Science Score: 20.0%
This score indicates how likely this project is to be science-related based on various indicators:
-
○CITATION.cff file
-
○codemeta.json file
-
○.zenodo.json file
-
○DOI references
-
✓Academic publication links
Links to: arxiv.org -
✓Committers with academic emails
2 of 4 committers (50.0%) from academic institutions -
○Institutional organization owner
-
○JOSS paper metadata
-
○Scientific vocabulary similarity
Low similarity (14.6%) to scientific vocabulary
Keywords
Repository
Conditional Auto-Regressive LASSO in R
Basic Info
- Host: GitHub
- Owner: YunyiShen
- License: gpl-3.0
- Language: C++
- Default Branch: master
- Homepage: https://yunyishen.github.io/CAR-LASSO
- Size: 284 MB
Statistics
- Stars: 17
- Watchers: 3
- Forks: 5
- Open Issues: 5
- Releases: 0
Topics
Metadata Files
README.md
CARlasso 
CARlasso: Conditional Auto-Regressive LASSO in R
With this package, users can infer a graph with two types of nodes: 1) for correlated responses (for example, microbial abundances) and 2) for predictors affecting the responses (for example, environmental or experimental conditions).
The advantages of our implementation is that:
- The edges in the graph correspond to conditional dependency (instead of marginal) which agree better with the biological intution behind experiments
- The graph is sparse
- Because the implementation is Bayesian, users can incorporate prior knowledge into the model
Installation
The package is on CRAN, so to install it, please use:
r
install.packages("CARlasso")
To install development version from github:
r
devtools::install_github("YunyiShen/CAR-LASSO")
Dependencies: The CAR-LASSO R package depends on the following R packages: Rcpp, RcppArmadillo, RcppProgress, coda, Matrix, igraph, ggraph, and ggplot2.
Documentation
For more information, please check out the tutorial.
Fitting a CAR-ALASSO network on human gut microbiome data
To run a reduced version of the analysis on human gut microbiome in our paper (with less predictors and responses):
```r library(CARlasso) gutres <- CARlasso(Alistipes+Bacteroides+ Eubacterium+Parabacteroides+allothers~ BMI+Age+Gender+Stratum, data = mgp154,link = "logit", adaptive = TRUE, niter = 5000, nburnin = 1000, thinby = 10)
horseshoe will take a while, as it needs to sample the latent normal too
gutres <- horseshoe(gutres) plot(gut_res) ``` It might take a little while due to the sampling process of the latent normal variable.
We are using the sample human gut microbiome data included in the package (mgp154).
If you want to run this model on your own data, check out the structure of mgp154 to put your data in the same format:
r
str(mgp154)
head(mgp154)
The color of the edge represents the type of correlation (negative=blue, positive=red) and the width of the edge corresponds to the effect size. Response nodes are represented by circles (in this case, microbes) and predictor nodes are represented by triangles (in this case, age, gender, and stratum).
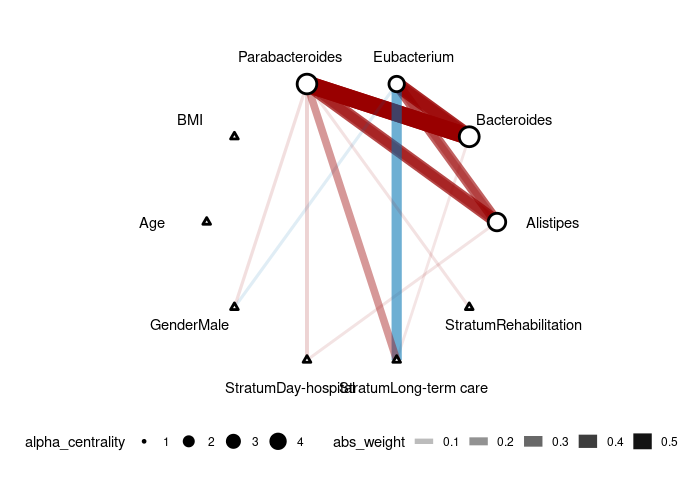
Though we don't recommend treating compositional data as counts, as a illustration, we can run the counting model (link = "log"):
```r gutres <- CARlasso(Alistipes+Bacteroides+ Eubacterium+Parabacteroides+allothers~ BMI+Age+Gender+Stratum, data = mgp154,link = "log", adaptive = TRUE, rbeta = 0.1, # default sometimes cause singularity in Poisson model due to exponential transformation, slightly change can fix it. niter = 5000, nburnin = 1000, thin_by = 10)
horseshoe will take a while, as it's currently implemented in R rather than C++
gutres <- horseshoe(gutres) plot(gut_res) ```
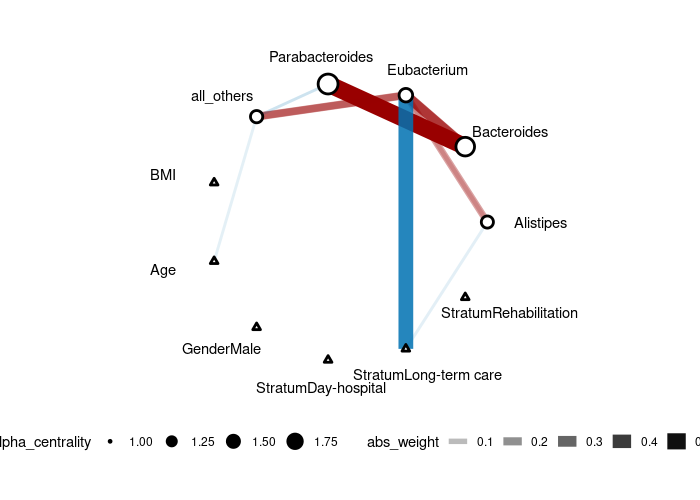
Fitting a CAR-ALASSO network on simulated data
We generate data from a 5-node AR1 model where each node has a specific treatment. Then, we use the adaptive version of CAR-LASSO (CAR-ALASSO) to reconstruct such network and plot the result:
```r set.seed(42) dt <- simuAR1(n=100, k=5, rho=0.7) carres <- CARlasso(y1+y2+y3+y4+y5~x1+x2+x3+x4+x5, data = dt, adaptive = TRUE) plot(car_res,tol = 0.05)
with horseshoe inference
carres <- horseshoe(carres) plot(car_res) ```
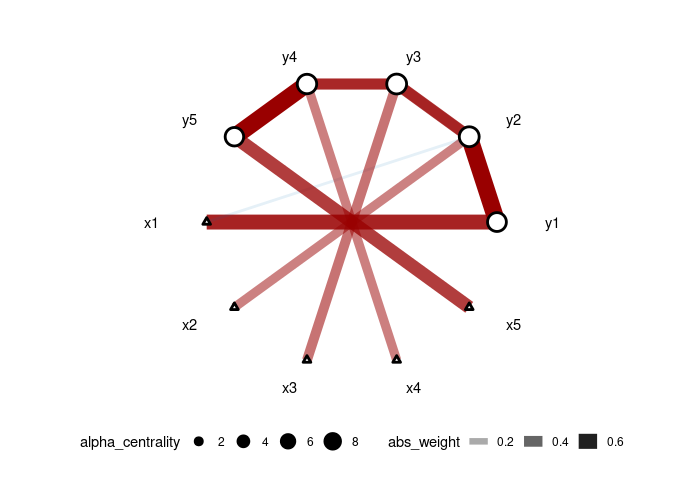
Fitting a standard Graphical LASSO network
Our package also includes functions to fit a standard graphical LASSO, see this page in the tutorial for more details.
Fitting your own hierarchical model
If you would like lower level interface of CAR-LASSO, see this page in the tutorial.
Contributions
Users interested in expanding functionalities in CAR-LASSO R package are welcome to do so. See details on how to contribute in CONTRIBUTING.md.
License
CAR-LASSO R package is licensed under the GNU General Public License v3.0 license.
Citation
If you use the CAR-LASSO R package in your work, we kindly ask that you cite the following paper:
Shen, Y., Solís-Lemus, C. (2020). Bayesian Conditional Auto-Regressive LASSO Models to Learn Sparse Networks with Predictors, arXiv:2012.08397
@article{Shen2020,
title = "Bayesian Conditional {Auto-Regressive} {LASSO} Models to
Learn Sparse Networks with Predictors",
author = "Shen, Yunyi and Solis-Lemus, Claudia",
month = dec,
year = 2020,
archivePrefix = "arXiv",
primaryClass = "stat.AP",
eprint = "2012.08397"
}
Feedback, issues and questions
Feedback, issues and questions are encouraged through the GitHub issue tracker.
Owner
- Name: Yunyi Shen
- Login: YunyiShen
- Kind: user
- Location: Cambridge
- Company: Massachusetts Institute of Technology
- Website: https://yunyishen.github.io
- Twitter: ShenRaphael
- Repositories: 9
- Profile: https://github.com/YunyiShen
PhD student at MIT/LIDS, working on Bayesian nonparametrics.
GitHub Events
Total
- Watch event: 3
Last Year
- Watch event: 3
Committers
Last synced: over 1 year ago
Top Committers
| Name | Commits | |
|---|---|---|
| YunyiShen | y****9@w****u | 340 |
| Yunyi Shen | 3****n | 29 |
| ClaudiaSL | s****s@w****u | 5 |
| Dirk Eddelbuettel | e****d@d****g | 1 |
Committer Domains (Top 20 + Academic)
Issues and Pull Requests
Last synced: almost 2 years ago
All Time
- Total issues: 5
- Total pull requests: 7
- Average time to close issues: N/A
- Average time to close pull requests: 1 day
- Total issue authors: 4
- Total pull request authors: 2
- Average comments per issue: 1.6
- Average comments per pull request: 0.29
- Merged pull requests: 7
- Bot issues: 0
- Bot pull requests: 0
Past Year
- Issues: 1
- Pull requests: 0
- Average time to close issues: N/A
- Average time to close pull requests: N/A
- Issue authors: 1
- Pull request authors: 0
- Average comments per issue: 1.0
- Average comments per pull request: 0
- Merged pull requests: 0
- Bot issues: 0
- Bot pull requests: 0
Top Authors
Issue Authors
- YunyiShen (2)
- connor-morozumi (1)
- Sophiayangg (1)
- kjmtaylor22 (1)
Pull Request Authors
- crsl4 (6)
- eddelbuettel (1)
Top Labels
Issue Labels
Pull Request Labels
Packages
- Total packages: 1
-
Total downloads:
- cran 180 last-month
- Total docker downloads: 74
- Total dependent packages: 0
- Total dependent repositories: 0
- Total versions: 3
- Total maintainers: 1
cran.r-project.org: CARlasso
Conditional Autoregressive LASSO
- Homepage: https://github.com/YunyiShen/CAR-LASSO
- Documentation: http://cran.r-project.org/web/packages/CARlasso/CARlasso.pdf
- License: GPL-3
- Status: removed
-
Latest release: 0.1.2
published almost 5 years ago
Rankings
Maintainers (1)
Dependencies
- R >= 3.5.0 depends
- MASS * imports
- Matrix * imports
- Rcpp * imports
- coda * imports
- ggplot2 * imports
- ggraph * imports
- igraph * imports
- methods * imports
- knitr * suggests
- rmarkdown * suggests
- testthat * suggests
- actions/cache v2 composite
- actions/checkout v2 composite
- actions/upload-artifact main composite
- r-lib/actions/setup-pandoc v1 composite
- r-lib/actions/setup-r v1 composite