)|
|
|
| Region plot (see example)| Compare regions (see example)|
| |
| |
| Merging and annotating calls (see example)|Call somatic CNAs (see example)|
|
|
| Merging and annotating calls (see example)|Call somatic CNAs (see example)|
|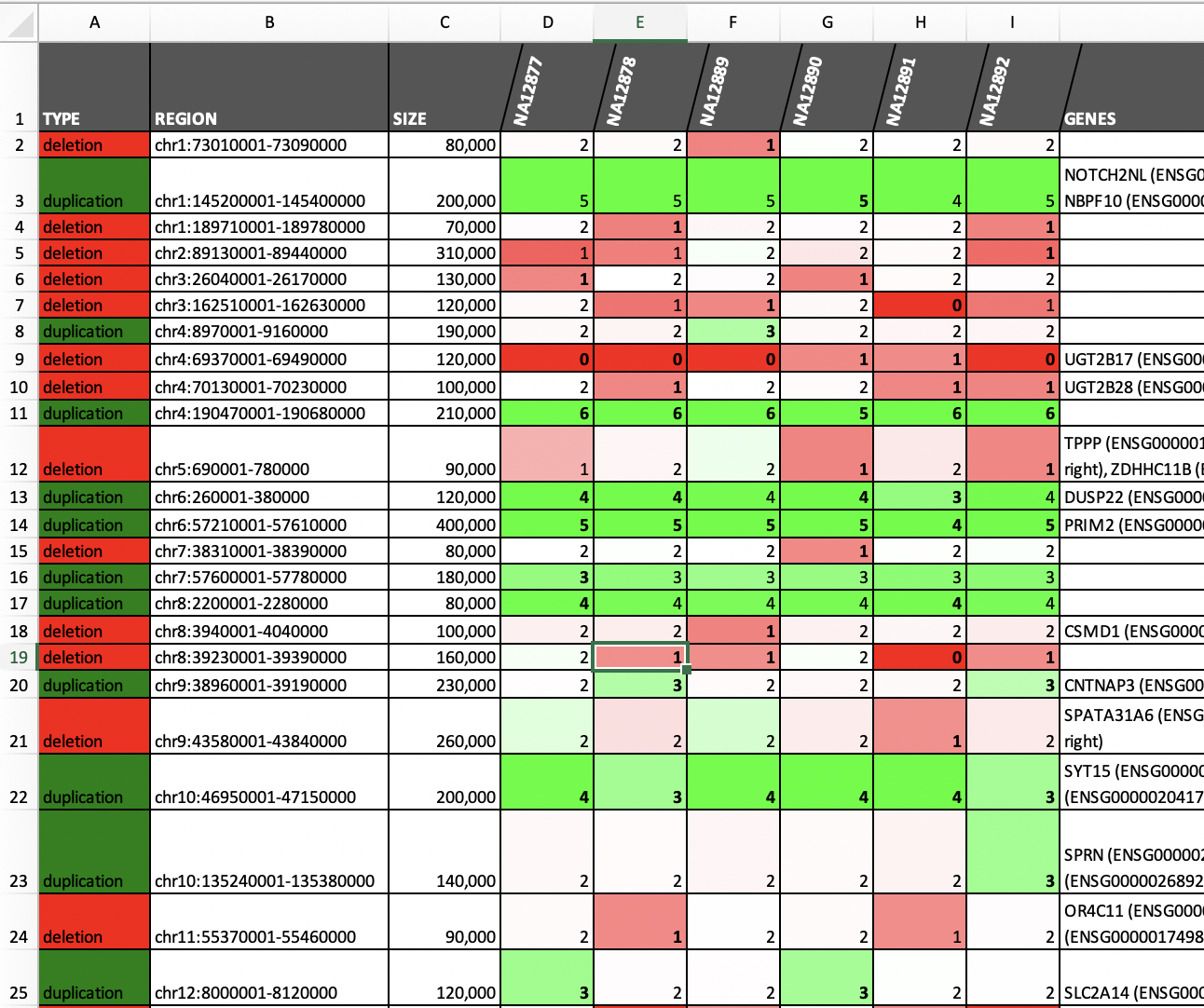 |
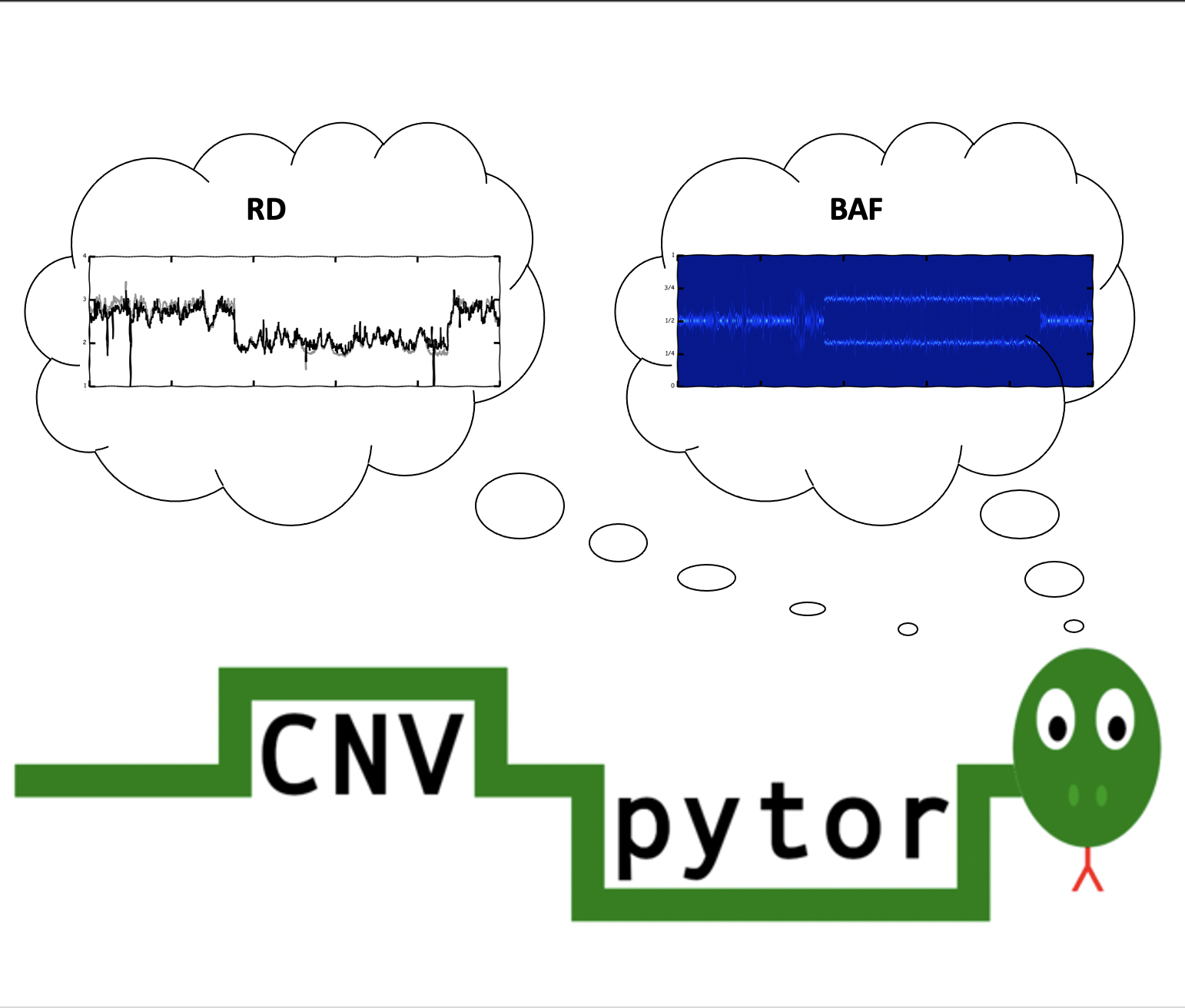
|
Install
Dependencies
- requests>=2.0
- gnureadline
- pathlib>=1.0
- pysam>=0.15
- numpy>=1.16
- scipy>=1.1
- matplotlib>=2.2
- h5py>=2.9
- xlsxwriter>=1.3
- pathlib>=1.0
Optional:
- pyBigWig - for JBrowse export functionality
- ROOT - for CNVnator root import/export functionality
- seaborn - for additional plotting styles
Install by cloning from GitHub
```
git clone https://github.com/abyzovlab/CNVpytor.git cd CNVpytor pip install .
For single user (without admin privileges) use:pip install --user . ```
Install using pip
Version (v1.2.1) is available using pip directly: ```
pip install cnvpytor cnvpytor -download ```
Use as a command line tool
Diagram made using Draw.io.
Call CNVs using read depth:
```
cnvpytor -root file.pytor -rd file.bam cnvpytor -root file.pytor -his 1000 10000 100000 cnvpytor -root file.pytor -partition 1000 10000 100000 cnvpytor -root file.pytor -call 1000 10000 100000 ```
Importing and using single nucleotide polymorphism data:
```
cnvpytor -root file.pytor -snp file.vcf -sample samplename cnvpytor -root file.pytor -pileup file.bam # OPTIONAL cnvpytor -root file.pytor -masksnps # OPTIONAL cnvpytor -root file.pytor -baf 10000 100000 ```
Filtering calls from view mode
```
cnvpytor -root file.pytor -view 100000 print calls set Q0range 0 0.5 set sizerange 100000 inf print calls set prange 0 0.00001 set printfilename output.xls print calls set printfilename output.vcf print calls
Annotating filtered calls:cnvpytor -root file.pytor -view 100000 set Q0range 0 0.5 set sizerange 100000 inf set printfilename output.tsv set annotate print calls ```
Merging calls from multiple samples
```
cnvpytor -root file1.pytor file2.pytor ... -view 100000 print mergedcalls set Q0range 0 0.5 set sizerange 100000 inf set printfilename output.xls print mergedcalls
Plotting all merged calls:cnvpytor -root file1.pytor file2.pytor ... -view 100000 set Q0range 0 0.5 set sizerange 100000 inf set printfilename output.xls set print set outputfilename prefix.png print mergedcallsAnnotating merged calls:cnvpytor -root file1.pytor file2.pytor ... -view 100000 set Q0range 0 0.5 set sizerange 100000 inf set printfilename output.xls set annotate print mergedcalls ```
Genotyping from command line
```
cnvpytor -root file.pytor -genotype 10000 100000 12:11396601-11436500 12:11396601-11436500 1.933261 1.937531 22:20999401-21300400 22:20999401-21300400 1.949186 1.957068 ```
Genotyping with additional informations: ```
cnvpytor -root file.pytor -genotype 10000 100000 -a 12:11396601-11436500 12:11396601-11436500 2.0152 1.629621e+04 9.670589e+08 0.0000 0.0000 4156900 1.0000 50 4 0.0000 1.000000e+00 ```
Genotyping using P filtered (1000 Genome Project strict mask) RD signal: ```
cnvpytor -root file.pytor -genotype 10000 100000 -a -rdusemask 1:800k-900k 1:800000-900000 2.3012 1.032124e+01 8.296037e+06 0.0021 0.0000 278700 0.8000 48 28 0.0000 1.000000e+00 ```
Plot from interactive mode
CNVpytor view interactive mode is implemented with
To enter interactive mode use '-view bin_size' option:
```
cnvpytor -root file.pytor -view 10000 cnvpytor> chr1:1M-50M cnvpytor> rd cnvpytor> set panels rd likelihood cnvpytor> show Parameters * bafcolors: ['gray', 'black', 'red', 'green', 'blue'] * binsize: 100000 * chrom: [] * contrast: 20 * dpi: 200 * filetitles: [] * grid: auto * lhcolors: ['yellow'] * markersize: auto * minsegmentsize: 0 * outputfilename: * panels: ['rd'] * plotfile: 0 * plotfiles: [0] 0: file.pytor * rdcall: True * rdcallmosaic: False * rdcircularcolors: ['#555555', '#aaaaaa'] * rdcolors: ['grey', 'black', 'red', 'green', 'blue'] * rdmanhattancall: False * rdmanhattanrange: [0, 2] * rdpartition: True * rdrange: [0, 3] * rdraw: True * rdusegccorr: True * rdusemask: False * snpcall: False * snpcircularcolors: ['#00ff00', '#0000ff'] * snpcolors: ['yellow', 'orange', 'cyan', 'blue', 'lime', 'green', 'yellow', 'orange'] * snpuseid: False * snpusemask: True * snpuse_phase: False * style: None * xkcd: False
cnvpytor> help markersize
markersize Size of markers used in scatter like plots (e.g. manhattan, snp).
TYPE float or str
DEFAULT auto
PLOTS AFFECTS manhattan, snp, region plot with snp panel
EXAMPLE(s) set markersize 10 set markersize auto
SEE ALSO rdcolors, snpcolors, bafcolors, lhcolors
cnvpytor> set bin_size 100000 cnvpytor> chr1:1M-50M chr2:60M-65M > filename.png ```
Plot from script
```
echo "rdstat" | cnvpytor -root file.pytor -view 100000 -o prefix.png
cnvpytor -root file.pytor -view 100000 <<ENDL set rdusemask set markersize 1 set grid vertical set output_filename prefix.png manhattan circular ENDL
cnvpytor -root file.pytor -view 100000 < script.spytor
```
Persistent history and viewer configuration (experimental)
CNVpytor will automatically store command line history into file ~/.cnvpytor/history if there is directory
~/.cnvpytor. To enable this functionality create this directory:
```
mkdir ~/.cnvpytor ```
To configure viewer parameters create file viewer.conf within same directory in following format:
{
'panels': ['rd', 'likelihood'],
'snp_colors': ['orange', 'brown', 'green', 'blue', 'green', 'blue', 'orange', 'brown']
}
This way you can set any parameter using python syntax. Any parameter specified here will overwrite
parameters provided in command line.
Use as a Python package
CNVpytor is not just command line tool but also Python package.
For more details check API Documentation or see examples in Jupyter notebook.
Bugs
Please report any bugs that you find on GitHub: https://github.com/abyzovlab/CNVpytor/issues
Or, even better, fork the repository on GitHub and create a pull request.
License
Released under MIT licence.
Owner
- Name: Abyzov lab
- Login: abyzovlab
- Kind: organization
- Location: Rochester, MN
- Website: http://abyzovlab.org
- Repositories: 11
- Profile: https://github.com/abyzovlab
Software packages developed in Abyzov's lab
GitHub Events
Total
- Create event: 2
- Release event: 1
- Issues event: 45
- Watch event: 20
- Issue comment event: 58
- Push event: 7
- Pull request event: 1
- Pull request review event: 1
- Fork event: 2
Last Year
- Create event: 2
- Release event: 1
- Issues event: 45
- Watch event: 20
- Issue comment event: 58
- Push event: 7
- Pull request event: 1
- Pull request review event: 1
- Fork event: 2
Committers
Last synced: over 2 years ago
Top Committers
| Name | Commits | |
|---|---|---|
| Milovan Suvakov | s****v@g****m | 276 |
| arpanda | a****a@g****m | 70 |
| Mikhail Dozmorov | m****v@g****m | 2 |
| Bradford Powell | b****w@u****u | 1 |
| abyzov | a****j@m****u | 1 |
| Chase Mateusiak | c****m@w****u | 1 |
| Bradford Powell | b****w@d****g | 1 |
| chase mateusiak | c****k@g****m | 1 |
Committer Domains (Top 20 + Academic)
Issues and Pull Requests
Last synced: 7 months ago
All Time
- Total issues: 169
- Total pull requests: 21
- Average time to close issues: about 2 months
- Average time to close pull requests: 29 days
- Total issue authors: 116
- Total pull request authors: 6
- Average comments per issue: 2.65
- Average comments per pull request: 0.24
- Merged pull requests: 19
- Bot issues: 0
- Bot pull requests: 0
Past Year
- Issues: 32
- Pull requests: 0
- Average time to close issues: 11 days
- Average time to close pull requests: N/A
- Issue authors: 28
- Pull request authors: 0
- Average comments per issue: 1.91
- Average comments per pull request: 0
- Merged pull requests: 0
- Bot issues: 0
- Bot pull requests: 0
Top Authors
Issue Authors
- zainabae (10)
- gevro (6)
- xiucz (4)
- GENEDECODERS (4)
- hayeon-pak (3)
- maryawood (3)
- KrithikaSubramanian (3)
- NinaGerrekens (3)
- lucsnip (3)
- ZYongQi (3)
- Swanthana-bioinfo (3)
- oceancongliu (2)
- mpiersonsmela (2)
- kaqisekuzi (2)
- NannapatS (2)
Pull Request Authors
- suvakov (14)
- arpanda (12)
- archmageirvine (2)
- bpow (1)
- ManavalanG (1)
- cmatKhan (1)
Top Labels
Issue Labels
Pull Request Labels
Packages
- Total packages: 1
-
Total downloads:
- pypi 577 last-month
- Total dependent packages: 0
- Total dependent repositories: 1
- Total versions: 7
- Total maintainers: 2
pypi.org: cnvpytor
Python extension of CNVnator
- Homepage: https://github.com/abyzovlab/CNVpytor
- Documentation: https://cnvpytor.readthedocs.io/
- License: MIT License
-
Latest release: 1.3.2
published about 1 year ago
Rankings
Dependencies
- gnureadline *
- h5py >=2.9
- matplotlib >=2.2
- numpy >=1.16
- pathlib >=1.0
- pysam >=0.15
- requests >=2.0
- scipy >=1.1
- xlsxwriter >=1.3
- gnureadline *
- h5py >=2.9
- matplotlib >=2.2
- numpy >=1.16
- pysam >=0.15
- requests >=2.0
- scipy >=1.1
- xlsxwriter >=1.3