magic-impute
MAGIC (Markov Affinity-based Graph Imputation of Cells), is a method for imputing missing values restoring structure of large biological datasets.
Science Score: 46.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
○CITATION.cff file
-
✓codemeta.json file
Found codemeta.json file -
○.zenodo.json file
-
✓DOI references
Found 1 DOI reference(s) in README -
✓Academic publication links
Links to: zenodo.org -
✓Committers with academic emails
3 of 19 committers (15.8%) from academic institutions -
○Institutional organization owner
-
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (12.3%) to scientific vocabulary
Keywords from Contributors
Repository
MAGIC (Markov Affinity-based Graph Imputation of Cells), is a method for imputing missing values restoring structure of large biological datasets.
Basic Info
Statistics
- Stars: 363
- Watchers: 21
- Forks: 98
- Open Issues: 33
- Releases: 9
Metadata Files
README.md
Markov Affinity-based Graph Imputation of Cells (MAGIC)
Markov Affinity-based Graph Imputation of Cells (MAGIC) is an algorithm for denoising high-dimensional data most commonly applied to single-cell RNA sequencing data. MAGIC learns the manifold data, using the resultant graph to smooth the features and restore the structure of the data.
To see how MAGIC can be applied to single-cell RNA-seq, elucidating the epithelial-to-mesenchymal transition, read our publication in Cell.
MAGIC has been implemented in Python, Matlab, and R.
To get started immediately, check out our tutorials:
Python
- Epithelial-to-Mesenchymal Transition Tutorial
- Bone Marrow Tutorial ##### R
- Epithelial-to-Mesenchymal Transition Tutorial
- Bone Marrow Tutorial
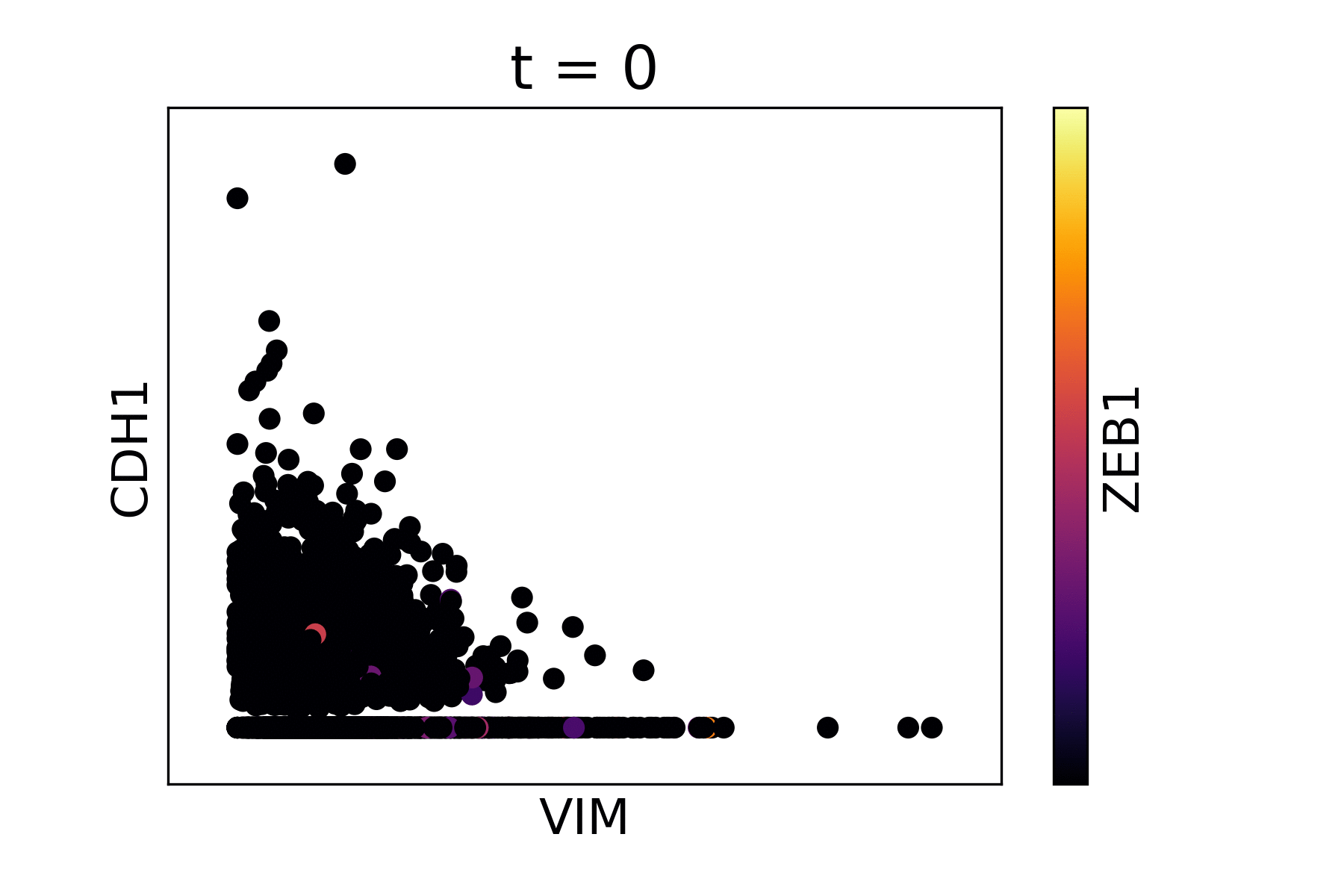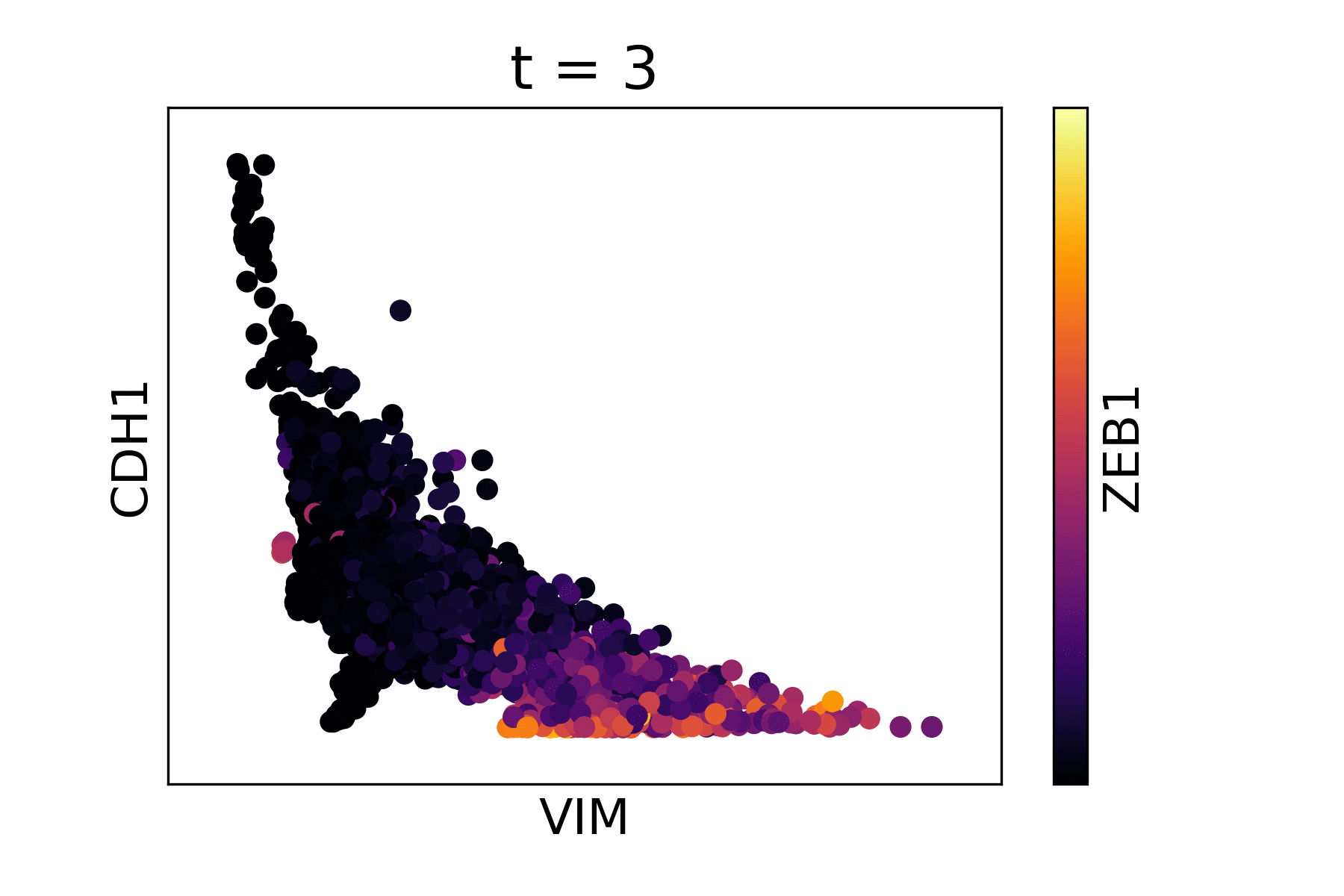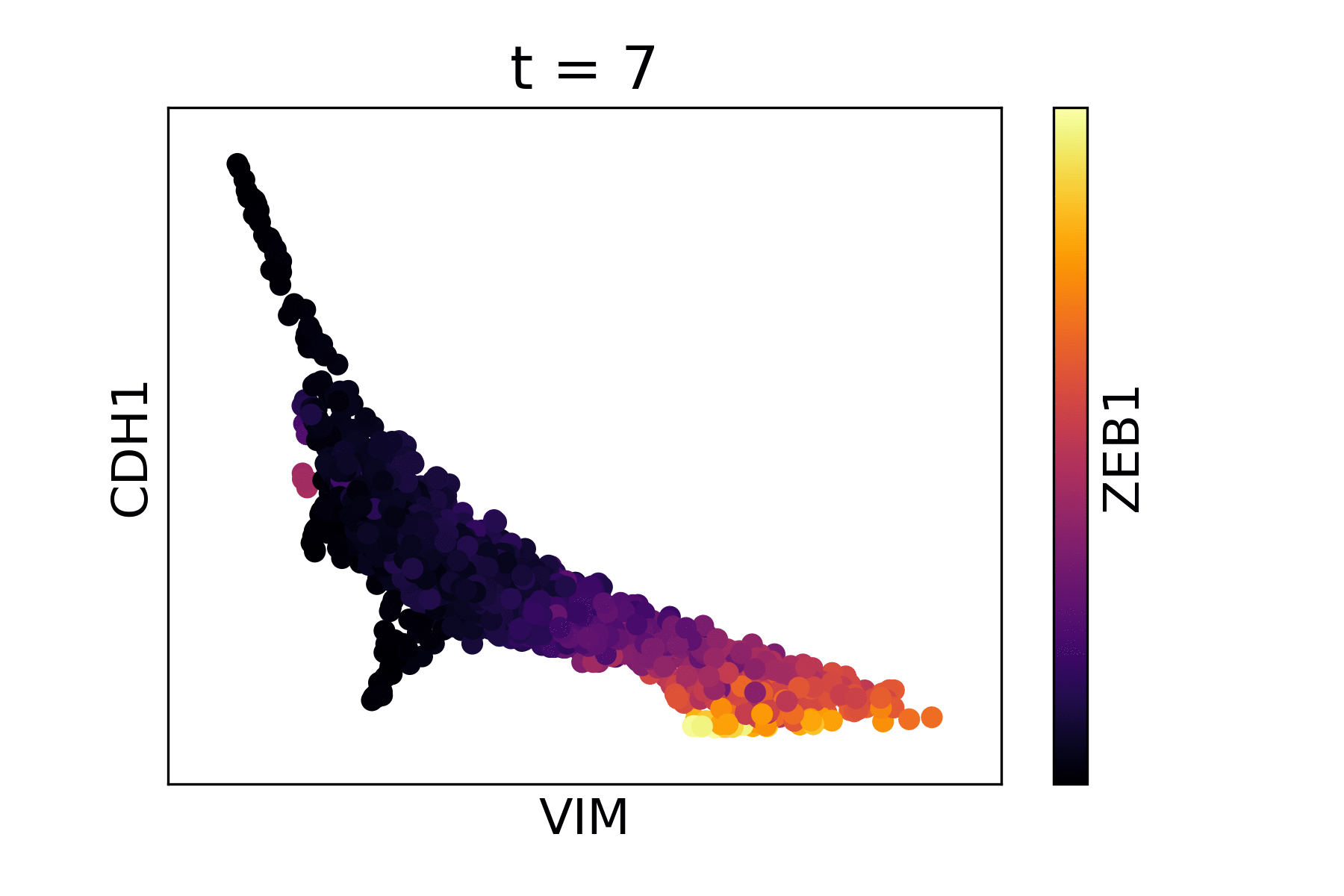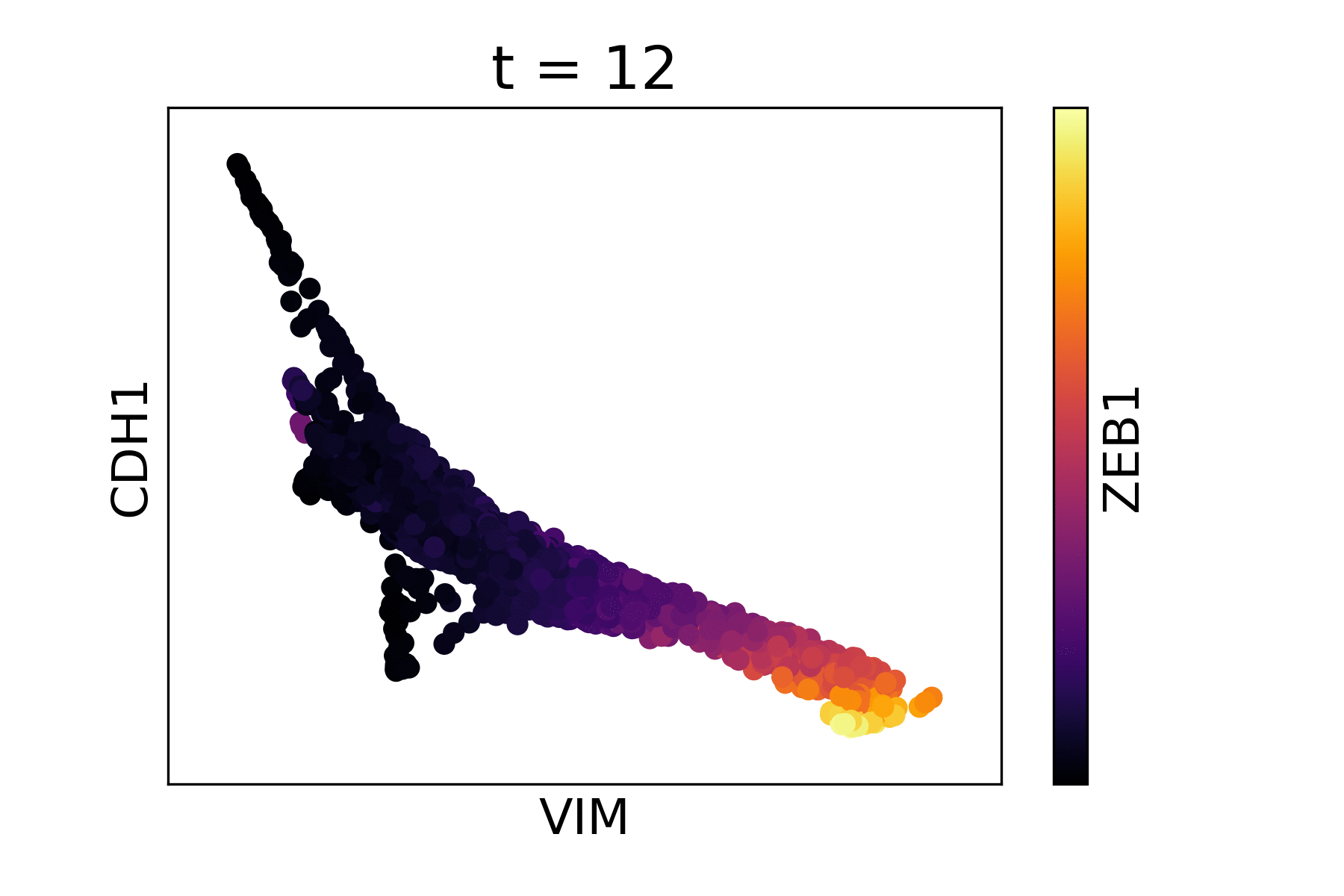
Magic reveals the interaction between Vimentin (VIM), Cadherin-1 (CDH1), and Zinc finger E-box-binding homeobox 1 (ZEB1, encoded by colors).
Table of Contents
Python
Installation
Installation with pip
To install with pip, run the following from a terminal:
pip install --user magic-impute
Installation from GitHub
To clone the repository and install manually, run the following from a terminal:
git clone git://github.com/KrishnaswamyLab/MAGIC.git
cd MAGIC/python
python setup.py install --user
Usage
Quick Start
The following code runs MAGIC on test data located in the MAGIC repository.
import magic
import pandas as pd
import matplotlib.pyplot as plt
X = pd.read_csv("MAGIC/data/test_data.csv")
magic_operator = magic.MAGIC()
X_magic = magic_operator.fit_transform(X, genes=['VIM', 'CDH1', 'ZEB1'])
plt.scatter(X_magic['VIM'], X_magic['CDH1'], c=X_magic['ZEB1'], s=1, cmap='inferno')
plt.show()
magic.plot.animate_magic(X, gene_x='VIM', gene_y='CDH1', gene_color='ZEB1', operator=magic_operator)
Tutorials
You can read the MAGIC documentation at https://magic.readthedocs.io/. We have included two tutorial notebooks on MAGIC usage and results visualization for single cell RNA-seq data.
EMT data notebook: http://nbviewer.jupyter.org/github/KrishnaswamyLab/MAGIC/blob/master/python/tutorialnotebooks/emttutorial.ipynb
Bone Marrow data notebook: http://nbviewer.jupyter.org/github/KrishnaswamyLab/MAGIC/blob/master/python/tutorialnotebooks/bonemarrowtutorial.ipynb
Matlab
Instructions for the Matlab version
- run_magic.m -- MAGIC imputation function
- testmagic.m -- Shows how to run MAGIC. Also included is a function for loading 10x format data (load10x.m)
R
Installation
To use MAGIC, you will need to install both the R and Python packages.
If python or pip are not installed, you will need to install them. We recommend
Miniconda3 to install Python and pip together,
or otherwise you can install pip from https://pip.pypa.io/en/stable/installing/.
Installation from CRAN
In R, run this command to install MAGIC and all dependencies:
install.packages("Rmagic")
In a terminal, run the following command to install the Python repository.
pip install --user magic-impute
Installation from GitHub
To clone the repository and install manually, run the following from a terminal:
git clone git://github.com/KrishnaswamyLab/MAGIC.git
cd MAGIC/python
python setup.py install --user
cd ../Rmagic
R CMD INSTALL .
Usage
Quick Start
After installing the package, MAGIC can be run by loading the library and calling magic():
library(Rmagic)
library(ggplot2)
data(magic_testdata)
MAGIC_data <- magic(magic_testdata, genes=c("VIM", "CDH1", "ZEB1"))
ggplot(MAGIC_data) +
geom_point(aes(x=VIM, y=CDH1, color=ZEB1))
Tutorials
You can read the MAGIC tutorial by running help(Rmagic::magic). For a working example, see the Rmarkdown tutorials at http://htmlpreview.github.io/?https://github.com/KrishnaswamyLab/MAGIC/blob/master/Rmagic/inst/examples/bonemarrow_tutorial.html and http://htmlpreview.github.io/?https://github.com/KrishnaswamyLab/MAGIC/blob/master/Rmagic/inst/examples/emt_tutorial.html or in Rmagic/inst/examples.
Help
If you have any questions or require assistance using MAGIC, please contact us at https://krishnaswamylab.org/get-help.
Owner
- Name: Krishnaswamy Lab
- Login: KrishnaswamyLab
- Kind: organization
- Email: smita.krishnaswamy@yale.edu
- Location: New Haven, CT
- Website: https://krishnaswamylab.org
- Repositories: 31
- Profile: https://github.com/KrishnaswamyLab
GitHub Events
Total
- Issues event: 3
- Watch event: 28
- Issue comment event: 3
- Fork event: 3
Last Year
- Issues event: 3
- Watch event: 28
- Issue comment event: 3
- Fork event: 3
Committers
Last synced: over 2 years ago
Top Committers
| Name | Commits | |
|---|---|---|
| Scott Gigante | s****e@g****m | 374 |
| Pooja Kathail | p****5@c****u | 98 |
| Ngan Vu | n****u@y****u | 40 |
| dvdijk | d****k@g****m | 36 |
| Manu Setty | m****i@g****m | 34 |
| Daniel Dager | d****r@y****u | 24 |
| Daniel Burkhardt | b****b@g****m | 10 |
| Scott Gigante | s****e | 5 |
| Paul Hoffman | p****n@n****g | 4 |
| Ruchira-Ray | 3****y | 4 |
| github-actions[bot] | 4****] | 4 |
| Fabio Zanini | f****i@f****m | 3 |
| pkathail | p****l | 2 |
| lkmklsmn | l****n@g****m | 1 |
| guilhermesena | g****4@g****m | 1 |
| EC2 Default User | e****r@i****l | 1 |
| Cao T | c****8@g****m | 1 |
| Alex Tong | a****g@g****m | 1 |
| Paul Hoffman | m****e | 1 |
Committer Domains (Top 20 + Academic)
Issues and Pull Requests
Last synced: 7 months ago
All Time
- Total issues: 81
- Total pull requests: 32
- Average time to close issues: 2 months
- Average time to close pull requests: 4 days
- Total issue authors: 73
- Total pull request authors: 6
- Average comments per issue: 2.8
- Average comments per pull request: 0.28
- Merged pull requests: 27
- Bot issues: 0
- Bot pull requests: 0
Past Year
- Issues: 3
- Pull requests: 1
- Average time to close issues: about 10 hours
- Average time to close pull requests: 7 minutes
- Issue authors: 2
- Pull request authors: 1
- Average comments per issue: 0.67
- Average comments per pull request: 0.0
- Merged pull requests: 0
- Bot issues: 0
- Bot pull requests: 0
Top Authors
Issue Authors
- HHHit (3)
- buutrg (2)
- yongjieliu (2)
- xingzhongyu (2)
- MichaelPeibo (2)
- abhay0305 (2)
- scottgigante (1)
- zvittorio (1)
- Roger-GOAT (1)
- prummer (1)
- dagarfield (1)
- Natacoman (1)
- k326xh (1)
- saeedfc (1)
- AVitriolo (1)
Pull Request Authors
- scottgigante (22)
- dburkhardt (5)
- dbhaskar92 (4)
- dvdijk (1)
- softbear (1)
- mojaveazure (1)
Top Labels
Issue Labels
Pull Request Labels
Packages
- Total packages: 3
-
Total downloads:
- pypi 1,598 last-month
- Total docker downloads: 485
-
Total dependent packages: 13
(may contain duplicates) -
Total dependent repositories: 22
(may contain duplicates) - Total versions: 53
- Total maintainers: 1
pypi.org: magic-impute
MAGIC
- Homepage: https://github.com/KrishnaswamyLab/MAGIC
- Documentation: https://magic-impute.readthedocs.io/
- License: GNU General Public License Version 2
-
Latest release: 3.0.0
published about 5 years ago
Rankings
Maintainers (1)
proxy.golang.org: github.com/KrishnaswamyLab/MAGIC
- Documentation: https://pkg.go.dev/github.com/KrishnaswamyLab/MAGIC#section-documentation
- License: gpl-2.0
-
Latest release: v3.0.0+incompatible
published about 5 years ago
Rankings
proxy.golang.org: github.com/krishnaswamylab/magic
- Documentation: https://pkg.go.dev/github.com/krishnaswamylab/magic#section-documentation
- License: gpl-2.0
-
Latest release: v3.0.0+incompatible
published about 5 years ago
Rankings
Dependencies
- Matrix >= 1.2 depends
- R >= 3.3 depends
- ggplot2 * imports
- methods * imports
- reticulate >= 1.4 imports
- stats * imports
- Seurat >= 3.0.0 suggests
- phateR * suggests
- readr * suggests
- viridis * suggests
- docopt *
- git2r *
- lintr *
- precommit *
- roxygen2 *
- styler *
- future *
- graphtools >=1.0.0
- matplotlib >=2.0.1
- numpy >=1.14.0
- pandas >=0.25
- scikit-learn >=0.19.1
- scipy >=1.1.0
- scprep >=1.0
- sphinx *
- sphinxcontrib-napoleon *
- future *
- graphtools >=1.4.0
- matplotlib *
- numpy >=1.14.0
- pandas >=0.25
- scikit-learn >=0.19.1
- scipy >=1.1.0
- scprep >=1.0
- tasklogger >=1.0.0
- future *
- graphtools >=1.4.0
- matplotlib *
- numpy >=1.14.0
- pandas >=0.25
- scikit-learn >=0.19.1
- scipy >=1.1.0
- scprep >=1.0
- tasklogger >=1.0.0
- actions/checkout master composite
- actions/setup-python v2 composite
- pypa/gh-action-pypi-publish master composite
- actions/cache v2 composite
- actions/checkout v2 composite
- actions/setup-python v2 composite
- ad-m/github-push-action master composite
- pre-commit/action v2.0.0 composite
- styfle/cancel-workflow-action 0.6.0 composite
- actions/cache v2 composite
- actions/checkout v2 composite
- actions/setup-python v2 composite
- actions/upload-artifact master composite
- r-lib/actions/setup-r v1 composite
- r-lib/actions/setup-tinytex v1 composite
- styfle/cancel-workflow-action 0.6.0 composite





